Shiyu Xia
Towards Understanding Feature Learning in Parameter Transfer
Sep 26, 2025
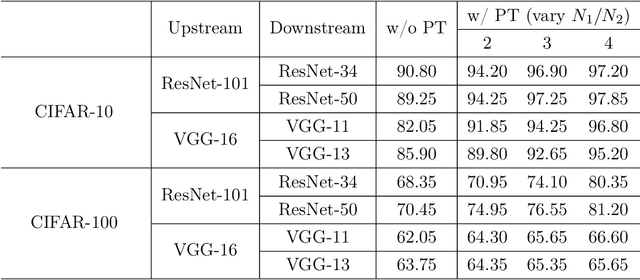
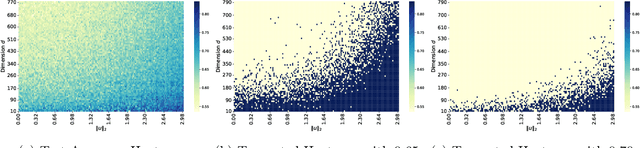
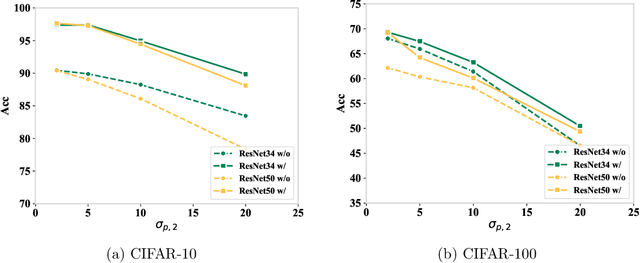
Abstract:Parameter transfer is a central paradigm in transfer learning, enabling knowledge reuse across tasks and domains by sharing model parameters between upstream and downstream models. However, when only a subset of parameters from the upstream model is transferred to the downstream model, there remains a lack of theoretical understanding of the conditions under which such partial parameter reuse is beneficial and of the factors that govern its effectiveness. To address this gap, we analyze a setting in which both the upstream and downstream models are ReLU convolutional neural networks (CNNs). Within this theoretical framework, we characterize how the inherited parameters act as carriers of universal knowledge and identify key factors that amplify their beneficial impact on the target task. Furthermore, our analysis provides insight into why, in certain cases, transferring parameters can lead to lower test accuracy on the target task than training a new model from scratch. Numerical experiments and real-world data experiments are conducted to empirically validate our theoretical findings.
Towards Video to Piano Music Generation with Chain-of-Perform Support Benchmarks
May 26, 2025

Abstract:Generating high-quality piano audio from video requires precise synchronization between visual cues and musical output, ensuring accurate semantic and temporal alignment.However, existing evaluation datasets do not fully capture the intricate synchronization required for piano music generation. A comprehensive benchmark is essential for two primary reasons: (1) existing metrics fail to reflect the complexity of video-to-piano music interactions, and (2) a dedicated benchmark dataset can provide valuable insights to accelerate progress in high-quality piano music generation. To address these challenges, we introduce the CoP Benchmark Dataset-a fully open-sourced, multimodal benchmark designed specifically for video-guided piano music generation. The proposed Chain-of-Perform (CoP) benchmark offers several compelling features: (1) detailed multimodal annotations, enabling precise semantic and temporal alignment between video content and piano audio via step-by-step Chain-of-Perform guidance; (2) a versatile evaluation framework for rigorous assessment of both general-purpose and specialized video-to-piano generation tasks; and (3) full open-sourcing of the dataset, annotations, and evaluation protocols. The dataset is publicly available at https://github.com/acappemin/Video-to-Audio-and-Piano, with a continuously updated leaderboard to promote ongoing research in this domain.
Towards Film-Making Production Dialogue, Narration, Monologue Adaptive Moving Dubbing Benchmarks
Apr 30, 2025Abstract:Movie dubbing has advanced significantly, yet assessing the real-world effectiveness of these models remains challenging. A comprehensive evaluation benchmark is crucial for two key reasons: 1) Existing metrics fail to fully capture the complexities of dialogue, narration, monologue, and actor adaptability in movie dubbing. 2) A practical evaluation system should offer valuable insights to improve movie dubbing quality and advancement in film production. To this end, we introduce Talking Adaptive Dubbing Benchmarks (TA-Dubbing), designed to improve film production by adapting to dialogue, narration, monologue, and actors in movie dubbing. TA-Dubbing offers several key advantages: 1) Comprehensive Dimensions: TA-Dubbing covers a variety of dimensions of movie dubbing, incorporating metric evaluations for both movie understanding and speech generation. 2) Versatile Benchmarking: TA-Dubbing is designed to evaluate state-of-the-art movie dubbing models and advanced multi-modal large language models. 3) Full Open-Sourcing: We fully open-source TA-Dubbing at https://github.com/woka- 0a/DeepDubber- V1 including all video suits, evaluation methods, annotations. We also continuously integrate new movie dubbing models into the TA-Dubbing leaderboard at https://github.com/woka- 0a/DeepDubber-V1 to drive forward the field of movie dubbing.
SpreadsheetLLM: Encoding Spreadsheets for Large Language Models
Jul 12, 2024



Abstract:Spreadsheets, with their extensive two-dimensional grids, various layouts, and diverse formatting options, present notable challenges for large language models (LLMs). In response, we introduce SpreadsheetLLM, pioneering an efficient encoding method designed to unleash and optimize LLMs' powerful understanding and reasoning capability on spreadsheets. Initially, we propose a vanilla serialization approach that incorporates cell addresses, values, and formats. However, this approach was limited by LLMs' token constraints, making it impractical for most applications. To tackle this challenge, we develop SheetCompressor, an innovative encoding framework that compresses spreadsheets effectively for LLMs. It comprises three modules: structural-anchor-based compression, inverse index translation, and data-format-aware aggregation. It significantly improves performance in spreadsheet table detection task, outperforming the vanilla approach by 25.6% in GPT4's in-context learning setting. Moreover, fine-tuned LLM with SheetCompressor has an average compression ratio of 25 times, but achieves a state-of-the-art 78.9% F1 score, surpassing the best existing models by 12.3%. Finally, we propose Chain of Spreadsheet for downstream tasks of spreadsheet understanding and validate in a new and demanding spreadsheet QA task. We methodically leverage the inherent layout and structure of spreadsheets, demonstrating that SpreadsheetLLM is highly effective across a variety of spreadsheet tasks.
Vision Language Models for Spreadsheet Understanding: Challenges and Opportunities
May 25, 2024



Abstract:This paper explores capabilities of Vision Language Models on spreadsheet comprehension. We propose three self-supervised challenges with corresponding evaluation metrics to comprehensively evaluate VLMs on Optical Character Recognition (OCR), spatial perception, and visual format recognition. Additionally, we utilize the spreadsheet table detection task to assess the overall performance of VLMs by integrating these challenges. To probe VLMs more finely, we propose three spreadsheet-to-image settings: column width adjustment, style change, and address augmentation. We propose variants of prompts to address the above tasks in different settings. Notably, to leverage the strengths of VLMs in understanding text rather than two-dimensional positioning, we propose to decode cell values on the four boundaries of the table in spreadsheet boundary detection. Our findings reveal that VLMs demonstrate promising OCR capabilities but produce unsatisfactory results due to cell omission and misalignment, and they notably exhibit insufficient spatial and format recognition skills, motivating future work to enhance VLMs' spreadsheet data comprehension capabilities using our methods to generate extensive spreadsheet-image pairs in various settings.
Transformer as Linear Expansion of Learngene
Dec 20, 2023



Abstract:We propose expanding the shared Transformer module to produce and initialize Transformers of varying depths, enabling adaptation to diverse resource constraints. Drawing an analogy to genetic expansibility, we term such module as learngene. To identify the expansion mechanism, we delve into the relationship between the layer's position and its corresponding weight value, and find that linear function appropriately approximates this relationship. Building on this insight, we present Transformer as Linear Expansion of learnGene (TLEG), a novel approach for flexibly producing and initializing Transformers of diverse depths. Specifically, to learn learngene, we firstly construct an auxiliary Transformer linearly expanded from learngene, after which we train it through employing soft distillation. Subsequently, we can produce and initialize Transformers of varying depths via linearly expanding the well-trained learngene, thereby supporting diverse downstream scenarios. Extensive experiments on ImageNet-1K demonstrate that TLEG achieves comparable or better performance in contrast to many individual models trained from scratch, while reducing around 2x training cost. When transferring to several downstream classification datasets, TLEG surpasses existing initialization methods by a large margin (e.g., +6.87% on iNat 2019 and +7.66% on CIFAR-100). Under the situation where we need to produce models of varying depths adapting for different resource constraints, TLEG achieves comparable results while reducing around 19x parameters stored to initialize these models and around 5x pre-training costs, in contrast to the pre-training and fine-tuning approach. When transferring a fixed set of parameters to initialize different models, TLEG presents better flexibility and competitive performance while reducing around 2.9x parameters stored to initialize, compared to the pre-training approach.
Building Variable-sized Models via Learngene Pool
Dec 12, 2023



Abstract:Recently, Stitchable Neural Networks (SN-Net) is proposed to stitch some pre-trained networks for quickly building numerous networks with different complexity and performance trade-offs. In this way, the burdens of designing or training the variable-sized networks, which can be used in application scenarios with diverse resource constraints, are alleviated. However, SN-Net still faces a few challenges. 1) Stitching from multiple independently pre-trained anchors introduces high storage resource consumption. 2) SN-Net faces challenges to build smaller models for low resource constraints. 3). SN-Net uses an unlearned initialization method for stitch layers, limiting the final performance. To overcome these challenges, motivated by the recently proposed Learngene framework, we propose a novel method called Learngene Pool. Briefly, Learngene distills the critical knowledge from a large pre-trained model into a small part (termed as learngene) and then expands this small part into a few variable-sized models. In our proposed method, we distill one pretrained large model into multiple small models whose network blocks are used as learngene instances to construct the learngene pool. Since only one large model is used, we do not need to store more large models as SN-Net and after distilling, smaller learngene instances can be created to build small models to satisfy low resource constraints. We also insert learnable transformation matrices between the instances to stitch them into variable-sized models to improve the performance of these models. Exhaustive experiments have been implemented and the results validate the effectiveness of the proposed Learngene Pool compared with SN-Net.
Towards Effective Visual Representations for Partial-Label Learning
May 10, 2023



Abstract:Under partial-label learning (PLL) where, for each training instance, only a set of ambiguous candidate labels containing the unknown true label is accessible, contrastive learning has recently boosted the performance of PLL on vision tasks, attributed to representations learned by contrasting the same/different classes of entities. Without access to true labels, positive points are predicted using pseudo-labels that are inherently noisy, and negative points often require large batches or momentum encoders, resulting in unreliable similarity information and a high computational overhead. In this paper, we rethink a state-of-the-art contrastive PLL method PiCO[24], inspiring the design of a simple framework termed PaPi (Partial-label learning with a guided Prototypical classifier), which demonstrates significant scope for improvement in representation learning, thus contributing to label disambiguation. PaPi guides the optimization of a prototypical classifier by a linear classifier with which they share the same feature encoder, thus explicitly encouraging the representation to reflect visual similarity between categories. It is also technically appealing, as PaPi requires only a few components in PiCO with the opposite direction of guidance, and directly eliminates the contrastive learning module that would introduce noise and consume computational resources. We empirically demonstrate that PaPi significantly outperforms other PLL methods on various image classification tasks.
Learngene: From Open-World to Your Learning Task
Jun 12, 2021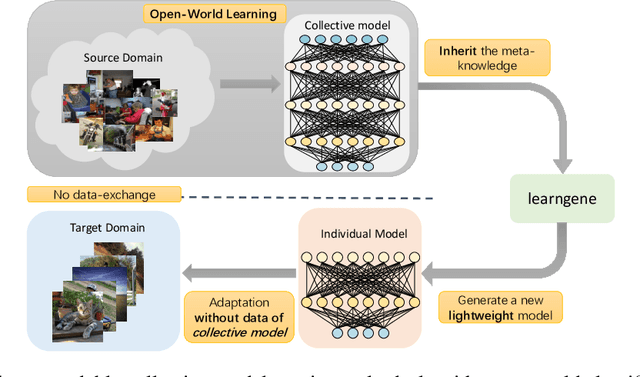
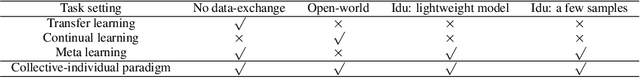
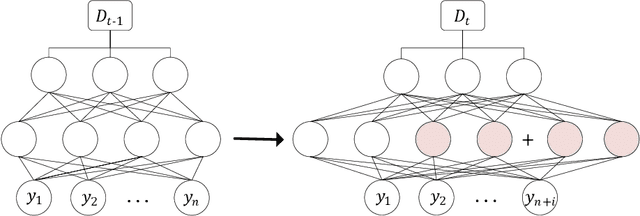
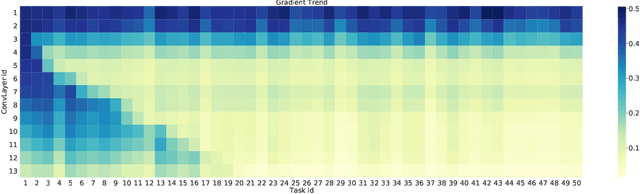
Abstract:Although deep learning has made significant progress on fixed large-scale datasets, it typically encounters challenges regarding improperly detecting new/unseen classes in the open-world classification, over-parametrized, and overfitting small samples. In contrast, biological systems can overcome the above difficulties very well. Individuals inherit an innate gene from collective creatures that have evolved over hundreds of millions of years, and can learn new skills through a few examples. Inspired by this, we propose a practical collective-individual paradigm where open-world tasks are trained in sequence using an evolution (expandable) network. To be specific, we innovatively introduce learngene that inherits the meta-knowledge from the collective model and reconstructs a new lightweight individual model for the target task, to realize the collective-individual paradigm. Particularly, we present a novel criterion that can discover the learngene in the collective model, according to the gradient information. Finally, the individual model is trained only with a few samples in the absence of the source data. We demonstrate the effectiveness of our approach in an extensive empirical study and theoretical analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge