Reza Razavi
School of Biomedical Engineering and Imaging Sciences, King's College London, London, UK, Department of Adult and Paediatric Cardiology, Guy's and St Thomas' NHS Foundation Trust, London, UK
Automated Quality Controlled Analysis of 2D Phase Contrast Cardiovascular Magnetic Resonance Imaging
Sep 28, 2022
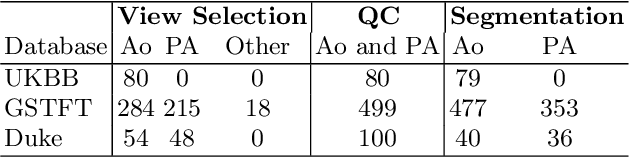
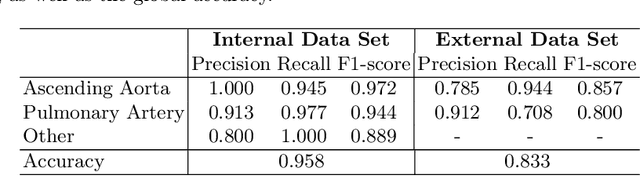
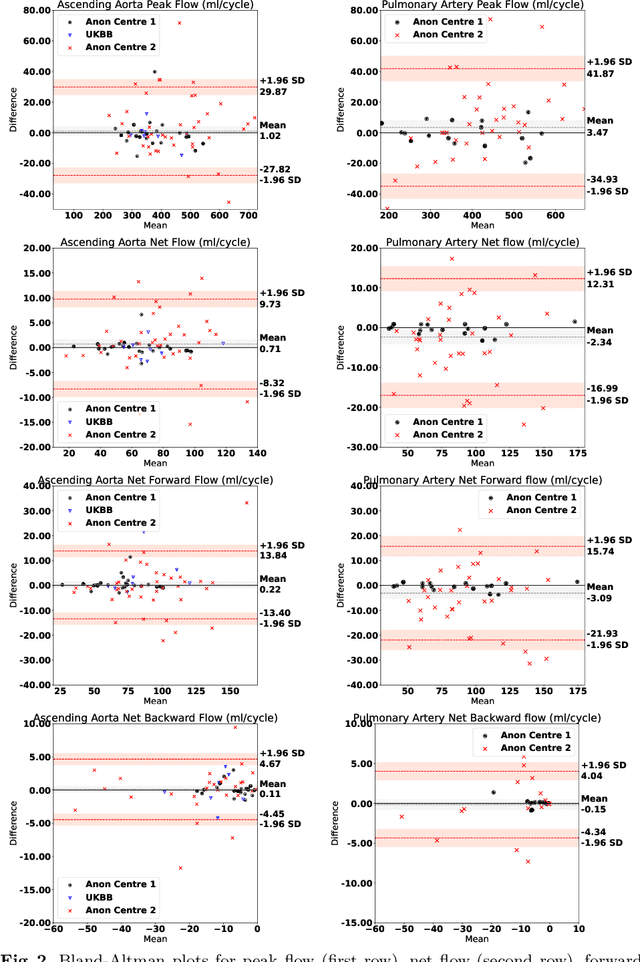
Abstract:Flow analysis carried out using phase contrast cardiac magnetic resonance imaging (PC-CMR) enables the quantification of important parameters that are used in the assessment of cardiovascular function. An essential part of this analysis is the identification of the correct CMR views and quality control (QC) to detect artefacts that could affect the flow quantification. We propose a novel deep learning based framework for the fully-automated analysis of flow from full CMR scans that first carries out these view selection and QC steps using two sequential convolutional neural networks, followed by automatic aorta and pulmonary artery segmentation to enable the quantification of key flow parameters. Accuracy values of 0.958 and 0.914 were obtained for view classification and QC, respectively. For segmentation, Dice scores were $>$0.969 and the Bland-Altman plots indicated excellent agreement between manual and automatic peak flow values. In addition, we tested our pipeline on an external validation data set, with results indicating good robustness of the pipeline. This work was carried out using multivendor clinical data consisting of 986 cases, indicating the potential for the use of this pipeline in a clinical setting.
Large-scale, multi-centre, multi-disease validation of an AI clinical tool for cine CMR analysis
Jun 15, 2022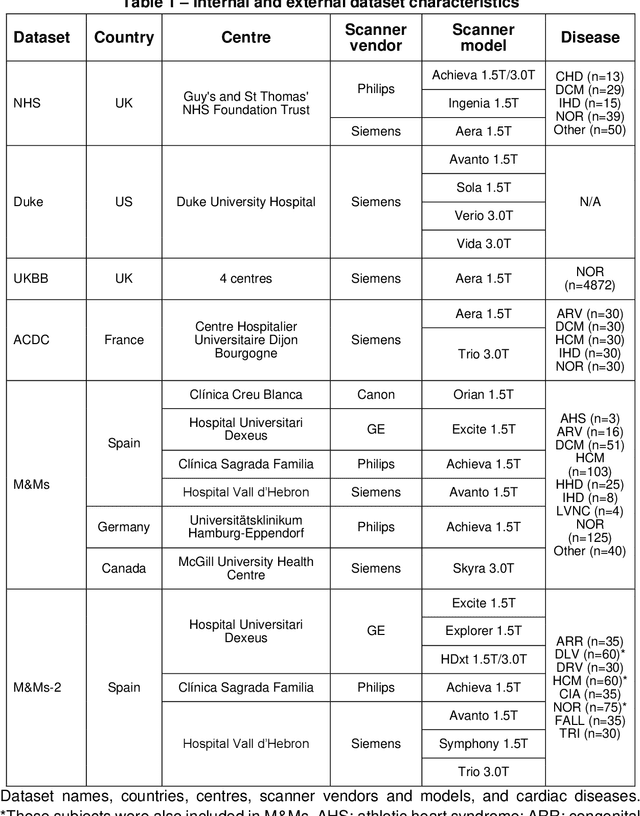
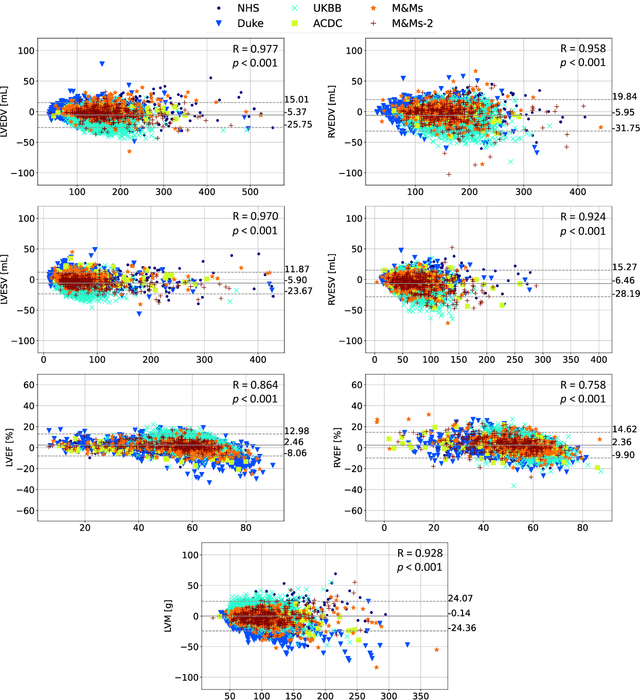
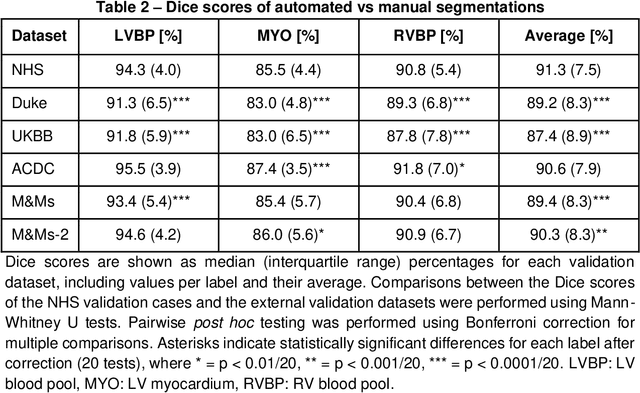
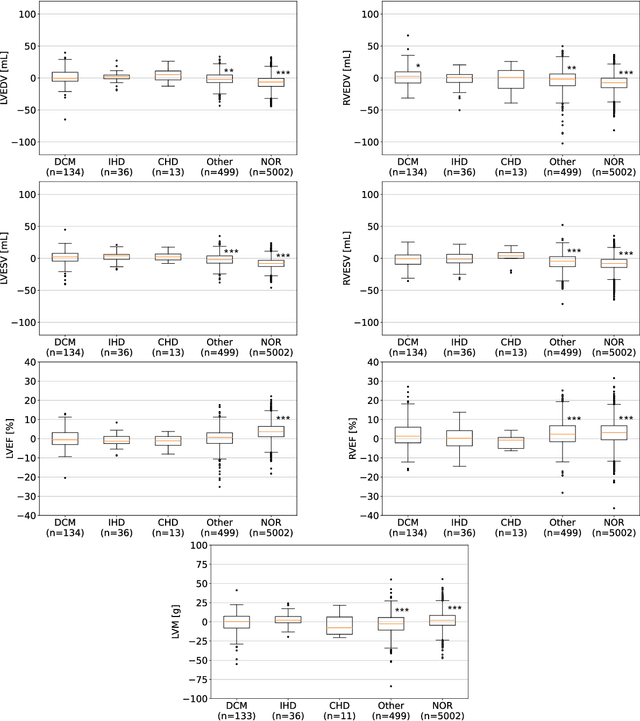
Abstract:INTRODUCTION: Artificial intelligence (AI) has the potential to facilitate the automation of CMR analysis for biomarker extraction. However, most AI algorithms are trained on a specific input domain (e.g., single scanner vendor or hospital-tailored imaging protocol) and lack the robustness to perform optimally when applied to CMR data from other input domains. METHODS: Our proposed framework consists of an AI-based algorithm for biventricular segmentation of short-axis images, followed by a post-analysis quality control to detect erroneous results. The segmentation algorithm was trained on a large dataset of clinical CMR scans from two NHS hospitals (n=2793) and validated on additional cases from this dataset (n=441) and on five external datasets (n=6808). The validation data included CMR scans of patients with a range of diseases acquired at 12 different centres using CMR scanners from all major vendors. RESULTS: Our method yielded median Dice scores over 87%, translating into median absolute errors in cardiac biomarkers within the range of inter-observer variability: <8.4mL (left ventricle), <9.2mL (right ventricle), <13.3g (left ventricular mass), and <5.9% (ejection fraction) across all datasets. Stratification of cases according to phenotypes of cardiac disease and scanner vendors showed good agreement. CONCLUSIONS: We show that our proposed tool, which combines a state-of-the-art AI algorithm trained on a large-scale multi-domain CMR dataset with a post-analysis quality control, allows us to robustly deal with routine clinical data from multiple centres, vendors, and cardiac diseases. This is a fundamental step for the clinical translation of AI algorithms. Moreover, our method yields a range of additional biomarkers of cardiac function (filling and ejection rates, regional wall motion, and strain) at no extra computational cost.
AI-enabled Assessment of Cardiac Systolic and Diastolic Function from Echocardiography
Mar 21, 2022
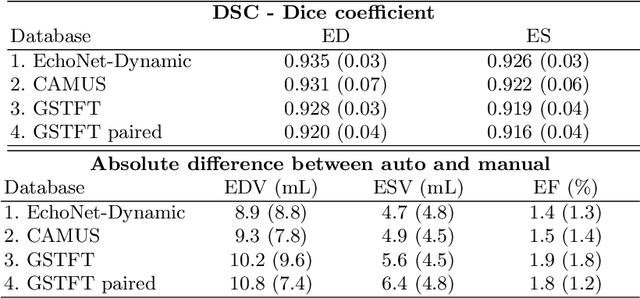
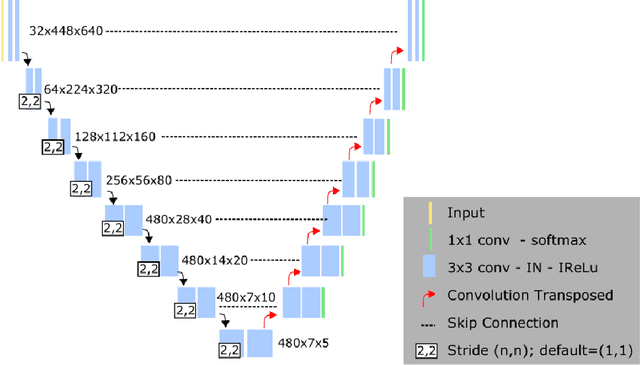
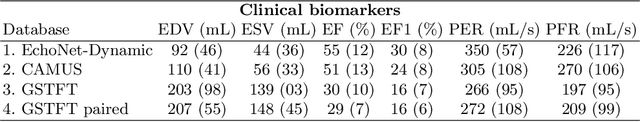
Abstract:Left ventricular (LV) function is an important factor in terms of patient management, outcome, and long-term survival of patients with heart disease. The most recently published clinical guidelines for heart failure recognise that over reliance on only one measure of cardiac function (LV ejection fraction) as a diagnostic and treatment stratification biomarker is suboptimal. Recent advances in AI-based echocardiography analysis have shown excellent results on automated estimation of LV volumes and LV ejection fraction. However, from time-varying 2-D echocardiography acquisition, a richer description of cardiac function can be obtained by estimating functional biomarkers from the complete cardiac cycle. In this work we propose for the first time an AI approach for deriving advanced biomarkers of systolic and diastolic LV function from 2-D echocardiography based on segmentations of the full cardiac cycle. These biomarkers will allow clinicians to obtain a much richer picture of the heart in health and disease. The AI model is based on the 'nn-Unet' framework and was trained and tested using four different databases. Results show excellent agreement between manual and automated analysis and showcase the potential of the advanced systolic and diastolic biomarkers for patient stratification. Finally, for a subset of 50 cases, we perform a correlation analysis between clinical biomarkers derived from echocardiography and CMR and we show excellent agreement between the two modalities.
Uncertainty-Aware Training for Cardiac Resynchronisation Therapy Response Prediction
Sep 22, 2021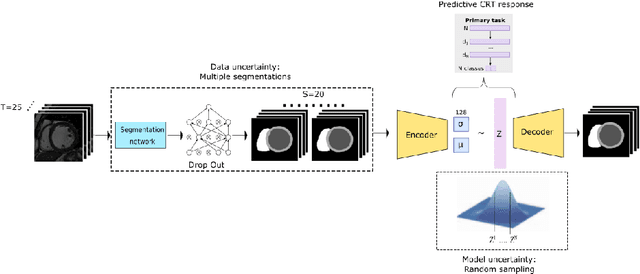
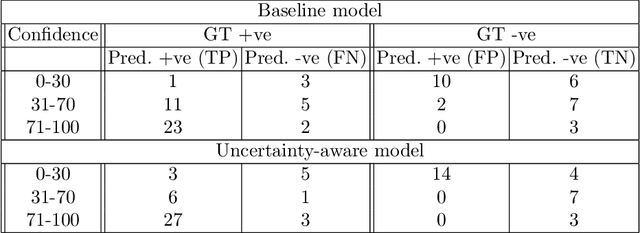
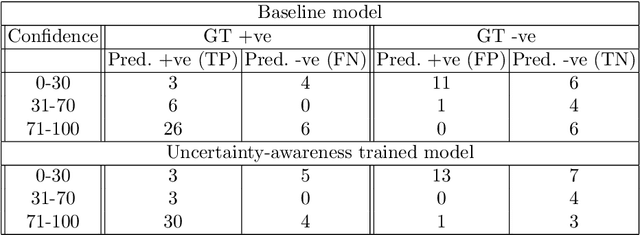
Abstract:Evaluation of predictive deep learning (DL) models beyond conventional performance metrics has become increasingly important for applications in sensitive environments like healthcare. Such models might have the capability to encode and analyse large sets of data but they often lack comprehensive interpretability methods, preventing clinical trust in predictive outcomes. Quantifying uncertainty of a prediction is one way to provide such interpretability and promote trust. However, relatively little attention has been paid to how to include such requirements into the training of the model. In this paper we: (i) quantify the data (aleatoric) and model (epistemic) uncertainty of a DL model for Cardiac Resynchronisation Therapy response prediction from cardiac magnetic resonance images, and (ii) propose and perform a preliminary investigation of an uncertainty-aware loss function that can be used to retrain an existing DL image-based classification model to encourage confidence in correct predictions and reduce confidence in incorrect predictions. Our initial results are promising, showing a significant increase in the (epistemic) confidence of true positive predictions, with some evidence of a reduction in false negative confidence.
Improved AI-based segmentation of apical and basal slices from clinical cine CMR
Sep 20, 2021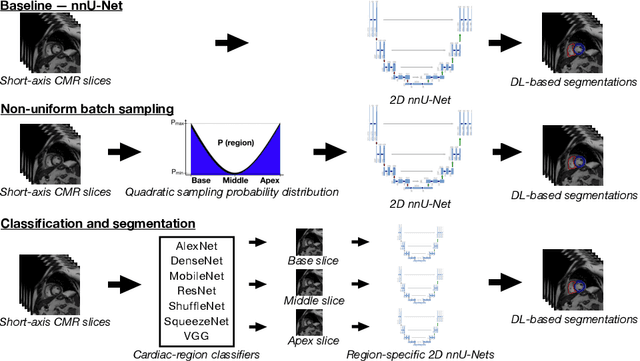
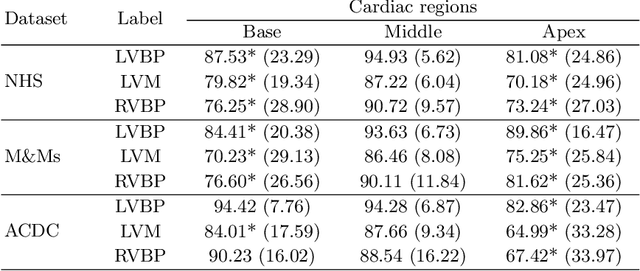
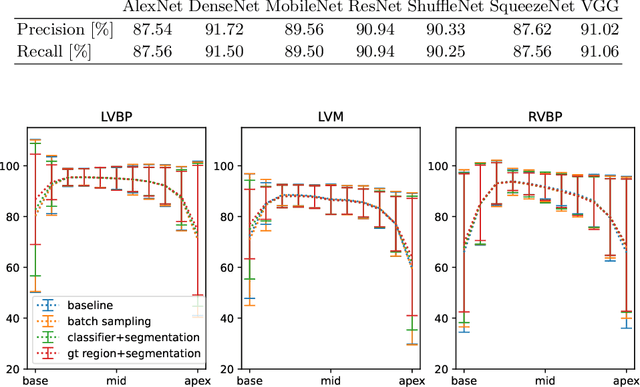
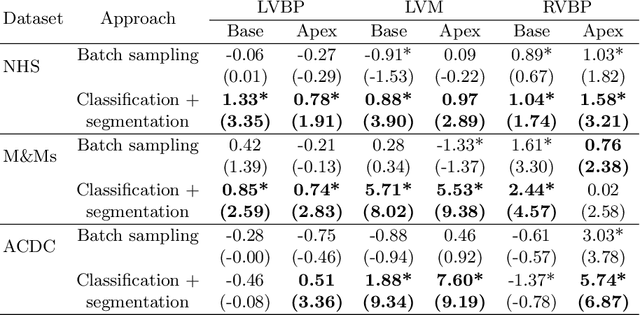
Abstract:Current artificial intelligence (AI) algorithms for short-axis cardiac magnetic resonance (CMR) segmentation achieve human performance for slices situated in the middle of the heart. However, an often-overlooked fact is that segmentation of the basal and apical slices is more difficult. During manual analysis, differences in the basal segmentations have been reported as one of the major sources of disagreement in human interobserver variability. In this work, we aim to investigate the performance of AI algorithms in segmenting basal and apical slices and design strategies to improve their segmentation. We trained all our models on a large dataset of clinical CMR studies obtained from two NHS hospitals (n=4,228) and evaluated them against two external datasets: ACDC (n=100) and M&Ms (n=321). Using manual segmentations as a reference, CMR slices were assigned to one of four regions: non-cardiac, base, middle, and apex. Using the nnU-Net framework as a baseline, we investigated two different approaches to reduce the segmentation performance gap between cardiac regions: (1) non-uniform batch sampling, which allows us to choose how often images from different regions are seen during training; and (2) a cardiac-region classification model followed by three (i.e. base, middle, and apex) region-specific segmentation models. We show that the classification and segmentation approach was best at reducing the performance gap across all datasets. We also show that improvements in the classification performance can subsequently lead to a significantly better performance in the segmentation task.
Can non-specialists provide high quality gold standard labels in challenging modalities?
Jul 30, 2021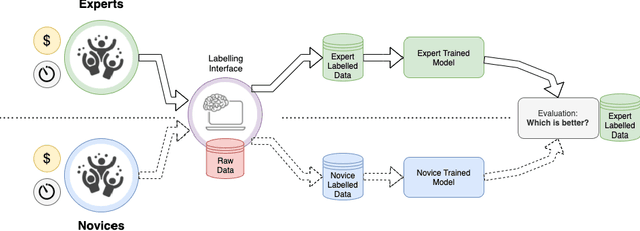
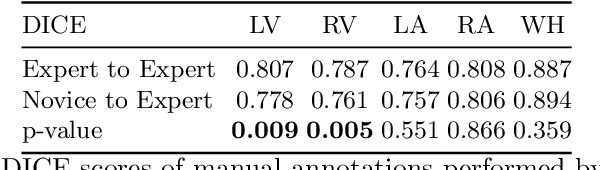
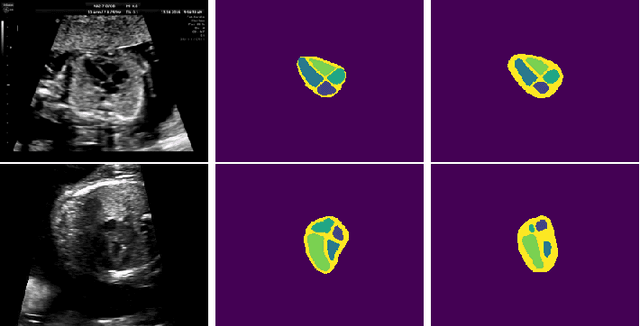
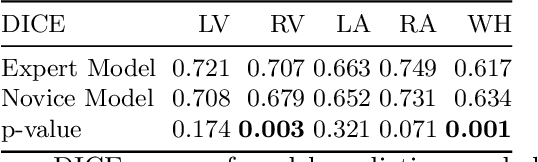
Abstract:Probably yes. -- Supervised Deep Learning dominates performance scores for many computer vision tasks and defines the state-of-the-art. However, medical image analysis lags behind natural image applications. One of the many reasons is the lack of well annotated medical image data available to researchers. One of the first things researchers are told is that we require significant expertise to reliably and accurately interpret and label such data. We see significant inter- and intra-observer variability between expert annotations of medical images. Still, it is a widely held assumption that novice annotators are unable to provide useful annotations for use by clinical Deep Learning models. In this work we challenge this assumption and examine the implications of using a minimally trained novice labelling workforce to acquire annotations for a complex medical image dataset. We study the time and cost implications of using novice annotators, the raw performance of novice annotators compared to gold-standard expert annotators, and the downstream effects on a trained Deep Learning segmentation model's performance for detecting a specific congenital heart disease (hypoplastic left heart syndrome) in fetal ultrasound imaging.
B-line Detection in Lung Ultrasound Videos: Cartesian vs Polar Representation
Jul 26, 2021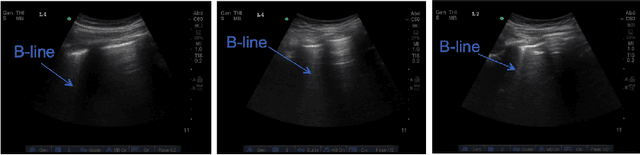

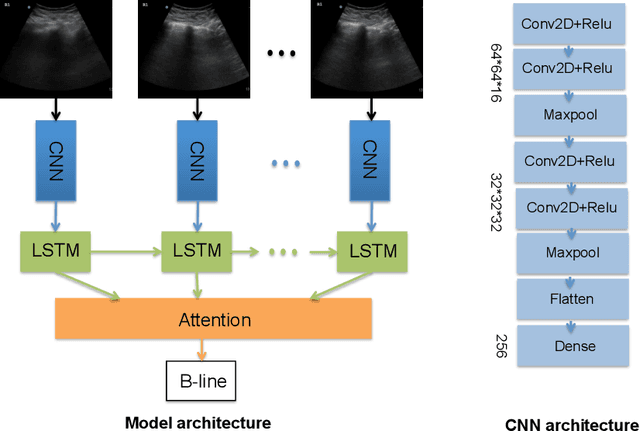
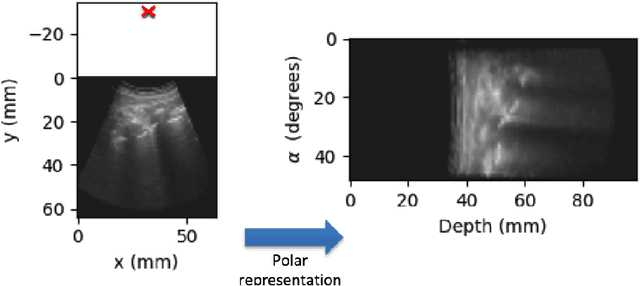
Abstract:Lung ultrasound (LUS) imaging is becoming popular in the intensive care units (ICU) for assessing lung abnormalities such as the appearance of B-line artefacts as a result of severe dengue. These artefacts appear in the LUS images and disappear quickly, making their manual detection very challenging. They also extend radially following the propagation of the sound waves. As a result, we hypothesize that a polar representation may be more adequate for automatic image analysis of these images. This paper presents an attention-based Convolutional+LSTM model to automatically detect B-lines in LUS videos, comparing performance when image data is taken in Cartesian and polar representations. Results indicate that the proposed framework with polar representation achieves competitive performance compared to the Cartesian representation for B-line classification and that attention mechanism can provide better localization.
Detecting Hypo-plastic Left Heart Syndrome in Fetal Ultrasound via Disease-specific Atlas Maps
Jul 06, 2021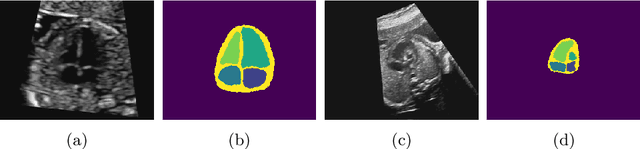
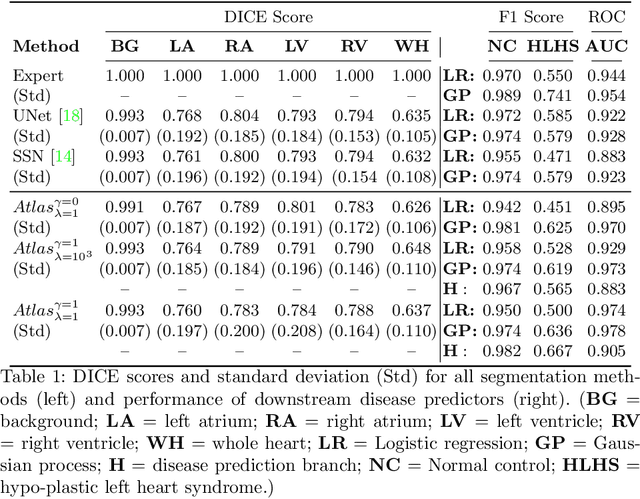
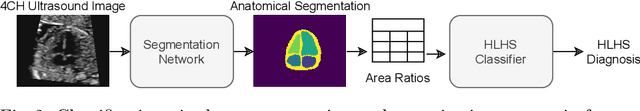
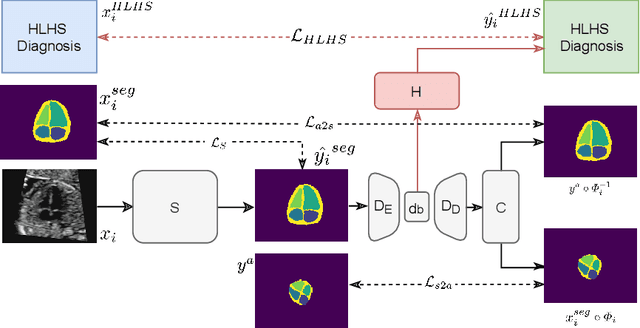
Abstract:Fetal ultrasound screening during pregnancy plays a vital role in the early detection of fetal malformations which have potential long-term health impacts. The level of skill required to diagnose such malformations from live ultrasound during examination is high and resources for screening are often limited. We present an interpretable, atlas-learning segmentation method for automatic diagnosis of Hypo-plastic Left Heart Syndrome (HLHS) from a single `4 Chamber Heart' view image. We propose to extend the recently introduced Image-and-Spatial Transformer Networks (Atlas-ISTN) into a framework that enables sensitising atlas generation to disease. In this framework we can jointly learn image segmentation, registration, atlas construction and disease prediction while providing a maximum level of clinical interpretability compared to direct image classification methods. As a result our segmentation allows diagnoses competitive with expert-derived manual diagnosis and yields an AUC-ROC of 0.978 (1043 cases for training, 260 for validation and 325 for testing).
Fairness in Cardiac MR Image Analysis: An Investigation of Bias Due to Data Imbalance in Deep Learning Based Segmentation
Jul 01, 2021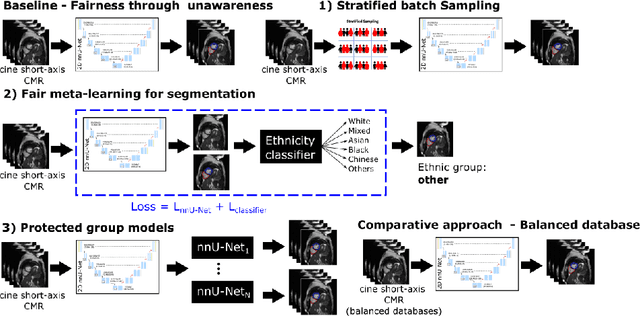
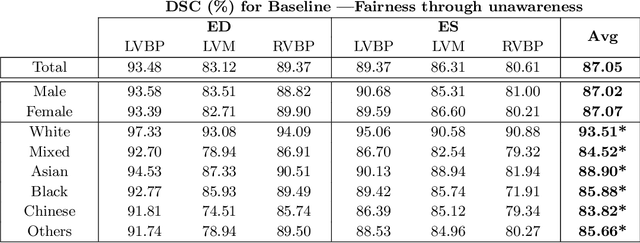
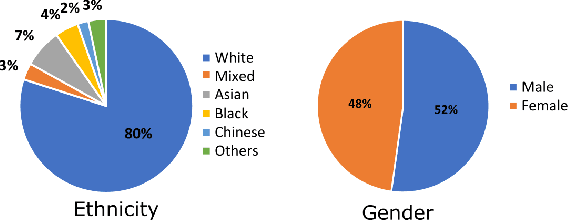
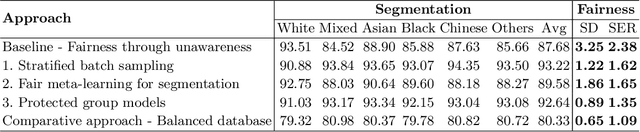
Abstract:The subject of "fairness" in artificial intelligence (AI) refers to assessing AI algorithms for potential bias based on demographic characteristics such as race and gender, and the development of algorithms to address this bias. Most applications to date have been in computer vision, although some work in healthcare has started to emerge. The use of deep learning (DL) in cardiac MR segmentation has led to impressive results in recent years, and such techniques are starting to be translated into clinical practice. However, no work has yet investigated the fairness of such models. In this work, we perform such an analysis for racial/gender groups, focusing on the problem of training data imbalance, using a nnU-Net model trained and evaluated on cine short axis cardiac MR data from the UK Biobank dataset, consisting of 5,903 subjects from 6 different racial groups. We find statistically significant differences in Dice performance between different racial groups. To reduce the racial bias, we investigated three strategies: (1) stratified batch sampling, in which batch sampling is stratified to ensure balance between racial groups; (2) fair meta-learning for segmentation, in which a DL classifier is trained to classify race and jointly optimized with the segmentation model; and (3) protected group models, in which a different segmentation model is trained for each racial group. We also compared the results to the scenario where we have a perfectly balanced database. To assess fairness we used the standard deviation (SD) and skewed error ratio (SER) of the average Dice values. Our results demonstrate that the racial bias results from the use of imbalanced training data, and that all proposed bias mitigation strategies improved fairness, with the best SD and SER resulting from the use of protected group models.
Automatic Detection of B-lines in Lung Ultrasound Videos From Severe Dengue Patients
Feb 01, 2021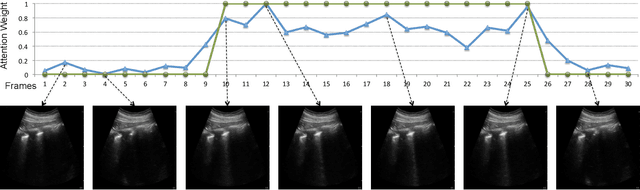
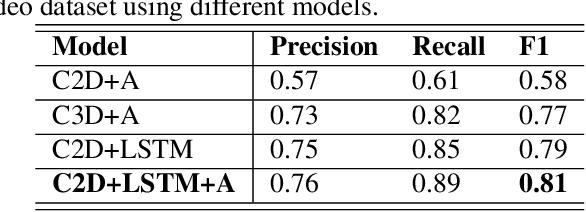
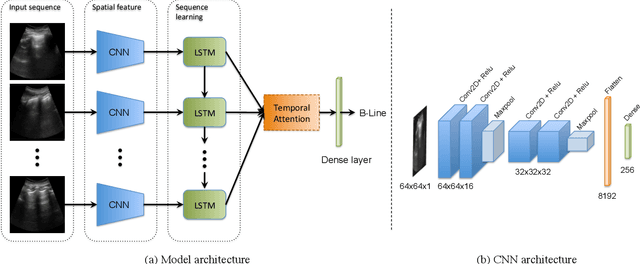
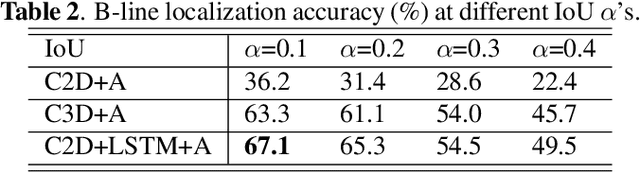
Abstract:Lung ultrasound (LUS) imaging is used to assess lung abnormalities, including the presence of B-line artefacts due to fluid leakage into the lungs caused by a variety of diseases. However, manual detection of these artefacts is challenging. In this paper, we propose a novel methodology to automatically detect and localize B-lines in LUS videos using deep neural networks trained with weak labels. To this end, we combine a convolutional neural network (CNN) with a long short-term memory (LSTM) network and a temporal attention mechanism. Four different models are compared using data from 60 patients. Results show that our best model can determine whether one-second clips contain B-lines or not with an F1 score of 0.81, and extracts a representative frame with B-lines with an accuracy of 87.5%.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge