S. Kevin Zhou
Mixed-order self-paced curriculum learning for universal lesion detection
Feb 09, 2023



Abstract:Self-paced curriculum learning (SCL) has demonstrated its great potential in computer vision, natural language processing, etc. During training, it implements easy-to-hard sampling based on online estimation of data difficulty. Most SCL methods commonly adopt a loss-based strategy of estimating data difficulty and deweighting the `hard' samples in the early training stage. While achieving success in a variety of applications, SCL stills confront two challenges in a medical image analysis task, such as universal lesion detection, featuring insufficient and highly class-imbalanced data: (i) the loss-based difficulty measurer is inaccurate; ii) the hard samples are under-utilized from a deweighting mechanism. To overcome these challenges, in this paper we propose a novel mixed-order self-paced curriculum learning (Mo-SCL) method. We integrate both uncertainty and loss to better estimate difficulty online and mix both hard and easy samples in the same mini-batch to appropriately alleviate the problem of under-utilization of hard samples. We provide a theoretical investigation of our method in the context of stochastic gradient descent optimization and extensive experiments based on the DeepLesion benchmark dataset for universal lesion detection (ULD). When applied to two state-of-the-art ULD methods, the proposed mixed-order SCL method can provide a free boost to lesion detection accuracy without extra special network designs.
Multi-site Organ Segmentation with Federated Partial Supervision and Site Adaptation
Feb 08, 2023



Abstract:Objective and Impact Statement: Accurate organ segmentation is critical for many clinical applications at different clinical sites, which may have their specific application requirements that concern different organs. Introduction: However, learning high-quality, site-specific organ segmentation models is challenging as it often needs on-site curation of a large number of annotated images. Security concerns further complicate the matter. Methods: The paper aims to tackle these challenges via a two-phase aggregation-then-adaptation approach. The first phase of federated aggregation learns a single multi-organ segmentation model by leveraging the strength of 'bigger data', which are formed by (i) aggregating together datasets from multiple sites that with different organ labels to provide partial supervision, and (ii) conducting partially supervised learning without data breach. The second phase of site adaptation is to transfer the federated multi-organ segmentation model to site-specific organ segmentation models, one model per site, in order to further improve the performance of each site's organ segmentation task. Furthermore, improved marginal loss and exclusion loss functions are used to avoid 'knowledge conflict' problem in a partially supervision mechanism. Results and Conclusion: Extensive experiments on five organ segmentation datasets demonstrate the effectiveness of our multi-site approach, significantly outperforming the site-per-se learned models and achieving the performance comparable to the centrally learned models.
MURPHY: Relations Matter in Surgical Workflow Analysis
Dec 24, 2022Abstract:Autonomous robotic surgery has advanced significantly based on analysis of visual and temporal cues in surgical workflow, but relational cues from domain knowledge remain under investigation. Complex relations in surgical annotations can be divided into intra- and inter-relations, both valuable to autonomous systems to comprehend surgical workflows. Intra- and inter-relations describe the relevance of various categories within a particular annotation type and the relevance of different annotation types, respectively. This paper aims to systematically investigate the importance of relational cues in surgery. First, we contribute the RLLS12M dataset, a large-scale collection of robotic left lateral sectionectomy (RLLS), by curating 50 videos of 50 patients operated by 5 surgeons and annotating a hierarchical workflow, which consists of 3 inter- and 6 intra-relations, 6 steps, 15 tasks, and 38 activities represented as the triplet of 11 instruments, 8 actions, and 16 objects, totaling 2,113,510 video frames and 12,681,060 annotation entities. Correspondingly, we propose a multi-relation purification hybrid network (MURPHY), which aptly incorporates novel relation modules to augment the feature representation by purifying relational features using the intra- and inter-relations embodied in annotations. The intra-relation module leverages a R-GCN to implant visual features in different graph relations, which are aggregated using a targeted relation purification with affinity information measuring label consistency and feature similarity. The inter-relation module is motivated by attention mechanisms to regularize the influence of relational features based on the hierarchy of annotation types from the domain knowledge. Extensive experimental results on the curated RLLS dataset confirm the effectiveness of our approach, demonstrating that relations matter in surgical workflow analysis.
LE-UDA: Label-efficient unsupervised domain adaptation for medical image segmentation
Dec 05, 2022Abstract:While deep learning methods hitherto have achieved considerable success in medical image segmentation, they are still hampered by two limitations: (i) reliance on large-scale well-labeled datasets, which are difficult to curate due to the expert-driven and time-consuming nature of pixel-level annotations in clinical practices, and (ii) failure to generalize from one domain to another, especially when the target domain is a different modality with severe domain shifts. Recent unsupervised domain adaptation~(UDA) techniques leverage abundant labeled source data together with unlabeled target data to reduce the domain gap, but these methods degrade significantly with limited source annotations. In this study, we address this underexplored UDA problem, investigating a challenging but valuable realistic scenario, where the source domain not only exhibits domain shift~w.r.t. the target domain but also suffers from label scarcity. In this regard, we propose a novel and generic framework called ``Label-Efficient Unsupervised Domain Adaptation"~(LE-UDA). In LE-UDA, we construct self-ensembling consistency for knowledge transfer between both domains, as well as a self-ensembling adversarial learning module to achieve better feature alignment for UDA. To assess the effectiveness of our method, we conduct extensive experiments on two different tasks for cross-modality segmentation between MRI and CT images. Experimental results demonstrate that the proposed LE-UDA can efficiently leverage limited source labels to improve cross-domain segmentation performance, outperforming state-of-the-art UDA approaches in the literature. Code is available at: https://github.com/jacobzhaoziyuan/LE-UDA.
MONAI: An open-source framework for deep learning in healthcare
Nov 04, 2022



Abstract:Artificial Intelligence (AI) is having a tremendous impact across most areas of science. Applications of AI in healthcare have the potential to improve our ability to detect, diagnose, prognose, and intervene on human disease. For AI models to be used clinically, they need to be made safe, reproducible and robust, and the underlying software framework must be aware of the particularities (e.g. geometry, physiology, physics) of medical data being processed. This work introduces MONAI, a freely available, community-supported, and consortium-led PyTorch-based framework for deep learning in healthcare. MONAI extends PyTorch to support medical data, with a particular focus on imaging, and provide purpose-specific AI model architectures, transformations and utilities that streamline the development and deployment of medical AI models. MONAI follows best practices for software-development, providing an easy-to-use, robust, well-documented, and well-tested software framework. MONAI preserves the simple, additive, and compositional approach of its underlying PyTorch libraries. MONAI is being used by and receiving contributions from research, clinical and industrial teams from around the world, who are pursuing applications spanning nearly every aspect of healthcare.
Active CT Reconstruction with a Learned Sampling Policy
Nov 03, 2022



Abstract:Computed tomography (CT) is a widely-used imaging technology that assists clinical decision-making with high-quality human body representations. To reduce the radiation dose posed by CT, sparse-view and limited-angle CT are developed with preserved image quality. However, these methods are still stuck with a fixed or uniform sampling strategy, which inhibits the possibility of acquiring a better image with an even reduced dose. In this paper, we explore this possibility via learning an active sampling policy that optimizes the sampling positions for patient-specific, high-quality reconstruction. To this end, we design an \textit{intelligent agent} for active recommendation of sampling positions based on on-the-fly reconstruction with obtained sinograms in a progressive fashion. With such a design, we achieve better performances on the NIH-AAPM dataset over popular uniform sampling, especially when the number of views is small. Finally, such a design also enables RoI-aware reconstruction with improved reconstruction quality within regions of interest (RoI's) that are clinically important. Experiments on the VerSe dataset demonstrate this ability of our sampling policy, which is difficult to achieve based on uniform sampling.
DA-VSR: Domain Adaptable Volumetric Super-Resolution For Medical Images
Oct 11, 2022Abstract:Medical image super-resolution (SR) is an active research area that has many potential applications, including reducing scan time, bettering visual understanding, increasing robustness in downstream tasks, etc. However, applying deep-learning-based SR approaches for clinical applications often encounters issues of domain inconsistency, as the test data may be acquired by different machines or on different organs. In this work, we present a novel algorithm called domain adaptable volumetric super-resolution (DA-VSR) to better bridge the domain inconsistency gap. DA-VSR uses a unified feature extraction backbone and a series of network heads to improve image quality over different planes. Furthermore, DA-VSR leverages the in-plane and through-plane resolution differences on the test data to achieve a self-learned domain adaptation. As such, DA-VSR combines the advantages of a strong feature generator learned through supervised training and the ability to tune to the idiosyncrasies of the test volumes through unsupervised learning. Through experiments, we demonstrate that DA-VSR significantly improves super-resolution quality across numerous datasets of different domains, thereby taking a further step toward real clinical applications.
A Survey of Fairness in Medical Image Analysis: Concepts, Algorithms, Evaluations, and Challenges
Sep 27, 2022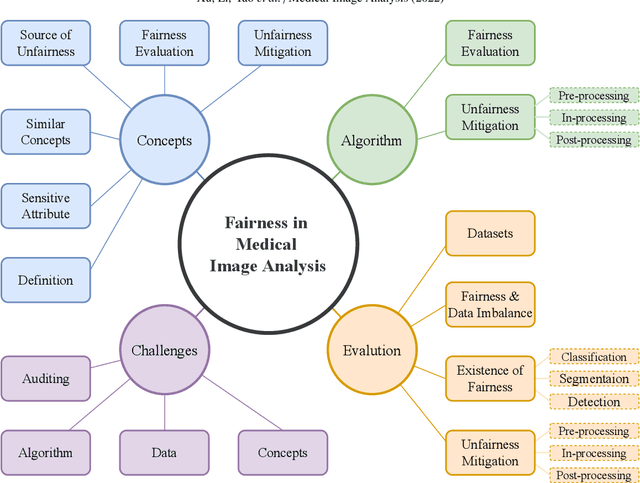
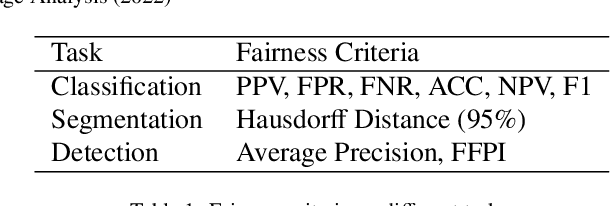

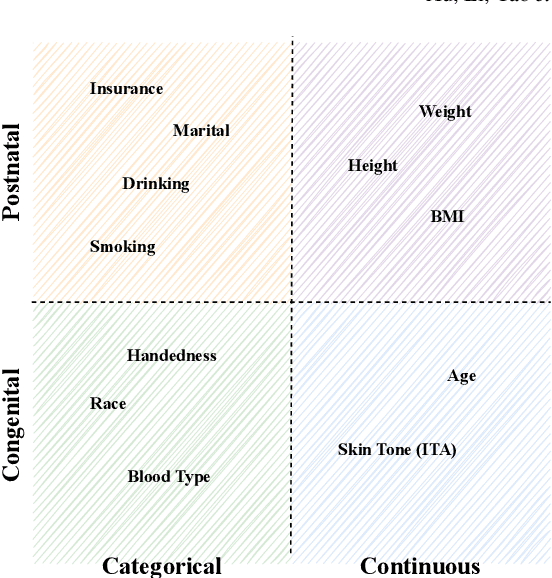
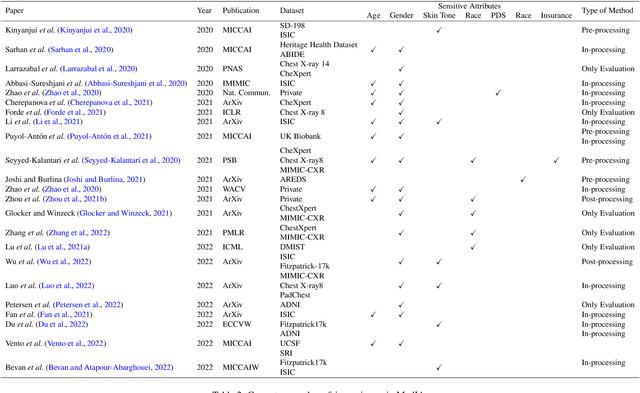
Abstract:Fairness, a criterion focuses on evaluating algorithm performance on different demographic groups, has gained attention in natural language processing, recommendation system and facial recognition. Since there are plenty of demographic attributes in medical image samples, it is important to understand the concepts of fairness, be acquainted with unfairness mitigation techniques, evaluate fairness degree of an algorithm and recognize challenges in fairness issues in medical image analysis (MedIA). In this paper, we first give a comprehensive and precise definition of fairness, following by introducing currently used techniques in fairness issues in MedIA. After that, we list public medical image datasets that contain demographic attributes for facilitating the fairness research and summarize current algorithms concerning fairness in MedIA. To help achieve a better understanding of fairness, and call attention to fairness related issues in MedIA, experiments are conducted comparing the difference between fairness and data imbalance, verifying the existence of unfairness in various MedIA tasks, especially in classification, segmentation and detection, and evaluating the effectiveness of unfairness mitigation algorithms. Finally, we conclude with opportunities and challenges in fairness in MedIA.
Stabilize, Decompose, and Denoise: Self-Supervised Fluoroscopy Denoising
Aug 30, 2022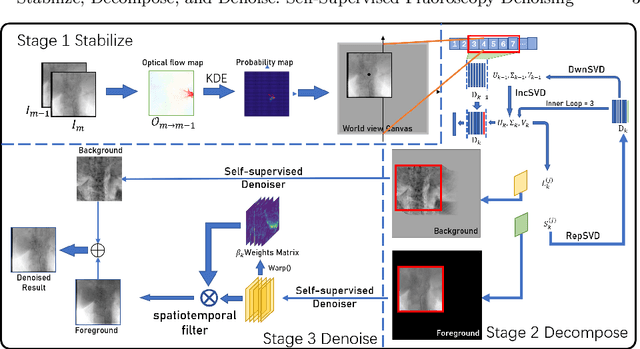
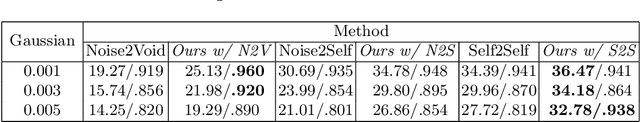
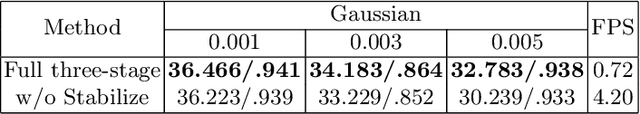
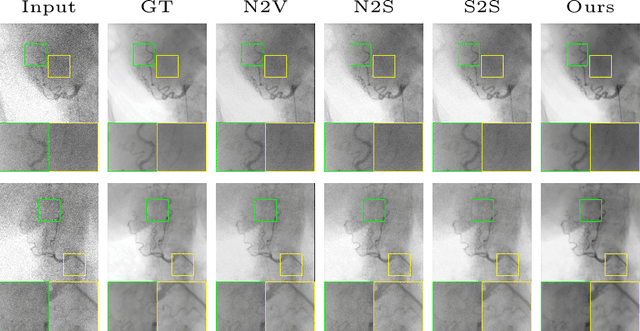
Abstract:Fluoroscopy is an imaging technique that uses X-ray to obtain a real-time 2D video of the interior of a 3D object, helping surgeons to observe pathological structures and tissue functions especially during intervention. However, it suffers from heavy noise that mainly arises from the clinical use of a low dose X-ray, thereby necessitating the technology of fluoroscopy denoising. Such denoising is challenged by the relative motion between the object being imaged and the X-ray imaging system. We tackle this challenge by proposing a self-supervised, three-stage framework that exploits the domain knowledge of fluoroscopy imaging. (i) Stabilize: we first construct a dynamic panorama based on optical flow calculation to stabilize the non-stationary background induced by the motion of the X-ray detector. (ii) Decompose: we then propose a novel mask-based Robust Principle Component Analysis (RPCA) decomposition method to separate a video with detector motion into a low-rank background and a sparse foreground. Such a decomposition accommodates the reading habit of experts. (iii) Denoise: we finally denoise the background and foreground separately by a self-supervised learning strategy and fuse the denoised parts into the final output via a bilateral, spatiotemporal filter. To assess the effectiveness of our work, we curate a dedicated fluoroscopy dataset of 27 videos (1,568 frames) and corresponding ground truth. Our experiments demonstrate that it achieves significant improvements in terms of denoising and enhancement effects when compared with standard approaches. Finally, expert rating confirms this efficacy.
REGAS: REspiratory-GAted Synthesis of Views for Multi-Phase CBCT Reconstruction from a single 3D CBCT Acquisition
Aug 17, 2022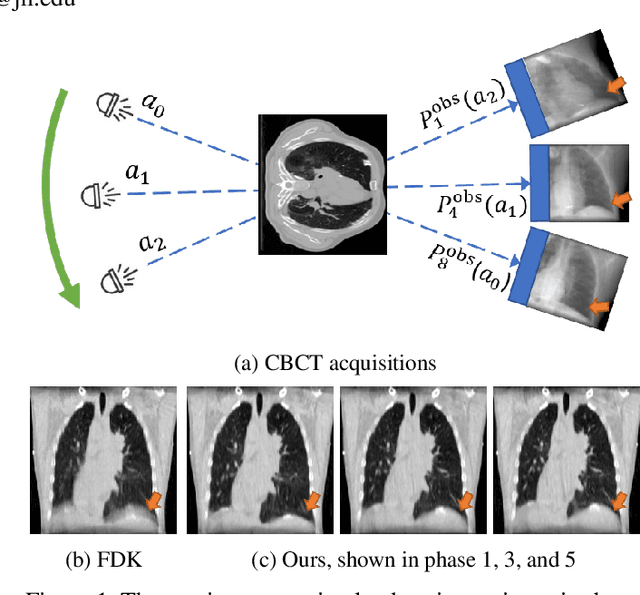
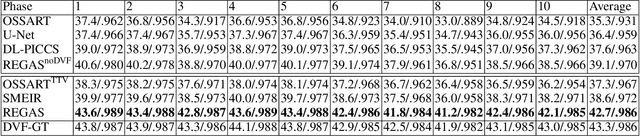
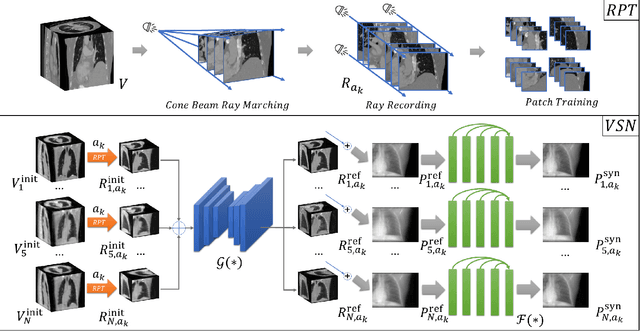
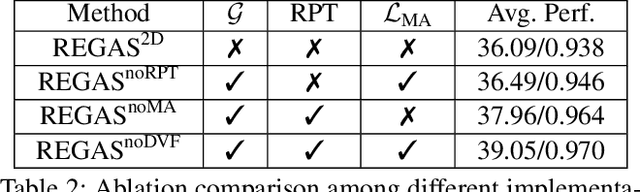
Abstract:It is a long-standing challenge to reconstruct Cone Beam Computed Tomography (CBCT) of the lung under respiratory motion. This work takes a step further to address a challenging setting in reconstructing a multi-phase}4D lung image from just a single}3D CBCT acquisition. To this end, we introduce REpiratory-GAted Synthesis of views, or REGAS. REGAS proposes a self-supervised method to synthesize the undersampled tomographic views and mitigate aliasing artifacts in reconstructed images. This method allows a much better estimation of between-phase Deformation Vector Fields (DVFs), which are used to enhance reconstruction quality from direct observations without synthesis. To address the large memory cost of deep neural networks on high resolution 4D data, REGAS introduces a novel Ray Path Transformation (RPT) that allows for distributed, differentiable forward projections. REGAS require no additional measurements like prior scans, air-flow volume, or breathing velocity. Our extensive experiments show that REGAS significantly outperforms comparable methods in quantitative metrics and visual quality.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge