Thomas Yu
Signal Processing Laboratory 5
The Brain Tumor Segmentation Challenge 2023: Brain MR Image Synthesis for Tumor Segmentation
May 20, 2023
Abstract:Automated brain tumor segmentation methods are well established, reaching performance levels with clear clinical utility. Most algorithms require four input magnetic resonance imaging (MRI) modalities, typically T1-weighted images with and without contrast enhancement, T2-weighted images, and FLAIR images. However, some of these sequences are often missing in clinical practice, e.g., because of time constraints and/or image artifacts (such as patient motion). Therefore, substituting missing modalities to recover segmentation performance in these scenarios is highly desirable and necessary for the more widespread adoption of such algorithms in clinical routine. In this work, we report the set-up of the Brain MR Image Synthesis Benchmark (BraSyn), organized in conjunction with the Medical Image Computing and Computer-Assisted Intervention (MICCAI) 2023. The objective of the challenge is to benchmark image synthesis methods that realistically synthesize missing MRI modalities given multiple available images to facilitate automated brain tumor segmentation pipelines. The image dataset is multi-modal and diverse, created in collaboration with various hospitals and research institutions.
Slice estimation in diffusion MRI of neonatal and fetal brains in image and spherical harmonics domains using autoencoders
Aug 29, 2022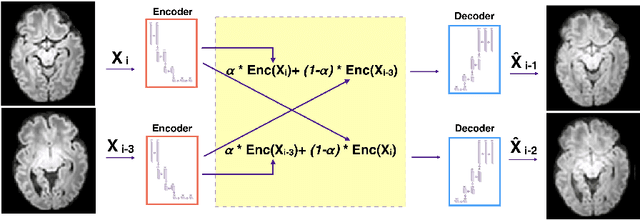
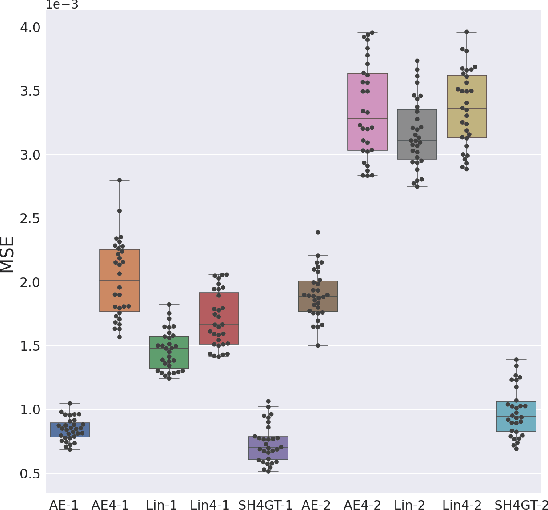
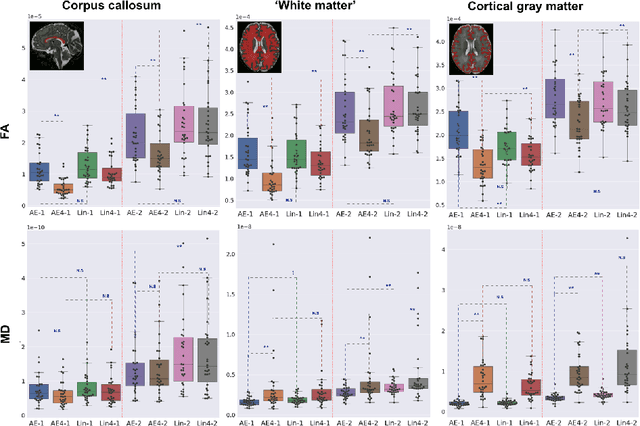
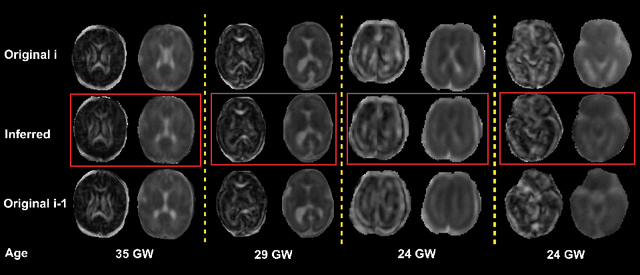
Abstract:Diffusion MRI (dMRI) of the developing brain can provide valuable insights into the white matter development. However, slice thickness in fetal dMRI is typically high (i.e., 3-5 mm) to freeze the in-plane motion, which reduces the sensitivity of the dMRI signal to the underlying anatomy. In this study, we aim at overcoming this problem by using autoencoders to learn unsupervised efficient representations of brain slices in a latent space, using raw dMRI signals and their spherical harmonics (SH) representation. We first learn and quantitatively validate the autoencoders on the developing Human Connectome Project pre-term newborn data, and further test the method on fetal data. Our results show that the autoencoder in the signal domain better synthesized the raw signal. Interestingly, the fractional anisotropy and, to a lesser extent, the mean diffusivity, are best recovered in missing slices by using the autoencoder trained with SH coefficients. A comparison was performed with the same maps reconstructed using an autoencoder trained with raw signals, as well as conventional interpolation methods of raw signals and SH coefficients. From these results, we conclude that the recovery of missing/corrupted slices should be performed in the signal domain if the raw signal is aimed to be recovered, and in the SH domain if diffusion tensor properties (i.e., fractional anisotropy) are targeted. Notably, the trained autoencoders were able to generalize to fetal dMRI data acquired using a much smaller number of diffusion gradients and a lower b-value, where we qualitatively show the consistency of the estimated diffusion tensor maps.
The NLP Sandbox: an efficient model-to-data system to enable federated and unbiased evaluation of clinical NLP models
Jun 28, 2022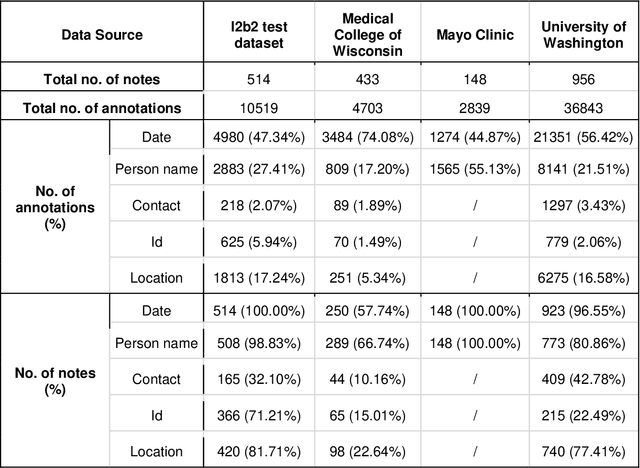
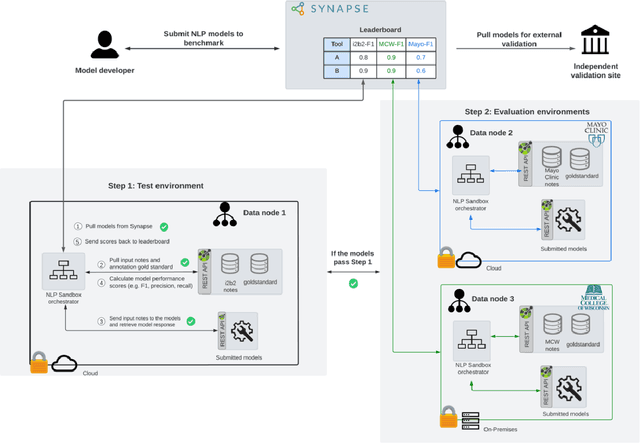
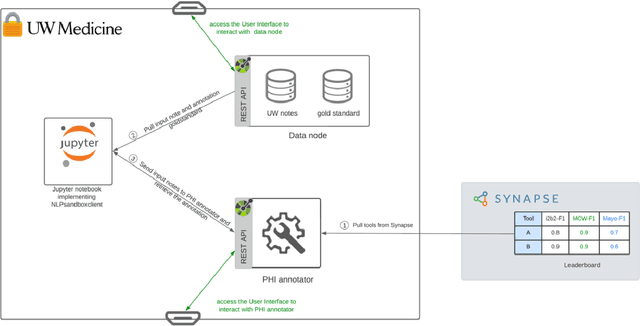
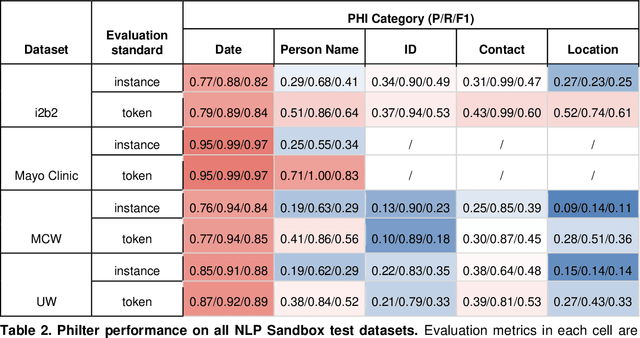
Abstract:Objective The evaluation of natural language processing (NLP) models for clinical text de-identification relies on the availability of clinical notes, which is often restricted due to privacy concerns. The NLP Sandbox is an approach for alleviating the lack of data and evaluation frameworks for NLP models by adopting a federated, model-to-data approach. This enables unbiased federated model evaluation without the need for sharing sensitive data from multiple institutions. Materials and Methods We leveraged the Synapse collaborative framework, containerization software, and OpenAPI generator to build the NLP Sandbox (nlpsandbox.io). We evaluated two state-of-the-art NLP de-identification focused annotation models, Philter and NeuroNER, using data from three institutions. We further validated model performance using data from an external validation site. Results We demonstrated the usefulness of the NLP Sandbox through de-identification clinical model evaluation. The external developer was able to incorporate their model into the NLP Sandbox template and provide user experience feedback. Discussion We demonstrated the feasibility of using the NLP Sandbox to conduct a multi-site evaluation of clinical text de-identification models without the sharing of data. Standardized model and data schemas enable smooth model transfer and implementation. To generalize the NLP Sandbox, work is required on the part of data owners and model developers to develop suitable and standardized schemas and to adapt their data or model to fit the schemas. Conclusions The NLP Sandbox lowers the barrier to utilizing clinical data for NLP model evaluation and facilitates federated, multi-site, unbiased evaluation of NLP models.
Validation and Generalizability of Self-Supervised Image Reconstruction Methods for Undersampled MRI
Jan 29, 2022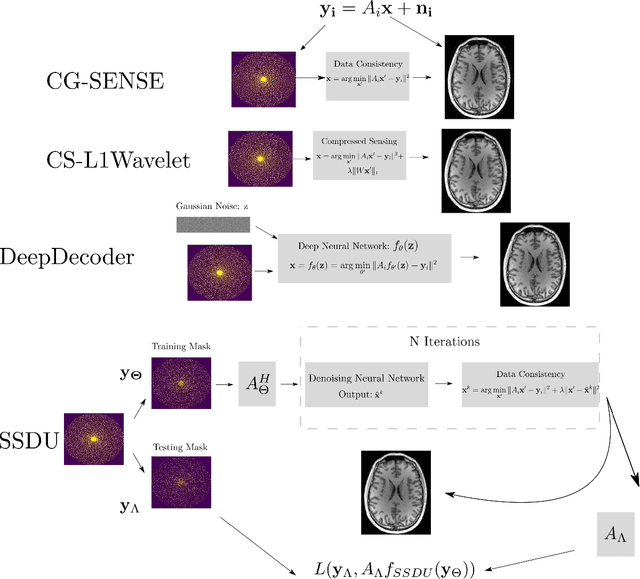
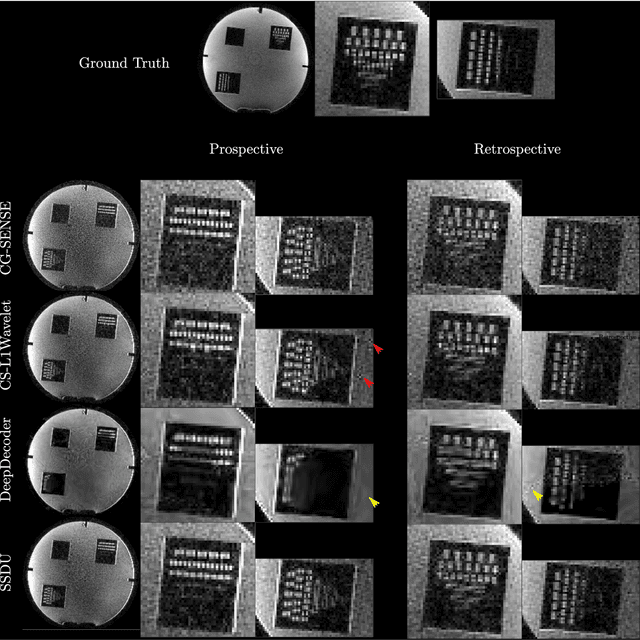
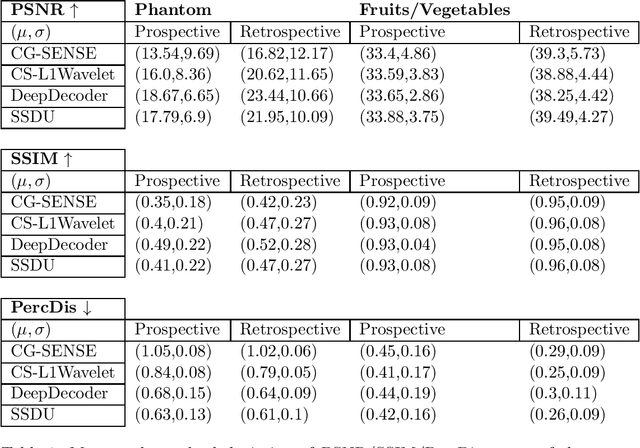

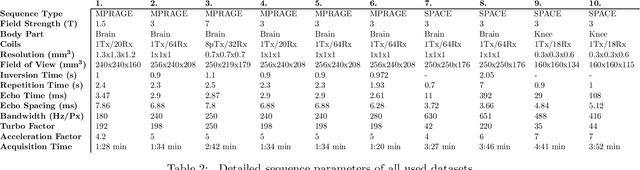
Abstract:Purpose: To investigate aspects of the validation of self-supervised algorithms for reconstruction of undersampled MR images: quantitative evaluation of prospective reconstructions, potential differences between prospective and retrospective reconstructions, suitability of commonly used quantitative metrics, and generalizability. Theory and Methods: Two self-supervised algorithms based on self-supervised denoising and neural network image priors were investigated. These methods are compared to a least squares fitting and a compressed sensing reconstruction using in-vivo and phantom data. Their generalizability was tested with prospectively under-sampled data from experimental conditions different to the training. Results: Prospective reconstructions can exhibit significant distortion relative to retrospective reconstructions/ground truth. Pixel-wise quantitative metrics may not capture differences in perceptual quality accurately, in contrast to a perceptual metric. All methods showed potential for generalization; generalizability is more affected by changes in anatomy/contrast than other changes. No-reference image metrics correspond well with human rating of image quality for studying generalizability. Compressed Sensing and learned denoising perform similarly well on all data. Conclusion: Self-supervised methods show promising results for accelerating image reconstruction in clinical routines. Nonetheless, more work is required to investigate standardized methods to validate reconstruction algorithms for future clinical use.
FaBiAN: A Fetal Brain magnetic resonance Acquisition Numerical phantom
Sep 06, 2021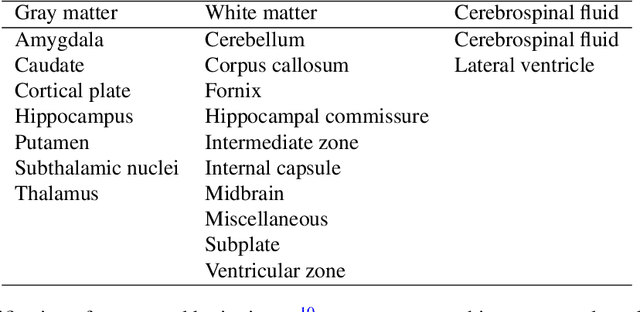
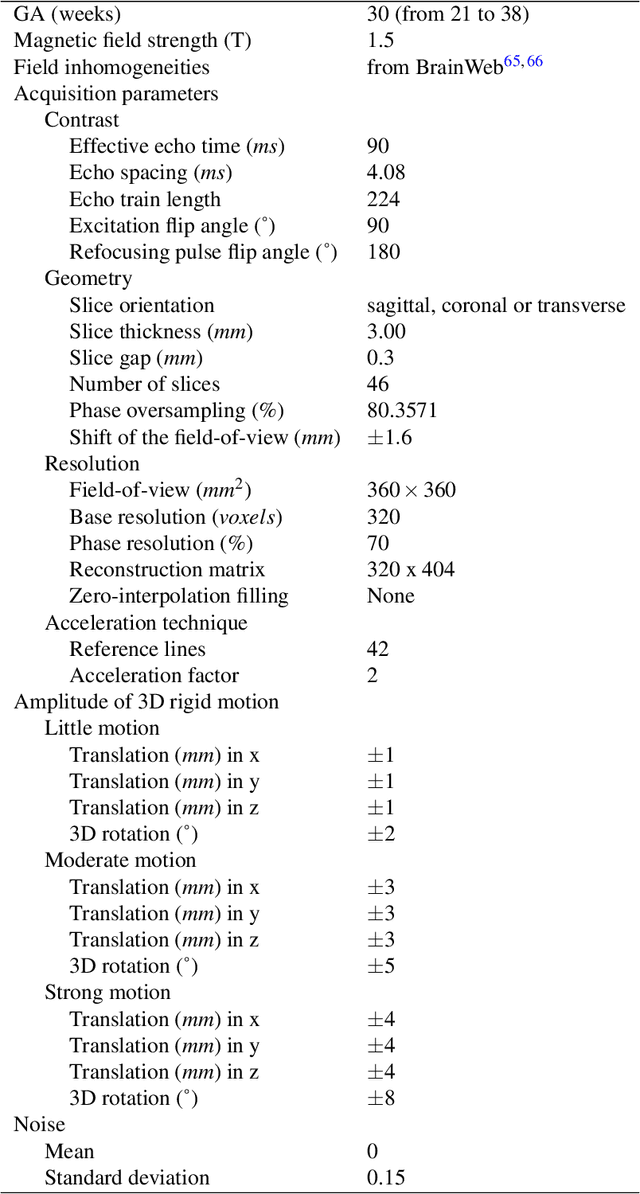
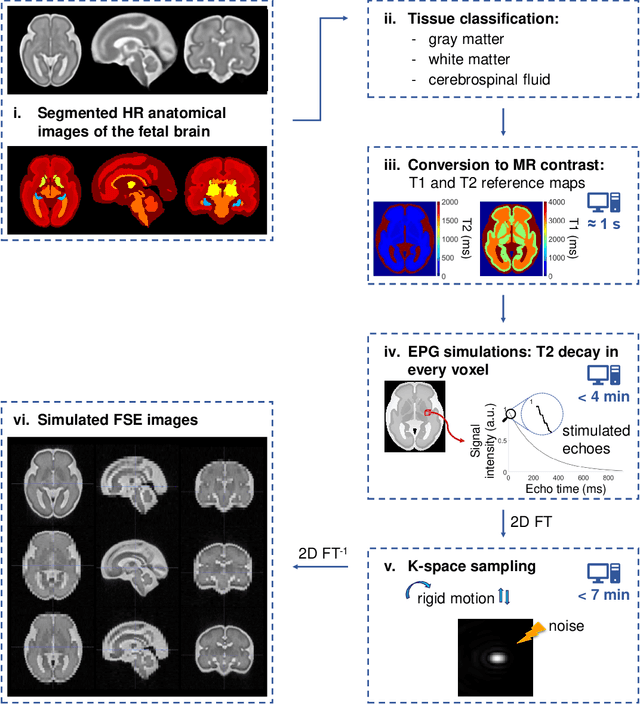

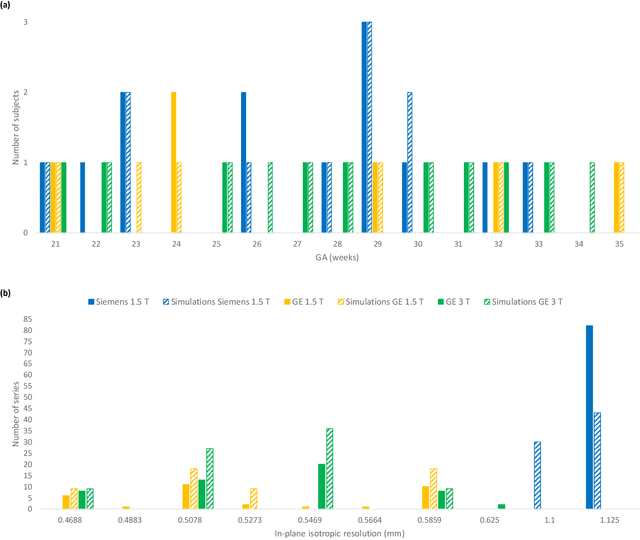
Abstract:Accurate characterization of in utero human brain maturation is critical as it involves complex and interconnected structural and functional processes that may influence health later in life. Magnetic resonance imaging is a powerful tool to investigate equivocal neurological patterns during fetal development. However, the number of acquisitions of satisfactory quality available in this cohort of sensitive subjects remains scarce, thus hindering the validation of advanced image processing techniques. Numerical phantoms can mitigate these limitations by providing a controlled environment with a known ground truth. In this work, we present FaBiAN, an open-source Fetal Brain magnetic resonance Acquisition Numerical phantom that simulates clinical T2-weighted fast spin echo sequences of the fetal brain. This unique tool is based on a general, flexible and realistic setup that includes stochastic fetal movements, thus providing images of the fetal brain throughout maturation comparable to clinical acquisitions. We demonstrate its value to evaluate the robustness and optimize the accuracy of an algorithm for super-resolution fetal brain magnetic resonance imaging from simulated motion-corrupted 2D low-resolution series as compared to a synthetic high-resolution reference volume. We also show that the images generated can complement clinical datasets to support data-intensive deep learning methods for fetal brain tissue segmentation.
The RSNA-ASNR-MICCAI BraTS 2021 Benchmark on Brain Tumor Segmentation and Radiogenomic Classification
Jul 05, 2021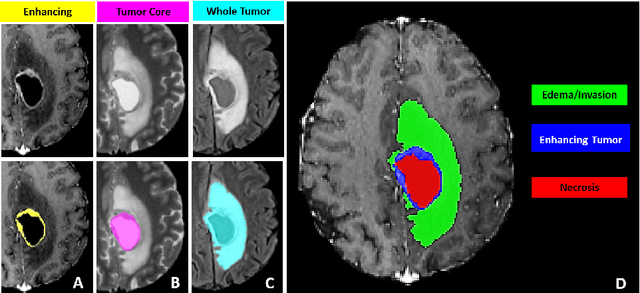
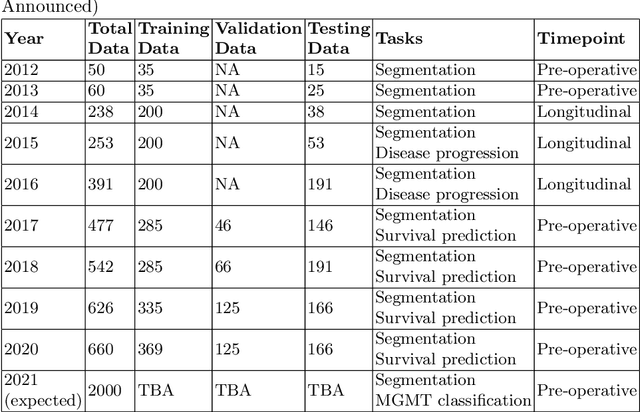
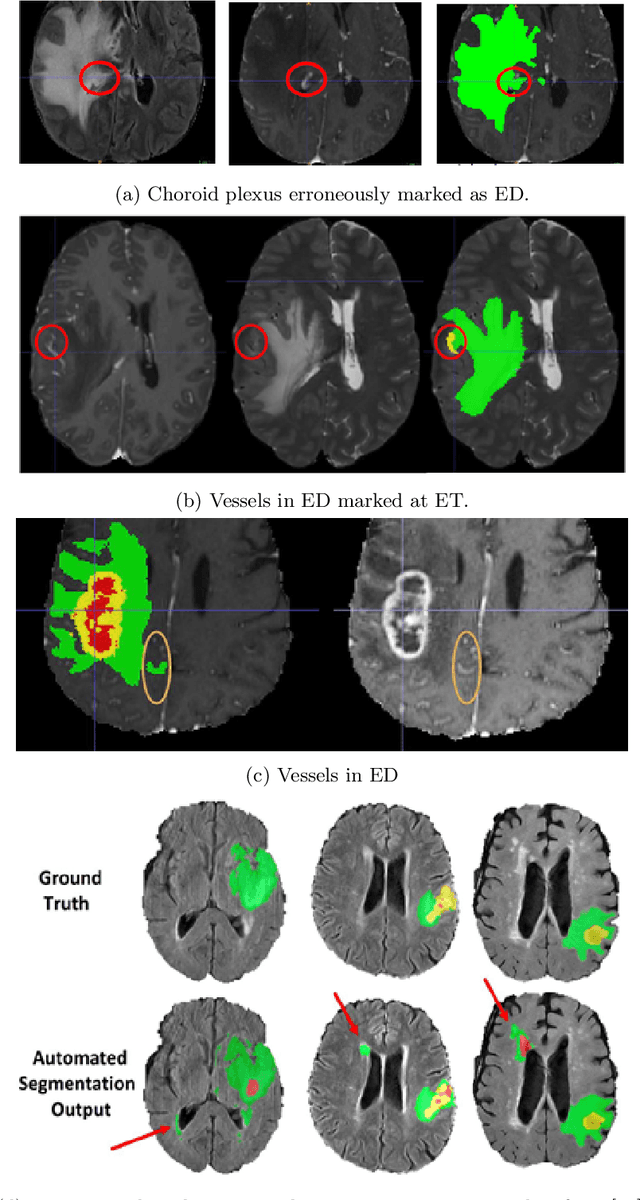
Abstract:The BraTS 2021 challenge celebrates its 10th anniversary and is jointly organized by the Radiological Society of North America (RSNA), the American Society of Neuroradiology (ASNR), and the Medical Image Computing and Computer Assisted Interventions (MICCAI) society. Since its inception, BraTS has been focusing on being a common benchmarking venue for brain glioma segmentation algorithms, with well-curated multi-institutional multi-parametric magnetic resonance imaging (mpMRI) data. Gliomas are the most common primary malignancies of the central nervous system, with varying degrees of aggressiveness and prognosis. The RSNA-ASNR-MICCAI BraTS 2021 challenge targets the evaluation of computational algorithms assessing the same tumor compartmentalization, as well as the underlying tumor's molecular characterization, in pre-operative baseline mpMRI data from 2,000 patients. Specifically, the two tasks that BraTS 2021 focuses on are: a) the segmentation of the histologically distinct brain tumor sub-regions, and b) the classification of the tumor's O[6]-methylguanine-DNA methyltransferase (MGMT) promoter methylation status. The performance evaluation of all participating algorithms in BraTS 2021 will be conducted through the Sage Bionetworks Synapse platform (Task 1) and Kaggle (Task 2), concluding in distributing to the top ranked participants monetary awards of $60,000 collectively.
Test-Time Adaptation for Super-Resolution: You Only Need to Overfit on a Few More Images
Apr 06, 2021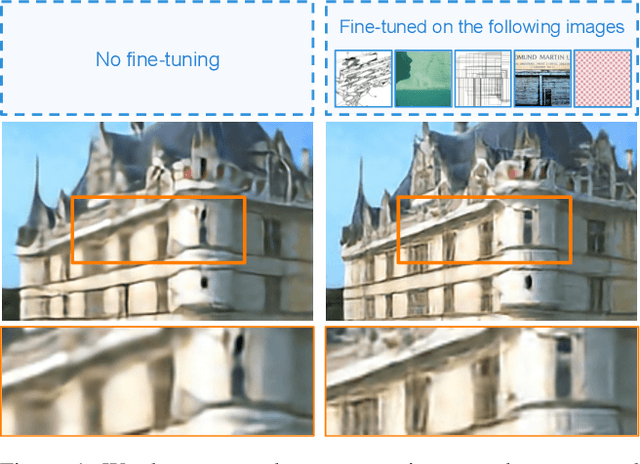
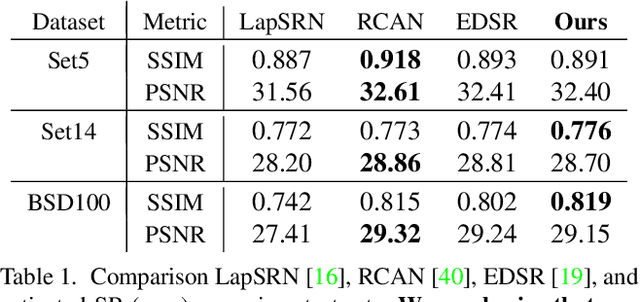
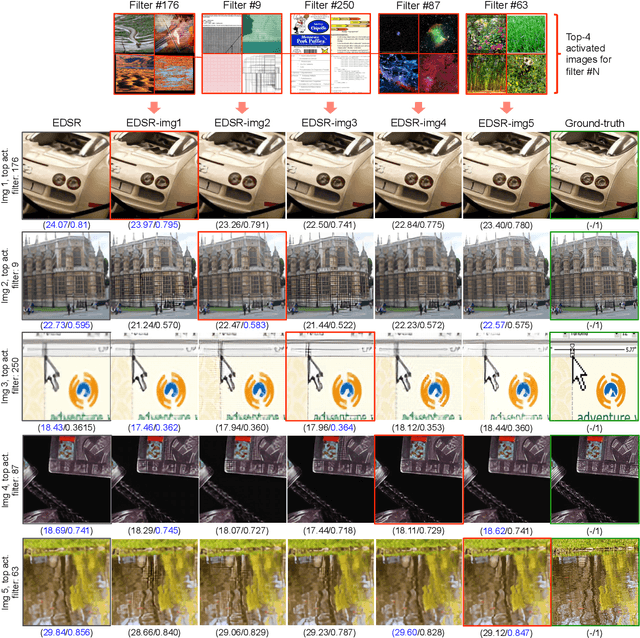
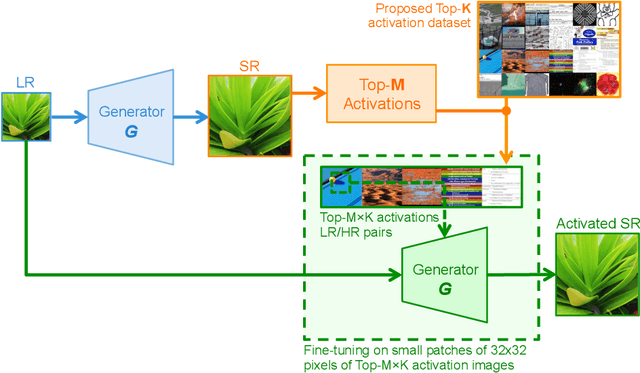
Abstract:Existing reference (RF)-based super-resolution (SR) models try to improve perceptual quality in SR under the assumption of the availability of high-resolution RF images paired with low-resolution (LR) inputs at testing. As the RF images should be similar in terms of content, colors, contrast, etc. to the test image, this hinders the applicability in a real scenario. Other approaches to increase the perceptual quality of images, including perceptual loss and adversarial losses, tend to dramatically decrease fidelity to the ground-truth through significant decreases in PSNR/SSIM. Addressing both issues, we propose a simple yet universal approach to improve the perceptual quality of the HR prediction from a pre-trained SR network on a given LR input by further fine-tuning the SR network on a subset of images from the training dataset with similar patterns of activation as the initial HR prediction, with respect to the filters of a feature extractor. In particular, we show the effects of fine-tuning on these images in terms of the perceptual quality and PSNR/SSIM values. Contrary to perceptually driven approaches, we demonstrate that the fine-tuned network produces a HR prediction with both greater perceptual quality and minimal changes to the PSNR/SSIM with respect to the initial HR prediction. Further, we present novel numerical experiments concerning the filters of SR networks, where we show through filter correlation, that the filters of the fine-tuned network from our method are closer to "ideal" filters, than those of the baseline network or a network fine-tuned on random images.
Benefitting from Bicubically Down-Sampled Images for Learning Real-World Image Super-Resolution
Jul 06, 2020
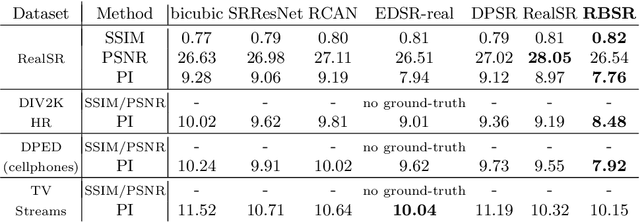
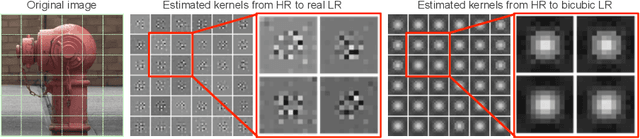
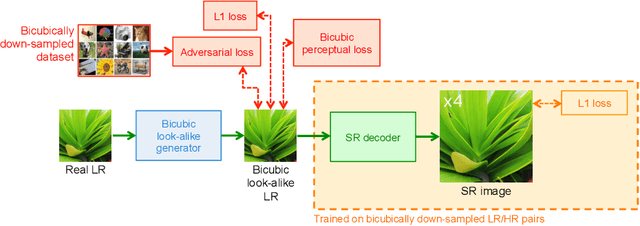
Abstract:Super-resolution (SR) has traditionally been based on pairs of high-resolution images (HR) and their low-resolution (LR) counterparts obtained artificially with bicubic downsampling. However, in real-world SR, there is a large variety of realistic image degradations and analytically modeling these realistic degradations can prove quite difficult. In this work, we propose to handle real-world SR by splitting this ill-posed problem into two comparatively more well-posed steps. First, we train a network to transform real LR images to the space of bicubically downsampled images in a supervised manner, by using both real LR/HR pairs and synthetic pairs. Second, we take a generic SR network trained on bicubically downsampled images to super-resolve the transformed LR image. The first step of the pipeline addresses the problem by registering the large variety of degraded images to a common, well understood space of images. The second step then leverages the already impressive performance of SR on bicubically downsampled images, sidestepping the issues of end-to-end training on datasets with many different image degradations. We demonstrate the effectiveness of our proposed method by comparing it to recent methods in real-world SR and show that our proposed approach outperforms the state-of-the-art works in terms of both qualitative and quantitative results, as well as results of an extensive user study conducted on several real image datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge