Zonghai Yao
Rethinking Patient Education as Multi-turn Multi-modal Interaction
Apr 16, 2026Abstract:Most medical multimodal benchmarks focus on static tasks such as image question answering, report generation, and plain-language rewriting. Patient education is more demanding: systems must identify relevant evidence across images, show patients where to look, explain findings in accessible language, and handle confusion or distress. Yet most patient education work remains text-only, even though combined image-and-text explanations may better support understanding. We introduce MedImageEdu, a benchmark for multi-turn, evidence-grounded radiology patient education. Each case provides a radiology report with report text and case images. A DoctorAgent interacts with a PatientAgent, conditioned on a hidden profile that captures factors such as education level, health literacy, and personality. When a patient question would benefit from visual support, the DoctorAgent can issue drawing instructions grounded in the report, case images, and the current question to a benchmark-provided drawing tool. The tool returns image(s), after which the DoctorAgent produces a final multimodal response consisting of the image(s) and a grounded plain-language explanation. MedImageEdu contains 150 cases from three sources and evaluates both the consultation process and the final multimodal response along five dimensions: Consultation, Safety and Scope, Language Quality, Drawing Quality, and Image-Text Response Quality. Across representative open- and closed-source vision-language model agents, we find three consistent gaps: fluent language often outpaces faithful visual grounding, safety is the weakest dimension across disease categories, and emotionally tense interactions are harder than low education or low health literacy. MedImageEdu provides a controlled testbed for assessing whether multimodal agents can teach from evidence rather than merely answer from text.
Efficient and Effective Internal Memory Retrieval for LLM-Based Healthcare Prediction
Apr 08, 2026Abstract:Large language models (LLMs) hold significant promise for healthcare, yet their reliability in high-stakes clinical settings is often compromised by hallucinations and a lack of granular medical context. While Retrieval Augmented Generation (RAG) can mitigate these issues, standard supervised pipelines require computationally intensive searches over massive external knowledge bases, leading to high latency that is impractical for time-sensitive care. To address this, we introduce Keys to Knowledge (K2K), a novel framework that replaces external retrieval with internal, key-based knowledge access. By encoding essential clinical information directly into the model's parameter space, K2K enables rapid retrieval from internal key-value memory without inference-time overhead. We further enhance retrieval quality through activation-guided probe construction and cross-attention reranking. Experimental results demonstrate that K2K achieves state-of-the-art performance across four benchmark healthcare outcome prediction datasets.
Knowing When to Abstain: Medical LLMs Under Clinical Uncertainty
Jan 18, 2026Abstract:Current evaluation of large language models (LLMs) overwhelmingly prioritizes accuracy; however, in real-world and safety-critical applications, the ability to abstain when uncertain is equally vital for trustworthy deployment. We introduce MedAbstain, a unified benchmark and evaluation protocol for abstention in medical multiple-choice question answering (MCQA) -- a discrete-choice setting that generalizes to agentic action selection -- integrating conformal prediction, adversarial question perturbations, and explicit abstention options. Our systematic evaluation of both open- and closed-source LLMs reveals that even state-of-the-art, high-accuracy models often fail to abstain with uncertain. Notably, providing explicit abstention options consistently increases model uncertainty and safer abstention, far more than input perturbations, while scaling model size or advanced prompting brings little improvement. These findings highlight the central role of abstention mechanisms for trustworthy LLM deployment and offer practical guidance for improving safety in high-stakes applications.
PRIME: Planning and Retrieval-Integrated Memory for Enhanced Reasoning
Sep 26, 2025Abstract:Inspired by the dual-process theory of human cognition from \textit{Thinking, Fast and Slow}, we introduce \textbf{PRIME} (Planning and Retrieval-Integrated Memory for Enhanced Reasoning), a multi-agent reasoning framework that dynamically integrates \textbf{System 1} (fast, intuitive thinking) and \textbf{System 2} (slow, deliberate thinking). PRIME first employs a Quick Thinking Agent (System 1) to generate a rapid answer; if uncertainty is detected, it then triggers a structured System 2 reasoning pipeline composed of specialized agents for \textit{planning}, \textit{hypothesis generation}, \textit{retrieval}, \textit{information integration}, and \textit{decision-making}. This multi-agent design faithfully mimics human cognitive processes and enhances both efficiency and accuracy. Experimental results with LLaMA 3 models demonstrate that PRIME enables open-source LLMs to perform competitively with state-of-the-art closed-source models like GPT-4 and GPT-4o on benchmarks requiring multi-hop and knowledge-grounded reasoning. This research establishes PRIME as a scalable solution for improving LLMs in domains requiring complex, knowledge-intensive reasoning.
DischargeSim: A Simulation Benchmark for Educational Doctor-Patient Communication at Discharge
Sep 10, 2025Abstract:Discharge communication is a critical yet underexplored component of patient care, where the goal shifts from diagnosis to education. While recent large language model (LLM) benchmarks emphasize in-visit diagnostic reasoning, they fail to evaluate models' ability to support patients after the visit. We introduce DischargeSim, a novel benchmark that evaluates LLMs on their ability to act as personalized discharge educators. DischargeSim simulates post-visit, multi-turn conversations between LLM-driven DoctorAgents and PatientAgents with diverse psychosocial profiles (e.g., health literacy, education, emotion). Interactions are structured across six clinically grounded discharge topics and assessed along three axes: (1) dialogue quality via automatic and LLM-as-judge evaluation, (2) personalized document generation including free-text summaries and structured AHRQ checklists, and (3) patient comprehension through a downstream multiple-choice exam. Experiments across 18 LLMs reveal significant gaps in discharge education capability, with performance varying widely across patient profiles. Notably, model size does not always yield better education outcomes, highlighting trade-offs in strategy use and content prioritization. DischargeSim offers a first step toward benchmarking LLMs in post-visit clinical education and promoting equitable, personalized patient support.
ChatThero: An LLM-Supported Chatbot for Behavior Change and Therapeutic Support in Addiction Recovery
Aug 28, 2025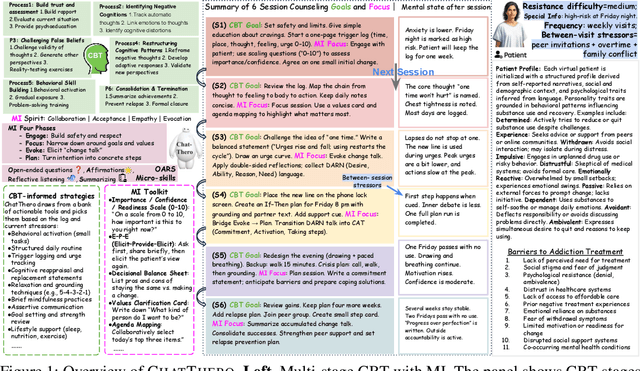
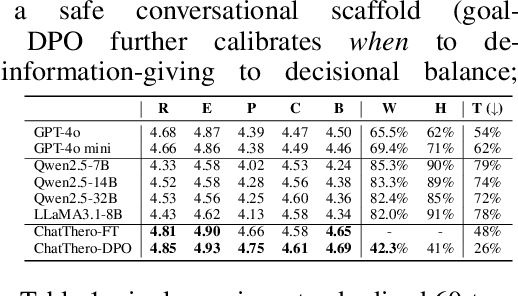
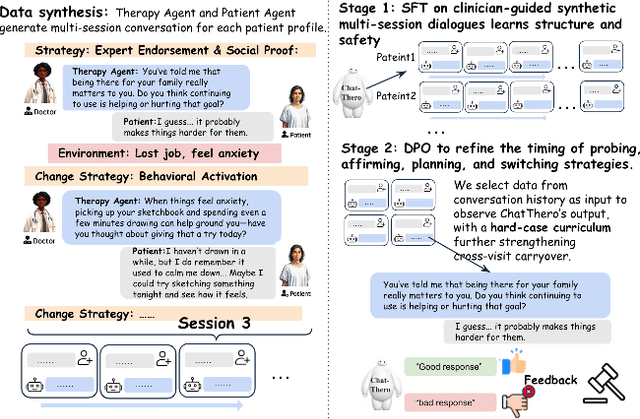

Abstract:Substance use disorders (SUDs) affect over 36 million people worldwide, yet few receive effective care due to stigma, motivational barriers, and limited personalized support. Although large language models (LLMs) show promise for mental-health assistance, most systems lack tight integration with clinically validated strategies, reducing effectiveness in addiction recovery. We present ChatThero, a multi-agent conversational framework that couples dynamic patient modeling with context-sensitive therapeutic dialogue and adaptive persuasive strategies grounded in cognitive behavioral therapy (CBT) and motivational interviewing (MI). We build a high-fidelity synthetic benchmark spanning Easy, Medium, and Hard resistance levels, and train ChatThero with a two-stage pipeline comprising supervised fine-tuning (SFT) followed by direct preference optimization (DPO). In evaluation, ChatThero yields a 41.5\% average gain in patient motivation, a 0.49\% increase in treatment confidence, and resolves hard cases with 26\% fewer turns than GPT-4o, and both automated and human clinical assessments rate it higher in empathy, responsiveness, and behavioral realism. The framework supports rigorous, privacy-preserving study of therapeutic conversation and provides a robust, replicable basis for research and clinical translation.
SynthEHR-Eviction: Enhancing Eviction SDoH Detection with LLM-Augmented Synthetic EHR Data
Jul 10, 2025Abstract:Eviction is a significant yet understudied social determinants of health (SDoH), linked to housing instability, unemployment, and mental health. While eviction appears in unstructured electronic health records (EHRs), it is rarely coded in structured fields, limiting downstream applications. We introduce SynthEHR-Eviction, a scalable pipeline combining LLMs, human-in-the-loop annotation, and automated prompt optimization (APO) to extract eviction statuses from clinical notes. Using this pipeline, we created the largest public eviction-related SDoH dataset to date, comprising 14 fine-grained categories. Fine-tuned LLMs (e.g., Qwen2.5, LLaMA3) trained on SynthEHR-Eviction achieved Macro-F1 scores of 88.8% (eviction) and 90.3% (other SDoH) on human validated data, outperforming GPT-4o-APO (87.8%, 87.3%), GPT-4o-mini-APO (69.1%, 78.1%), and BioBERT (60.7%, 68.3%), while enabling cost-effective deployment across various model sizes. The pipeline reduces annotation effort by over 80%, accelerates dataset creation, enables scalable eviction detection, and generalizes to other information extraction tasks.
MedReadCtrl: Personalizing medical text generation with readability-controlled instruction learning
Jul 10, 2025Abstract:Generative AI has demonstrated strong potential in healthcare, from clinical decision support to patient-facing chatbots that improve outcomes. A critical challenge for deployment is effective human-AI communication, where content must be both personalized and understandable. We introduce MedReadCtrl, a readability-controlled instruction tuning framework that enables LLMs to adjust output complexity without compromising meaning. Evaluations of nine datasets and three tasks across medical and general domains show that MedReadCtrl achieves significantly lower readability instruction-following errors than GPT-4 (e.g., 1.39 vs. 1.59 on ReadMe, p<0.001) and delivers substantial gains on unseen clinical tasks (e.g., +14.7 ROUGE-L, +6.18 SARI on MTSamples). Experts consistently preferred MedReadCtrl (71.7% vs. 23.3%), especially at low literacy levels. These gains reflect MedReadCtrl's ability to restructure clinical content into accessible, readability-aligned language while preserving medical intent, offering a scalable solution to support patient education and expand equitable access to AI-enabled care.
Enhancing LLMs for Identifying and Prioritizing Important Medical Jargons from Electronic Health Record Notes Utilizing Data Augmentation
Feb 25, 2025Abstract:OpenNotes enables patients to access EHR notes, but medical jargon can hinder comprehension. To improve understanding, we evaluated closed- and open-source LLMs for extracting and prioritizing key medical terms using prompting, fine-tuning, and data augmentation. We assessed LLMs on 106 expert-annotated EHR notes, experimenting with (i) general vs. structured prompts, (ii) zero-shot vs. few-shot prompting, (iii) fine-tuning, and (iv) data augmentation. To enhance open-source models in low-resource settings, we used ChatGPT for data augmentation and applied ranking techniques. We incrementally increased the augmented dataset size (10 to 10,000) and conducted 5-fold cross-validation, reporting F1 score and Mean Reciprocal Rank (MRR). Our result show that fine-tuning and data augmentation improved performance over other strategies. GPT-4 Turbo achieved the highest F1 (0.433), while Mistral7B with data augmentation had the highest MRR (0.746). Open-source models, when fine-tuned or augmented, outperformed closed-source models. Notably, the best F1 and MRR scores did not always align. Few-shot prompting outperformed zero-shot in vanilla models, and structured prompts yielded different preferences across models. Fine-tuning improved zero-shot performance but sometimes degraded few-shot performance. Data augmentation performed comparably or better than other methods. Our evaluation highlights the effectiveness of prompting, fine-tuning, and data augmentation in improving model performance for medical jargon extraction in low-resource scenarios.
RARE: Retrieval-Augmented Reasoning Enhancement for Large Language Models
Dec 05, 2024Abstract:This work introduces RARE (Retrieval-Augmented Reasoning Enhancement), a versatile extension to the mutual reasoning framework (rStar), aimed at enhancing reasoning accuracy and factual integrity across large language models (LLMs) for complex, knowledge-intensive tasks such as commonsense and medical reasoning. RARE incorporates two innovative actions within the Monte Carlo Tree Search (MCTS) framework: A6, which generates search queries based on the initial problem statement, performs information retrieval using those queries, and augments reasoning with the retrieved data to formulate the final answer; and A7, which leverages information retrieval specifically for generated sub-questions and re-answers these sub-questions with the relevant contextual information. Additionally, a Retrieval-Augmented Factuality Scorer is proposed to replace the original discriminator, prioritizing reasoning paths that meet high standards of factuality. Experimental results with LLaMA 3.1 show that RARE enables open-source LLMs to achieve competitive performance with top open-source models like GPT-4 and GPT-4o. This research establishes RARE as a scalable solution for improving LLMs in domains where logical coherence and factual integrity are critical.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge