Vassilis N. Ioannidis
SpotTarget: Rethinking the Effect of Target Edges for Link Prediction in Graph Neural Networks
Jun 01, 2023



Abstract:Graph Neural Networks (GNNs) have demonstrated promising outcomes across various tasks, including node classification and link prediction. Despite their remarkable success in various high-impact applications, we have identified three common pitfalls in message passing for link prediction. Particularly, in prevalent GNN frameworks (e.g., DGL and PyTorch-Geometric), the target edges (i.e., the edges being predicted) consistently exist as message passing edges in the graph during training. Consequently, this results in overfitting and distribution shift, both of which adversely impact the generalizability to test the target edges. Additionally, during test time, the failure to exclude the test target edges leads to implicit test leakage caused by neighborhood aggregation. In this paper, we analyze these three pitfalls and investigate the impact of including or excluding target edges on the performance of nodes with varying degrees during training and test phases. Our theoretical and empirical analysis demonstrates that low-degree nodes are more susceptible to these pitfalls. These pitfalls can have detrimental consequences when GNNs are implemented in production systems. To systematically address these pitfalls, we propose SpotTarget, an effective and efficient GNN training framework. During training, SpotTarget leverages our insight regarding low-degree nodes and excludes train target edges connected to at least one low-degree node. During test time, it emulates real-world scenarios of GNN usage in production and excludes all test target edges. Our experiments conducted on diverse real-world datasets, demonstrate that SpotTarget significantly enhances GNNs, achieving up to a 15x increase in accuracy in sparse graphs. Furthermore, SpotTarget consistently and dramatically improves the performance for low-degree nodes in dense graphs.
Train Your Own GNN Teacher: Graph-Aware Distillation on Textual Graphs
Apr 20, 2023



Abstract:How can we learn effective node representations on textual graphs? Graph Neural Networks (GNNs) that use Language Models (LMs) to encode textual information of graphs achieve state-of-the-art performance in many node classification tasks. Yet, combining GNNs with LMs has not been widely explored for practical deployments due to its scalability issues. In this work, we tackle this challenge by developing a Graph-Aware Distillation framework (GRAD) to encode graph structures into an LM for graph-free, fast inference. Different from conventional knowledge distillation, GRAD jointly optimizes a GNN teacher and a graph-free student over the graph's nodes via a shared LM. This encourages the graph-free student to exploit graph information encoded by the GNN teacher while at the same time, enables the GNN teacher to better leverage textual information from unlabeled nodes. As a result, the teacher and the student models learn from each other to improve their overall performance. Experiments in eight node classification benchmarks in both transductive and inductive settings showcase GRAD's superiority over existing distillation approaches for textual graphs.
OrthoReg: Improving Graph-regularized MLPs via Orthogonality Regularization
Jan 31, 2023



Abstract:Graph Neural Networks (GNNs) are currently dominating in modeling graph-structure data, while their high reliance on graph structure for inference significantly impedes them from widespread applications. By contrast, Graph-regularized MLPs (GR-MLPs) implicitly inject the graph structure information into model weights, while their performance can hardly match that of GNNs in most tasks. This motivates us to study the causes of the limited performance of GR-MLPs. In this paper, we first demonstrate that node embeddings learned from conventional GR-MLPs suffer from dimensional collapse, a phenomenon in which the largest a few eigenvalues dominate the embedding space, through empirical observations and theoretical analysis. As a result, the expressive power of the learned node representations is constrained. We further propose OrthoReg, a novel GR-MLP model to mitigate the dimensional collapse issue. Through a soft regularization loss on the correlation matrix of node embeddings, OrthoReg explicitly encourages orthogonal node representations and thus can naturally avoid dimensionally collapsed representations. Experiments on traditional transductive semi-supervised classification tasks and inductive node classification for cold-start scenarios demonstrate its effectiveness and superiority.
Predicting Cellular Responses with Variational Causal Inference and Refined Relational Information
Sep 30, 2022
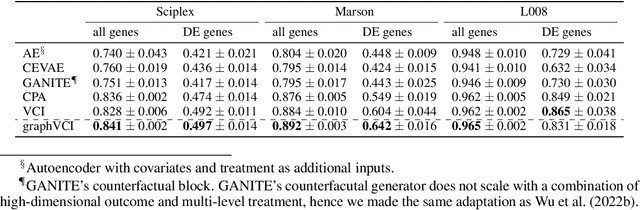

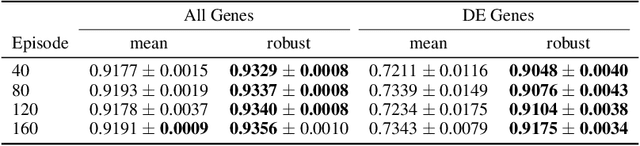
Abstract:Predicting the responses of a cell under perturbations may bring important benefits to drug discovery and personalized therapeutics. In this work, we propose a novel graph variational Bayesian causal inference framework to predict a cell's gene expressions under counterfactual perturbations (perturbations that this cell did not factually receive), leveraging information representing biological knowledge in the form of gene regulatory networks (GRNs) to aid individualized cellular response predictions. Aiming at a data-adaptive GRN, we also developed an adjacency matrix updating technique for graph convolutional networks and used it to refine GRNs during pre-training, which generated more insights on gene relations and enhanced model performance. Additionally, we propose a robust estimator within our framework for the asymptotically efficient estimation of marginal perturbation effect, which is yet to be carried out in previous works. With extensive experiments, we exhibited the advantage of our approach over state-of-the-art deep learning models for individual response prediction.
Variational Causal Inference
Sep 13, 2022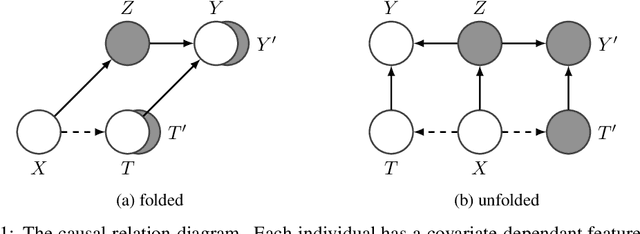
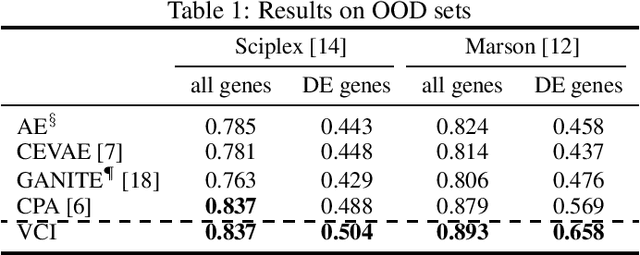
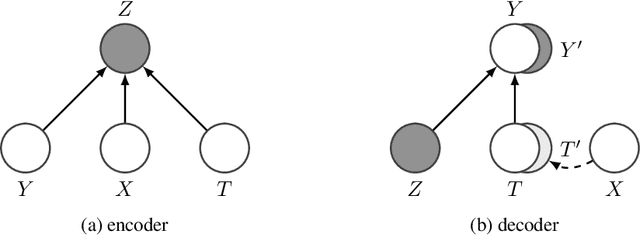
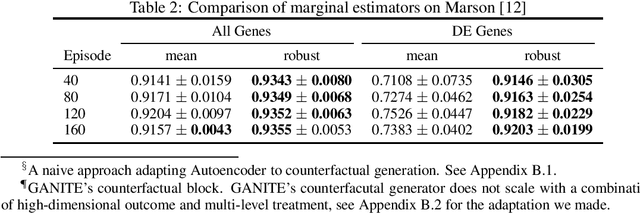
Abstract:Estimating an individual's potential outcomes under counterfactual treatments is a challenging task for traditional causal inference and supervised learning approaches when the outcome is high-dimensional (e.g. gene expressions, impulse responses, human faces) and covariates are relatively limited. In this case, to construct one's outcome under a counterfactual treatment, it is crucial to leverage individual information contained in its observed factual outcome on top of the covariates. We propose a deep variational Bayesian framework that rigorously integrates two main sources of information for outcome construction under a counterfactual treatment: one source is the individual features embedded in the high-dimensional factual outcome; the other source is the response distribution of similar subjects (subjects with the same covariates) that factually received this treatment of interest.
Efficient and effective training of language and graph neural network models
Jun 22, 2022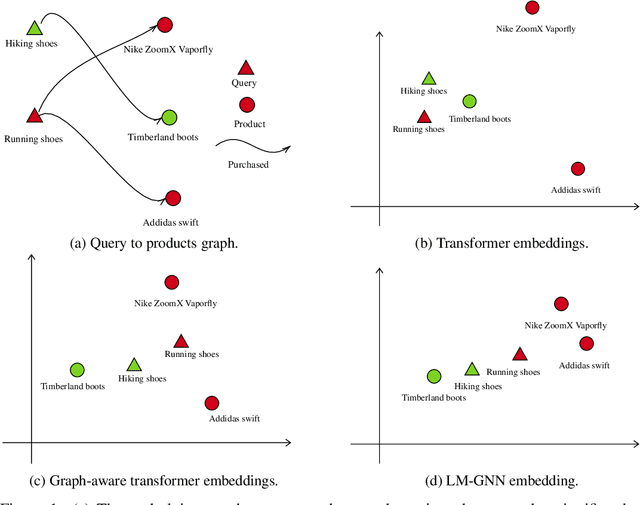
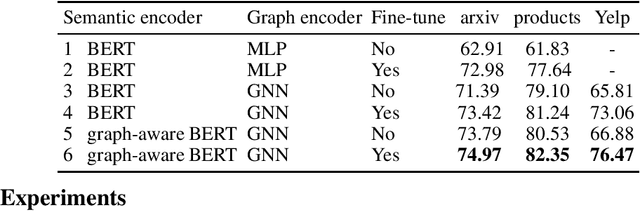
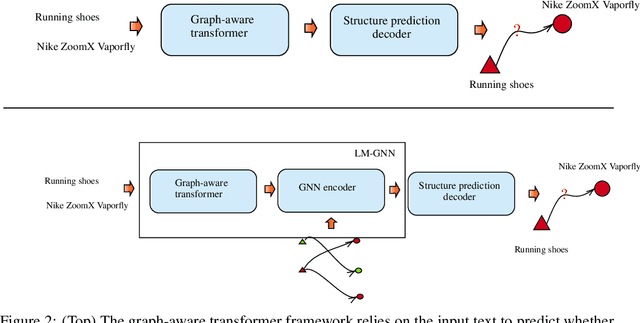
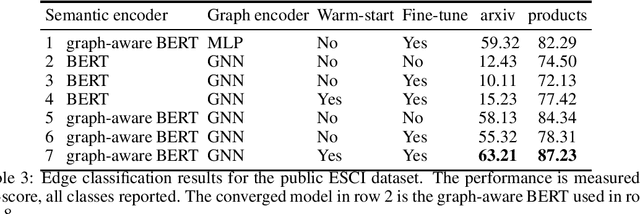
Abstract:Can we combine heterogenous graph structure with text to learn high-quality semantic and behavioural representations? Graph neural networks (GNN)s encode numerical node attributes and graph structure to achieve impressive performance in a variety of supervised learning tasks. Current GNN approaches are challenged by textual features, which typically need to be encoded to a numerical vector before provided to the GNN that may incur some information loss. In this paper, we put forth an efficient and effective framework termed language model GNN (LM-GNN) to jointly train large-scale language models and graph neural networks. The effectiveness in our framework is achieved by applying stage-wise fine-tuning of the BERT model first with heterogenous graph information and then with a GNN model. Several system and design optimizations are proposed to enable scalable and efficient training. LM-GNN accommodates node and edge classification as well as link prediction tasks. We evaluate the LM-GNN framework in different datasets performance and showcase the effectiveness of the proposed approach. LM-GNN provides competitive results in an Amazon query-purchase-product application.
A Robust Stacking Framework for Training Deep Graph Models with Multifaceted Node Features
Jun 16, 2022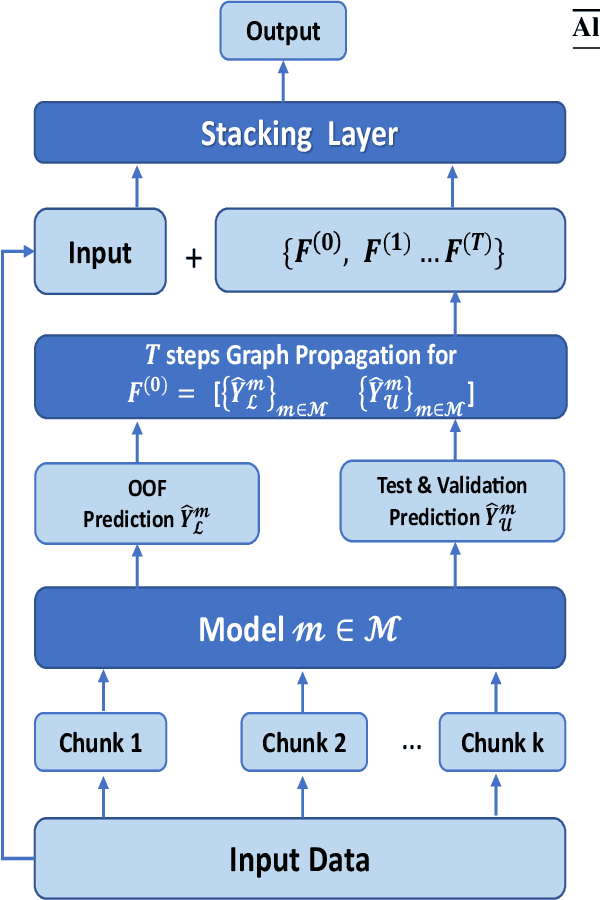
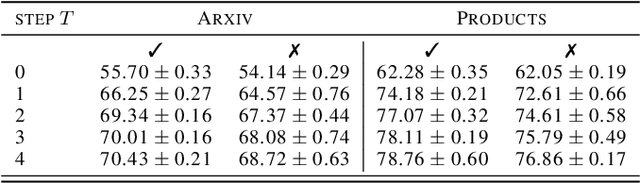
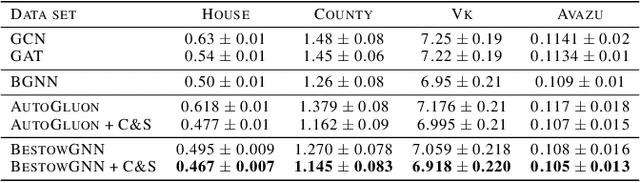
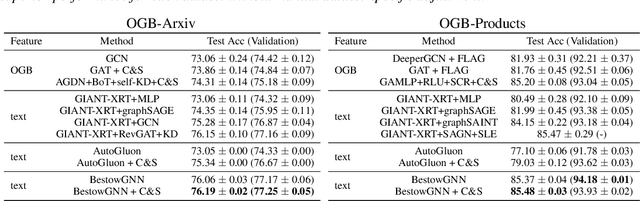

Abstract:Graph Neural Networks (GNNs) with numerical node features and graph structure as inputs have demonstrated superior performance on various supervised learning tasks with graph data. However the numerical node features utilized by GNNs are commonly extracted from raw data which is of text or tabular (numeric/categorical) type in most real-world applications. The best models for such data types in most standard supervised learning settings with IID (non-graph) data are not simple neural network layers and thus are not easily incorporated into a GNN. Here we propose a robust stacking framework that fuses graph-aware propagation with arbitrary models intended for IID data, which are ensembled and stacked in multiple layers. Our layer-wise framework leverages bagging and stacking strategies to enjoy strong generalization, in a manner which effectively mitigates label leakage and overfitting. Across a variety of graph datasets with tabular/text node features, our method achieves comparable or superior performance relative to both tabular/text and graph neural network models, as well as existing state-of-the-art hybrid strategies that combine the two.
TempoQR: Temporal Question Reasoning over Knowledge Graphs
Dec 10, 2021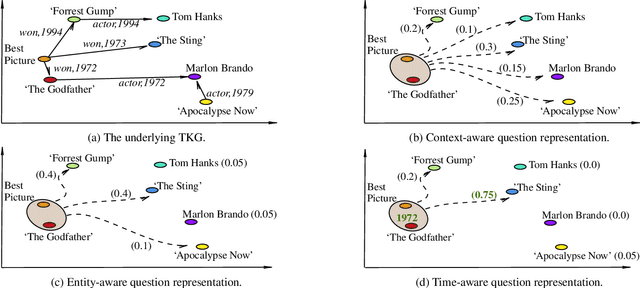

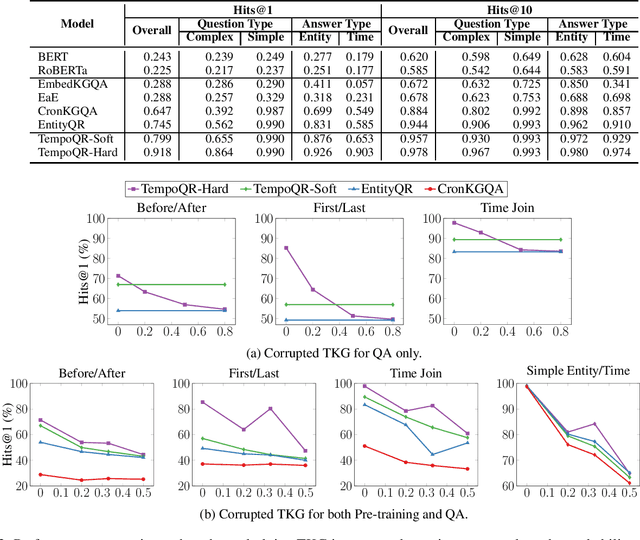
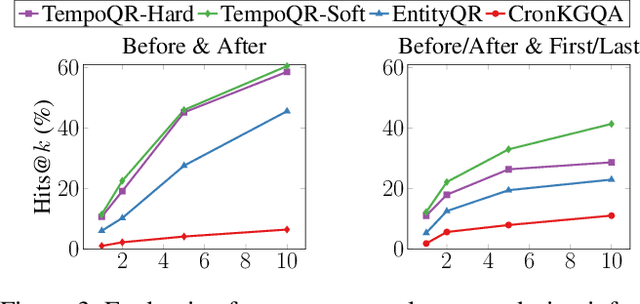
Abstract:Knowledge Graph Question Answering (KGQA) involves retrieving facts from a Knowledge Graph (KG) using natural language queries. A KG is a curated set of facts consisting of entities linked by relations. Certain facts include also temporal information forming a Temporal KG (TKG). Although many natural questions involve explicit or implicit time constraints, question answering (QA) over TKGs has been a relatively unexplored area. Existing solutions are mainly designed for simple temporal questions that can be answered directly by a single TKG fact. This paper puts forth a comprehensive embedding-based framework for answering complex questions over TKGs. Our method termed temporal question reasoning (TempoQR) exploits TKG embeddings to ground the question to the specific entities and time scope it refers to. It does so by augmenting the question embeddings with context, entity and time-aware information by employing three specialized modules. The first computes a textual representation of a given question, the second combines it with the entity embeddings for entities involved in the question, and the third generates question-specific time embeddings. Finally, a transformer-based encoder learns to fuse the generated temporal information with the question representation, which is used for answer predictions. Extensive experiments show that TempoQR improves accuracy by 25--45 percentage points on complex temporal questions over state-of-the-art approaches and it generalizes better to unseen question types.
Convergent Boosted Smoothing for Modeling Graph Data with Tabular Node Features
Oct 26, 2021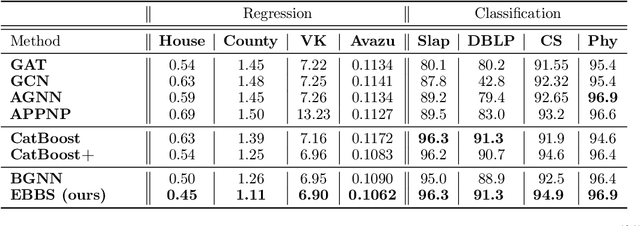
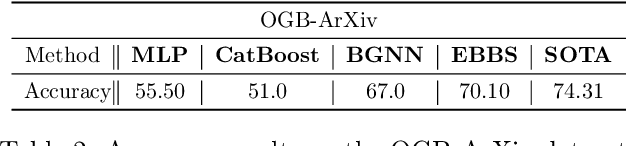
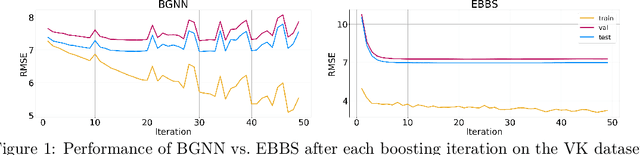

Abstract:For supervised learning with tabular data, decision tree ensembles produced via boosting techniques generally dominate real-world applications involving iid training/test sets. However for graph data where the iid assumption is violated due to structured relations between samples, it remains unclear how to best incorporate this structure within existing boosting pipelines. To this end, we propose a generalized framework for iterating boosting with graph propagation steps that share node/sample information across edges connecting related samples. Unlike previous efforts to integrate graph-based models with boosting, our approach is anchored in a principled meta loss function such that provable convergence can be guaranteed under relatively mild assumptions. Across a variety of non-iid graph datasets with tabular node features, our method achieves comparable or superior performance than both tabular and graph neural network models, as well as existing hybrid strategies that combine the two. Beyond producing better predictive performance than recently proposed graph models, our proposed techniques are easy to implement, computationally more efficient, and enjoy stronger theoretical guarantees (which make our results more reproducible).
Scalable Consistency Training for Graph Neural Networks via Self-Ensemble Self-Distillation
Oct 12, 2021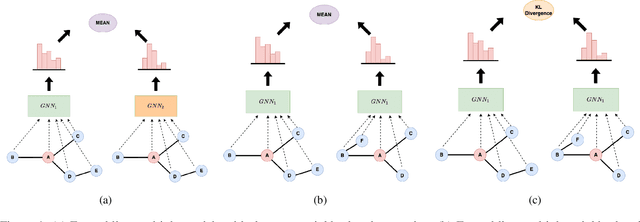
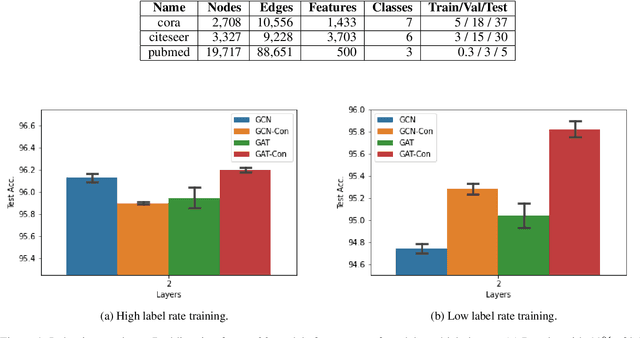
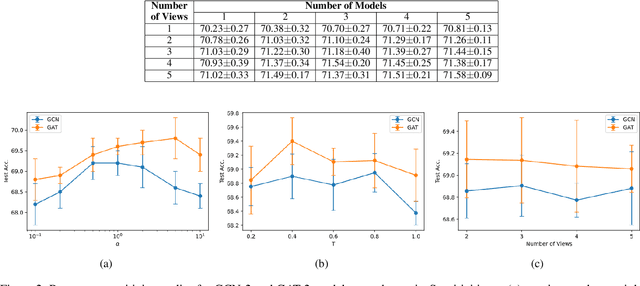
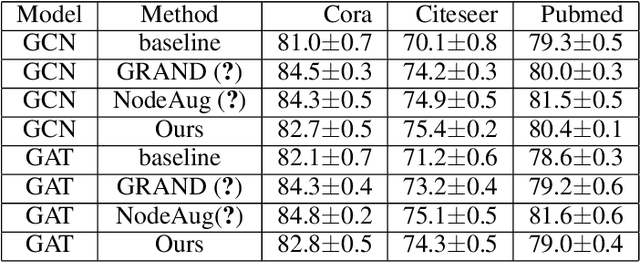
Abstract:Consistency training is a popular method to improve deep learning models in computer vision and natural language processing. Graph neural networks (GNNs) have achieved remarkable performance in a variety of network science learning tasks, but to date no work has studied the effect of consistency training on large-scale graph problems. GNNs scale to large graphs by minibatch training and subsample node neighbors to deal with high degree nodes. We utilize the randomness inherent in the subsampling of neighbors and introduce a novel consistency training method to improve accuracy. For a target node we generate different neighborhood expansions, and distill the knowledge of the average of the predictions to the GNN. Our method approximates the expected prediction of the possible neighborhood samples and practically only requires a few samples. We demonstrate that our training method outperforms standard GNN training in several different settings, and yields the largest gains when label rates are low.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge