Stefan Bauer
Institute for Surgical Technology and Biomechanics, University of Bern, Switzerland
Spatial Dependency Networks: Neural Layers for Improved Generative Image Modeling
Mar 16, 2021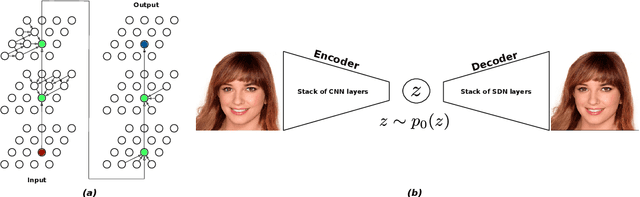
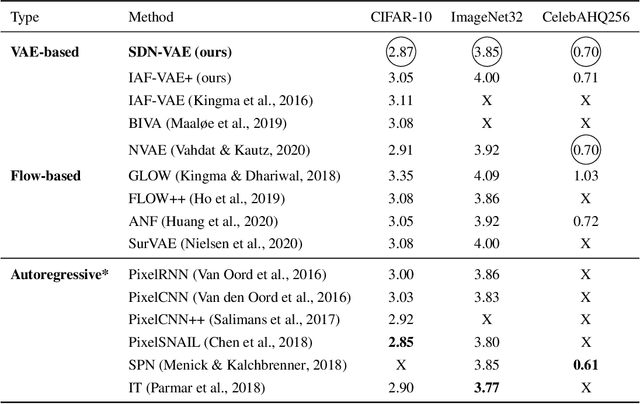
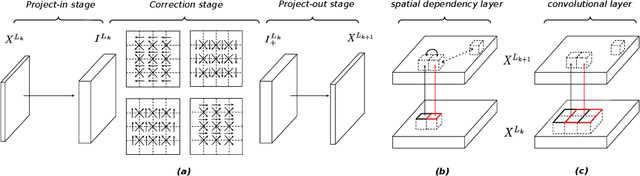
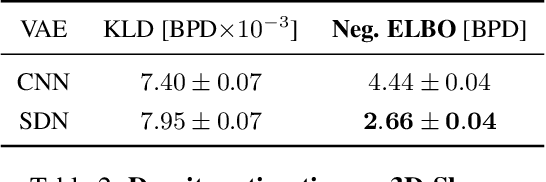
Abstract:How to improve generative modeling by better exploiting spatial regularities and coherence in images? We introduce a novel neural network for building image generators (decoders) and apply it to variational autoencoders (VAEs). In our spatial dependency networks (SDNs), feature maps at each level of a deep neural net are computed in a spatially coherent way, using a sequential gating-based mechanism that distributes contextual information across 2-D space. We show that augmenting the decoder of a hierarchical VAE by spatial dependency layers considerably improves density estimation over baseline convolutional architectures and the state-of-the-art among the models within the same class. Furthermore, we demonstrate that SDN can be applied to large images by synthesizing samples of high quality and coherence. In a vanilla VAE setting, we find that a powerful SDN decoder also improves learning disentangled representations, indicating that neural architectures play an important role in this task. Our results suggest favoring spatial dependency over convolutional layers in various VAE settings. The accompanying source code is given at https://github.com/djordjemila/sdn.
Towards Causal Representation Learning
Feb 22, 2021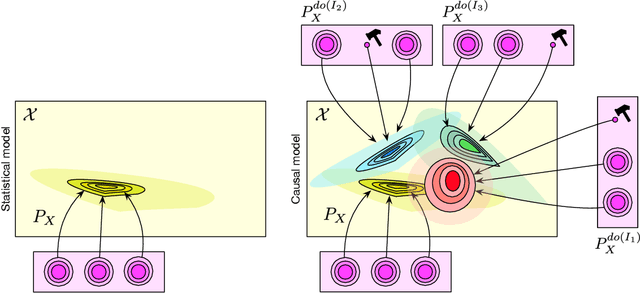
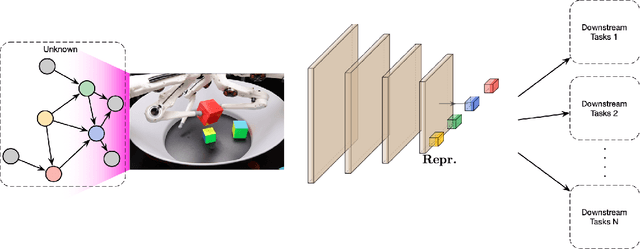
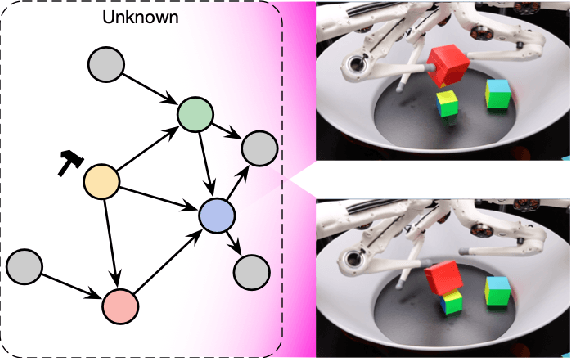
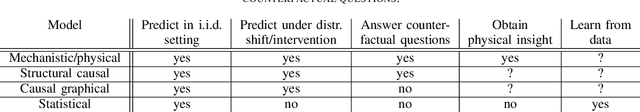
Abstract:The two fields of machine learning and graphical causality arose and developed separately. However, there is now cross-pollination and increasing interest in both fields to benefit from the advances of the other. In the present paper, we review fundamental concepts of causal inference and relate them to crucial open problems of machine learning, including transfer and generalization, thereby assaying how causality can contribute to modern machine learning research. This also applies in the opposite direction: we note that most work in causality starts from the premise that the causal variables are given. A central problem for AI and causality is, thus, causal representation learning, the discovery of high-level causal variables from low-level observations. Finally, we delineate some implications of causality for machine learning and propose key research areas at the intersection of both communities.
Overcoming Barriers to Data Sharing with Medical Image Generation: A Comprehensive Evaluation
Nov 29, 2020
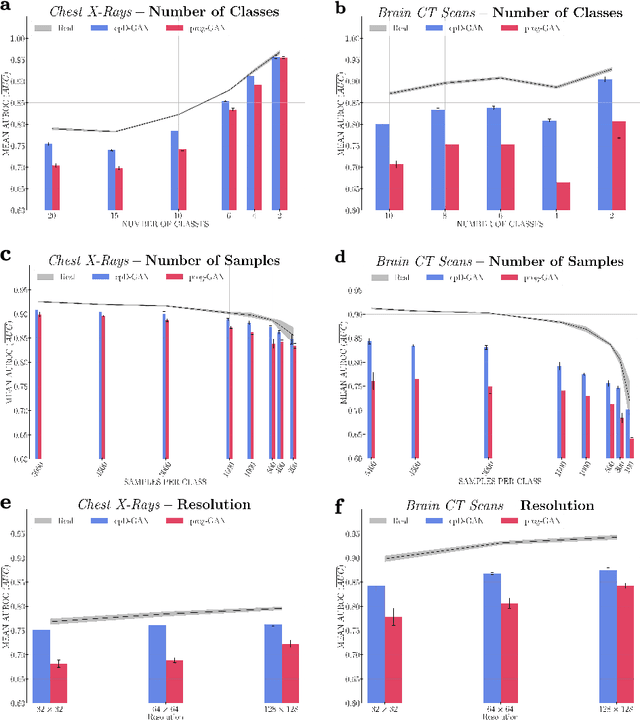
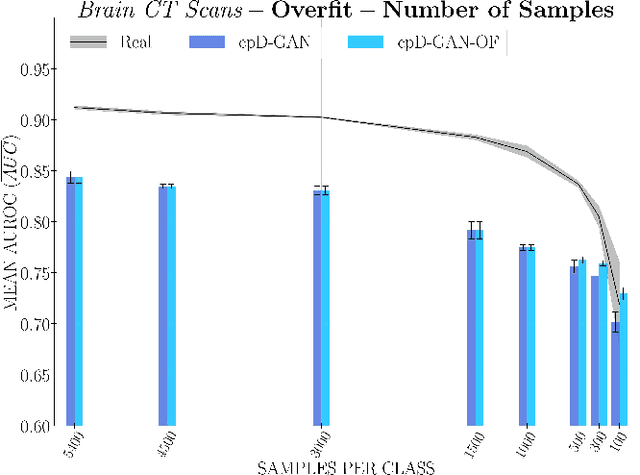
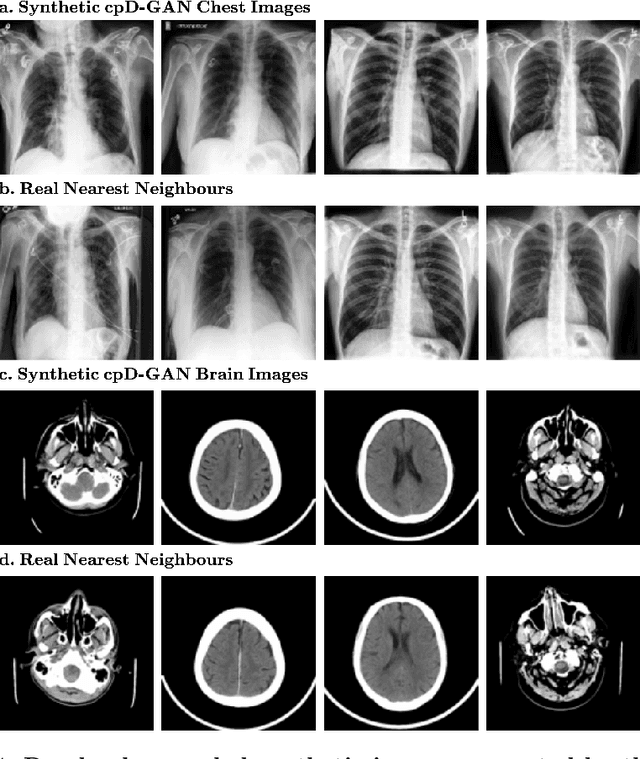
Abstract:Privacy concerns around sharing personally identifiable information are a major practical barrier to data sharing in medical research. However, in many cases, researchers have no interest in a particular individual's information but rather aim to derive insights at the level of cohorts. Here, we utilize Generative Adversarial Networks (GANs) to create derived medical imaging datasets consisting entirely of synthetic patient data. The synthetic images ideally have, in aggregate, similar statistical properties to those of a source dataset but do not contain sensitive personal information. We assess the quality of synthetic data generated by two GAN models for chest radiographs with 14 different radiology findings and brain computed tomography (CT) scans with six types of intracranial hemorrhages. We measure the synthetic image quality by the performance difference of predictive models trained on either the synthetic or the real dataset. We find that synthetic data performance disproportionately benefits from a reduced number of unique label combinations and determine at what number of samples per class overfitting effects start to dominate GAN training. Our open-source benchmark findings also indicate that synthetic data generation can benefit from higher levels of spatial resolution. We additionally conducted a reader study in which trained radiologists do not perform better than random on discriminating between synthetic and real medical images for both data modalities to a statistically significant extent. Our study offers valuable guidelines and outlines practical conditions under which insights derived from synthetic medical images are similar to those that would have been derived from real imaging data. Our results indicate that synthetic data sharing may be an attractive and privacy-preserving alternative to sharing real patient-level data in the right settings.
On the Transfer of Disentangled Representations in Realistic Settings
Oct 27, 2020



Abstract:Learning meaningful representations that disentangle the underlying structure of the data generating process is considered to be of key importance in machine learning. While disentangled representations were found to be useful for diverse tasks such as abstract reasoning and fair classification, their scalability and real-world impact remain questionable. We introduce a new high-resolution dataset with 1M simulated images and over 1,800 annotated real-world images of the same robotic setup. In contrast to previous work, this new dataset exhibits correlations, a complex underlying structure, and allows to evaluate transfer to unseen simulated and real-world settings where the encoder i) remains in distribution or ii) is out of distribution. We propose new architectures in order to scale disentangled representation learning to realistic high-resolution settings and conduct a large-scale empirical study of disentangled representations on this dataset. We observe that disentanglement is a good predictor for out-of-distribution (OOD) task performance.
A Sober Look at the Unsupervised Learning of Disentangled Representations and their Evaluation
Oct 27, 2020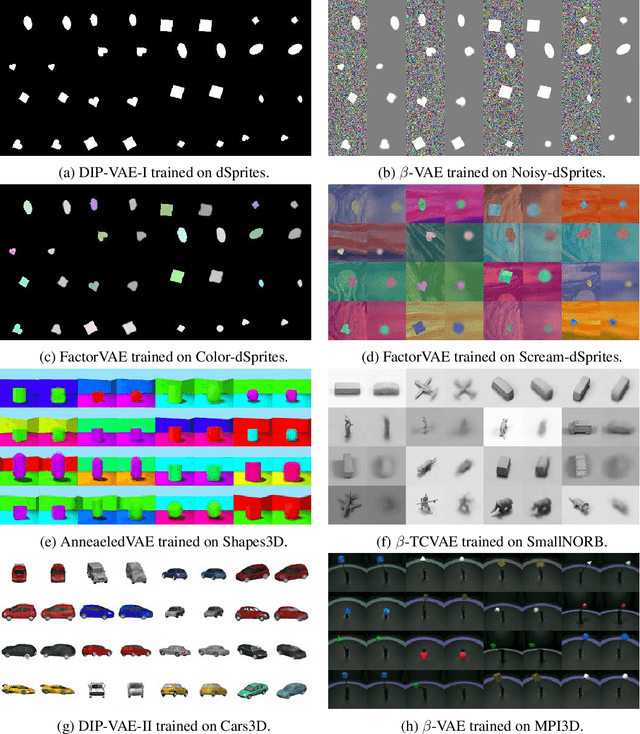
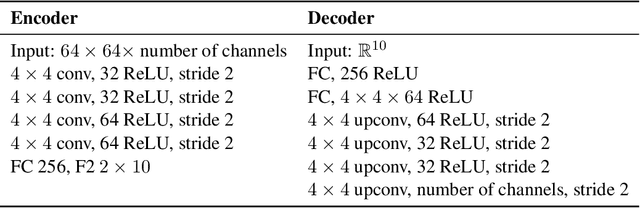

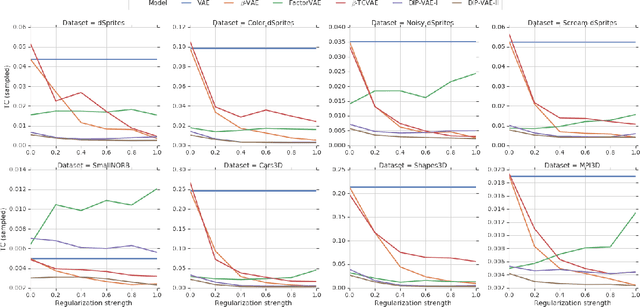

Abstract:The idea behind the \emph{unsupervised} learning of \emph{disentangled} representations is that real-world data is generated by a few explanatory factors of variation which can be recovered by unsupervised learning algorithms. In this paper, we provide a sober look at recent progress in the field and challenge some common assumptions. We first theoretically show that the unsupervised learning of disentangled representations is fundamentally impossible without inductive biases on both the models and the data. Then, we train over $14000$ models covering most prominent methods and evaluation metrics in a reproducible large-scale experimental study on eight data sets. We observe that while the different methods successfully enforce properties "encouraged" by the corresponding losses, well-disentangled models seemingly cannot be identified without supervision. Furthermore, different evaluation metrics do not always agree on what should be considered "disentangled" and exhibit systematic differences in the estimation. Finally, increased disentanglement does not seem to necessarily lead to a decreased sample complexity of learning for downstream tasks. Our results suggest that future work on disentanglement learning should be explicit about the role of inductive biases and (implicit) supervision, investigate concrete benefits of enforcing disentanglement of the learned representations, and consider a reproducible experimental setup covering several data sets.
* arXiv admin note: substantial text overlap with arXiv:1811.12359
Function Contrastive Learning of Transferable Representations
Oct 14, 2020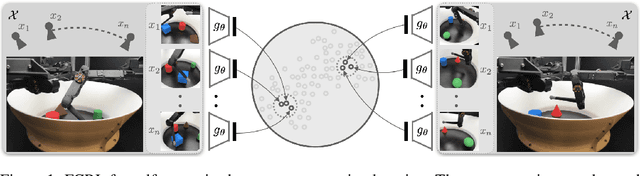
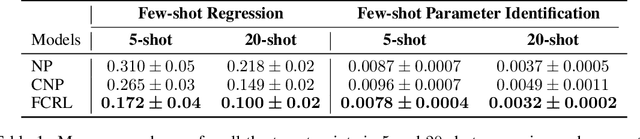
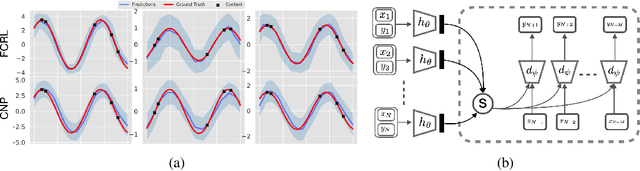
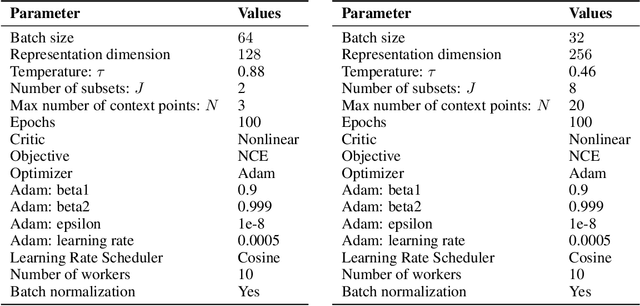
Abstract:Few-shot-learning seeks to find models that are capable of fast-adaptation to novel tasks. Unlike typical few-shot learning algorithms, we propose a contrastive learning method which is not trained to solve a set of tasks, but rather attempts to find a good representation of the underlying data-generating processes (\emph{functions}). This allows for finding representations which are useful for an entire series of tasks sharing the same function. In particular, our training scheme is driven by the self-supervision signal indicating whether two sets of samples stem from the same underlying function. Our experiments on a number of synthetic and real-world datasets show that the representations we obtain can outperform strong baselines in terms of downstream performance and noise robustness, even when these baselines are trained in an end-to-end manner.
CausalWorld: A Robotic Manipulation Benchmark for Causal Structure and Transfer Learning
Oct 08, 2020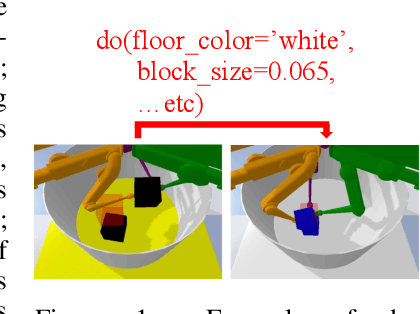
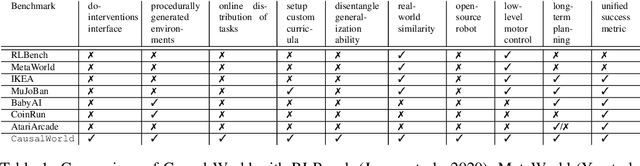
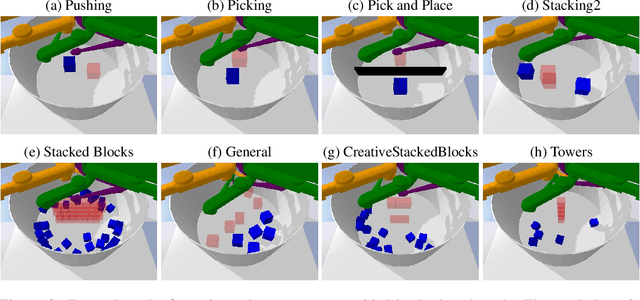
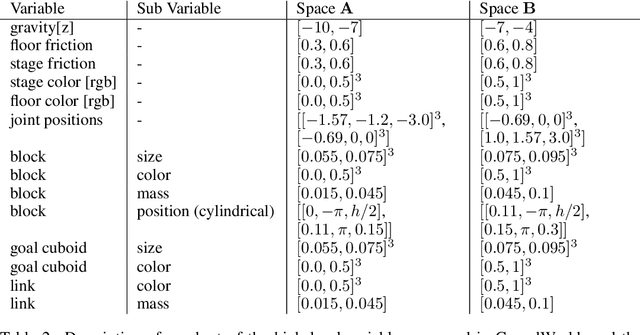
Abstract:Despite recent successes of reinforcement learning (RL), it remains a challenge for agents to transfer learned skills to related environments. To facilitate research addressing this problem, we propose CausalWorld, a benchmark for causal structure and transfer learning in a robotic manipulation environment. The environment is a simulation of an open-source robotic platform, hence offering the possibility of sim-to-real transfer. Tasks consist of constructing 3D shapes from a given set of blocks - inspired by how children learn to build complex structures. The key strength of CausalWorld is that it provides a combinatorial family of such tasks with common causal structure and underlying factors (including, e.g., robot and object masses, colors, sizes). The user (or the agent) may intervene on all causal variables, which allows for fine-grained control over how similar different tasks (or task distributions) are. One can thus easily define training and evaluation distributions of a desired difficulty level, targeting a specific form of generalization (e.g., only changes in appearance or object mass). Further, this common parametrization facilitates defining curricula by interpolating between an initial and a target task. While users may define their own task distributions, we present eight meaningful distributions as concrete benchmarks, ranging from simple to very challenging, all of which require long-horizon planning as well as precise low-level motor control. Finally, we provide baseline results for a subset of these tasks on distinct training curricula and corresponding evaluation protocols, verifying the feasibility of the tasks in this benchmark.
Real-time Prediction of COVID-19 related Mortality using Electronic Health Records
Aug 31, 2020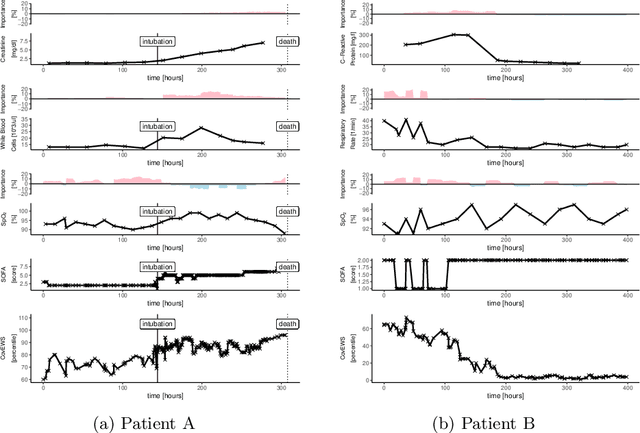
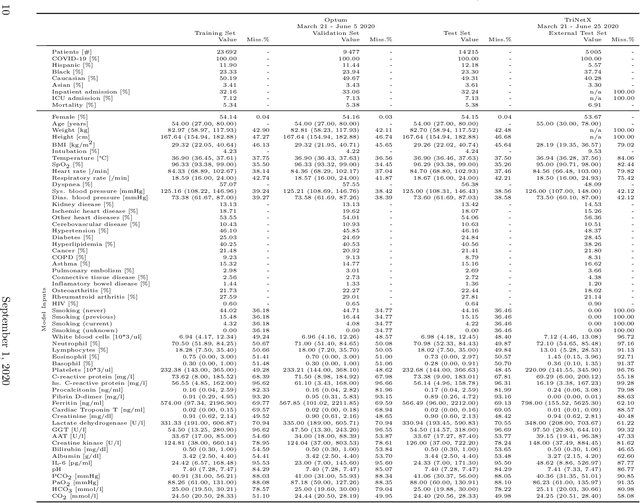
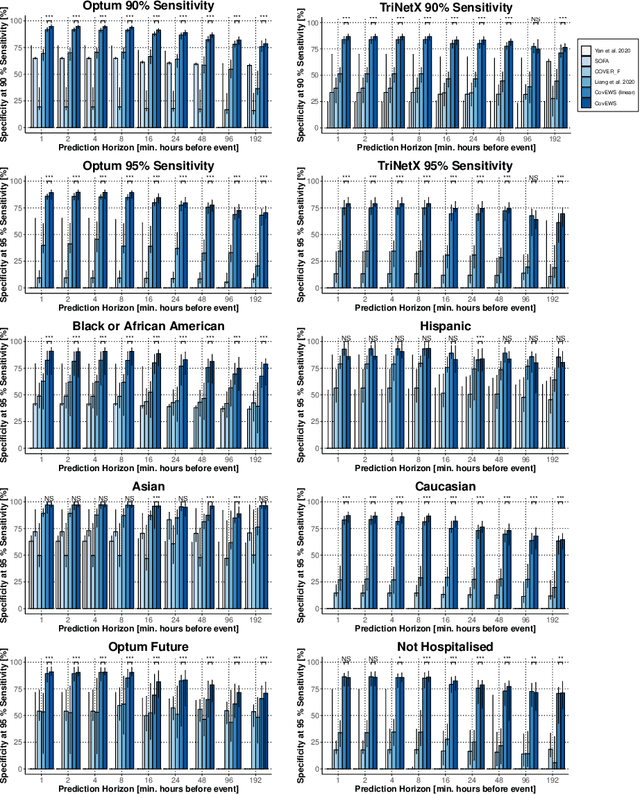
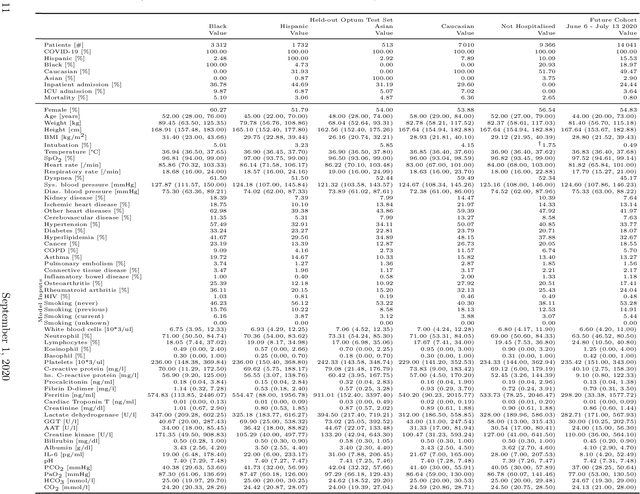
Abstract:Coronavirus Disease 2019 (COVID-19) is an emerging respiratory disease caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) with rapid human-to-human transmission and a high case fatality rate particularly in older patients. Due to the exponential growth of infections, many healthcare systems across the world are under pressure to care for increasing amounts of at-risk patients. Given the high number of infected patients, identifying patients with the highest mortality risk early is critical to enable effective intervention and optimal prioritisation of care. Here, we present the COVID-19 Early Warning System (CovEWS), a clinical risk scoring system for assessing COVID-19 related mortality risk. CovEWS provides continuous real-time risk scores for individual patients with clinically meaningful predictive performance up to 192 hours (8 days) in advance, and is automatically derived from patients' electronic health records (EHRs) using machine learning. We trained and evaluated CovEWS using de-identified data from a cohort of 66430 COVID-19 positive patients seen at over 69 healthcare institutions in the United States (US), Australia, Malaysia and India amounting to an aggregated total of over 2863 years of patient observation time. On an external test cohort of 5005 patients, CovEWS predicts COVID-19 related mortality from $78.8\%$ ($95\%$ confidence interval [CI]: $76.0$, $84.7\%$) to $69.4\%$ ($95\%$ CI: $57.6, 75.2\%$) specificity at a sensitivity greater than $95\%$ between respectively 1 and 192 hours prior to observed mortality events - significantly outperforming existing generic and COVID-19 specific clinical risk scores. CovEWS could enable clinicians to intervene at an earlier stage, and may therefore help in preventing or mitigating COVID-19 related mortality.
TriFinger: An Open-Source Robot for Learning Dexterity
Aug 08, 2020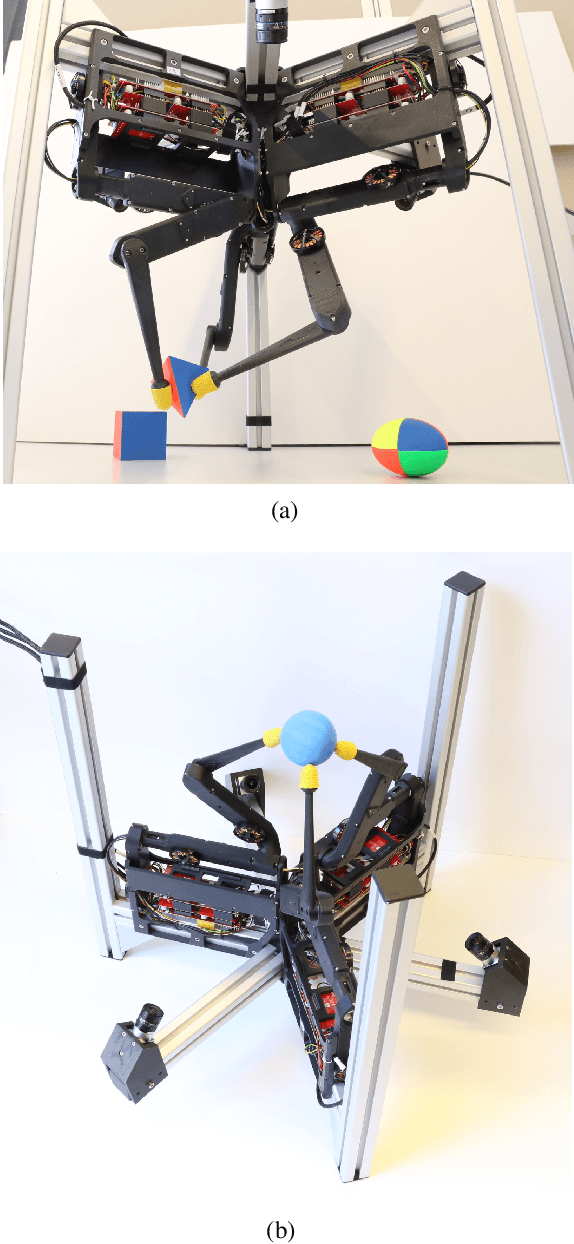
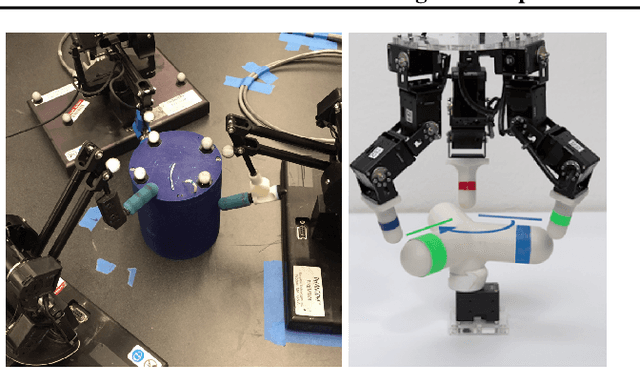
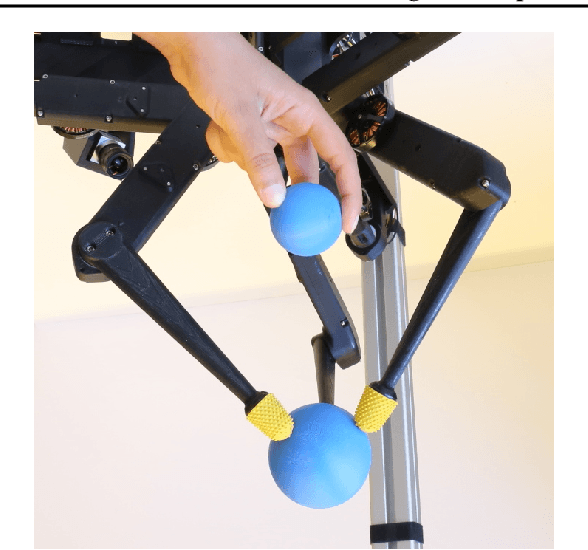
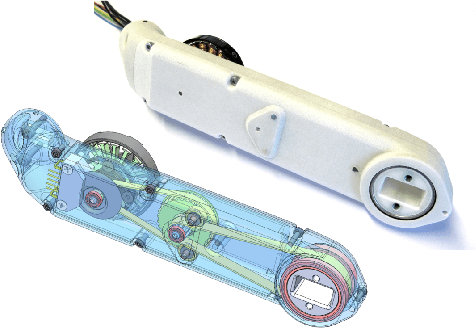



Abstract:Dexterous object manipulation remains an open problem in robotics, despite the rapid progress in machine learning during the past decade. We argue that a hindrance is the high cost of experimentation on real systems, in terms of both time and money. We address this problem by proposing an open-source robotic platform which can safely operate without human supervision. The hardware is inexpensive (about \SI{5000}[\$]{}) yet highly dynamic, robust, and capable of complex interaction with external objects. The software operates at 1-kilohertz and performs safety checks to prevent the hardware from breaking. The easy-to-use front-end (in C++ and Python) is suitable for real-time control as well as deep reinforcement learning. In addition, the software framework is largely robot-agnostic and can hence be used independently of the hardware proposed herein. Finally, we illustrate the potential of the proposed platform through a number of experiments, including real-time optimal control, deep reinforcement learning from scratch, throwing, and writing.
A Commentary on the Unsupervised Learning of Disentangled Representations
Jul 28, 2020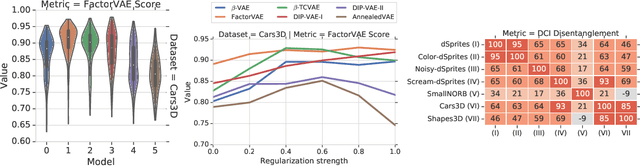
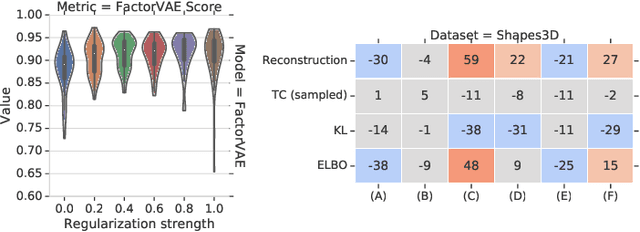
Abstract:The goal of the unsupervised learning of disentangled representations is to separate the independent explanatory factors of variation in the data without access to supervision. In this paper, we summarize the results of Locatello et al., 2019, and focus on their implications for practitioners. We discuss the theoretical result showing that the unsupervised learning of disentangled representations is fundamentally impossible without inductive biases and the practical challenges it entails. Finally, we comment on our experimental findings, highlighting the limitations of state-of-the-art approaches and directions for future research.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge