Regina Barzilay
Path-Augmented Graph Transformer Network
May 29, 2019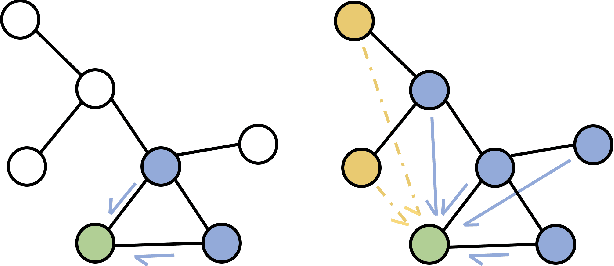
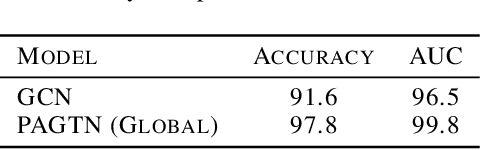
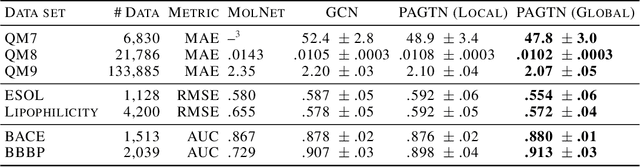

Abstract:Much of the recent work on learning molecular representations has been based on Graph Convolution Networks (GCN). These models rely on local aggregation operations and can therefore miss higher-order graph properties. To remedy this, we propose Path-Augmented Graph Transformer Networks (PAGTN) that are explicitly built on longer-range dependencies in graph-structured data. Specifically, we use path features in molecular graphs to create global attention layers. We compare our PAGTN model against the GCN model and show that our model consistently outperforms GCNs on molecular property prediction datasets including quantum chemistry (QM7, QM8, QM9), physical chemistry (ESOL, Lipophilictiy) and biochemistry (BACE, BBBP).
Using Machine Learning and Natural Language Processing to Review and Classify the Medical Literature on Cancer Susceptibility Genes
Apr 24, 2019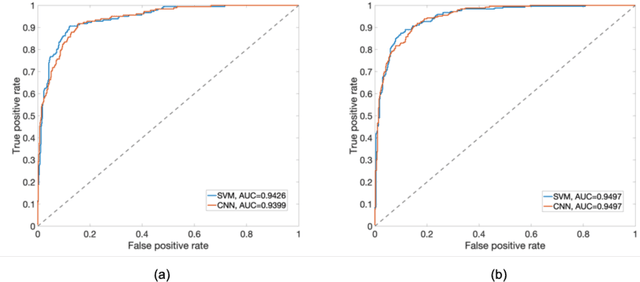
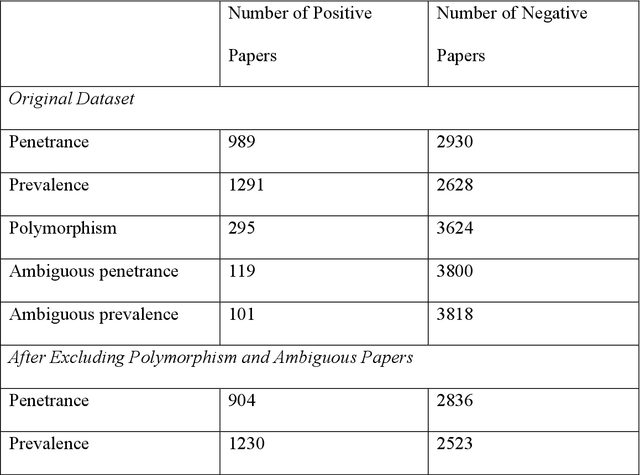
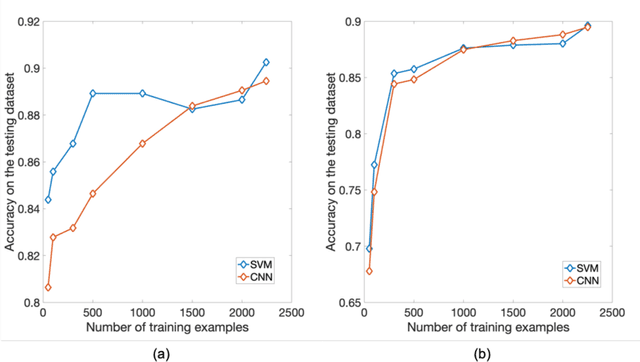
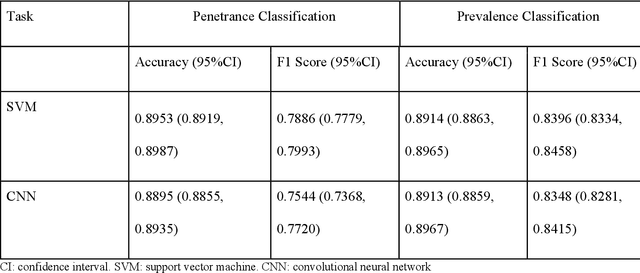
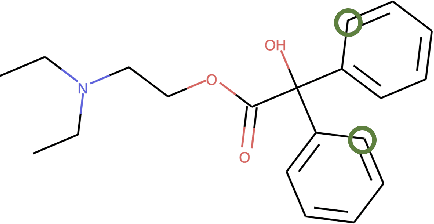
Abstract:PURPOSE: The medical literature relevant to germline genetics is growing exponentially. Clinicians need tools monitoring and prioritizing the literature to understand the clinical implications of the pathogenic genetic variants. We developed and evaluated two machine learning models to classify abstracts as relevant to the penetrance (risk of cancer for germline mutation carriers) or prevalence of germline genetic mutations. METHODS: We conducted literature searches in PubMed and retrieved paper titles and abstracts to create an annotated dataset for training and evaluating the two machine learning classification models. Our first model is a support vector machine (SVM) which learns a linear decision rule based on the bag-of-ngrams representation of each title and abstract. Our second model is a convolutional neural network (CNN) which learns a complex nonlinear decision rule based on the raw title and abstract. We evaluated the performance of the two models on the classification of papers as relevant to penetrance or prevalence. RESULTS: For penetrance classification, we annotated 3740 paper titles and abstracts and used 60% for training the model, 20% for tuning the model, and 20% for evaluating the model. The SVM model achieves 89.53% accuracy (percentage of papers that were correctly classified) while the CNN model achieves 88.95 % accuracy. For prevalence classification, we annotated 3753 paper titles and abstracts. The SVM model achieves 89.14% accuracy while the CNN model achieves 89.13 % accuracy. CONCLUSION: Our models achieve high accuracy in classifying abstracts as relevant to penetrance or prevalence. By facilitating literature review, this tool could help clinicians and researchers keep abreast of the burgeoning knowledge of gene-cancer associations and keep the knowledge bases for clinical decision support tools up to date.
Inferring Which Medical Treatments Work from Reports of Clinical Trials
Apr 04, 2019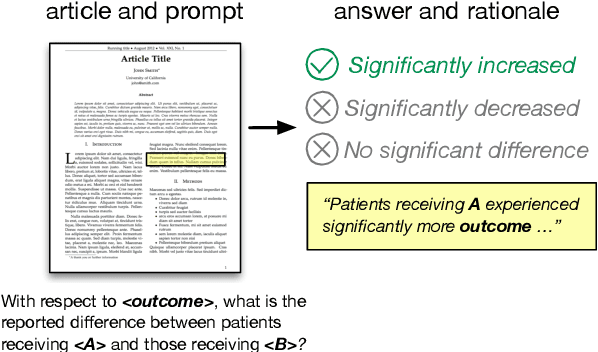

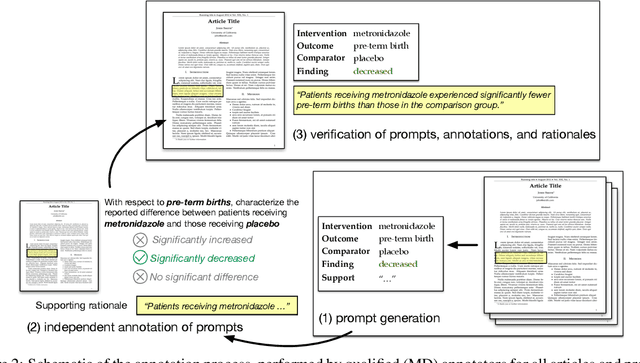
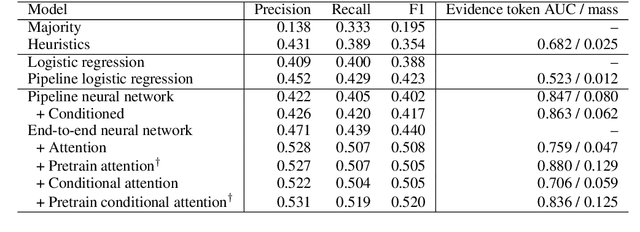
Abstract:How do we know if a particular medical treatment actually works? Ideally one would consult all available evidence from relevant clinical trials. Unfortunately, such results are primarily disseminated in natural language scientific articles, imposing substantial burden on those trying to make sense of them. In this paper, we present a new task and corpus for making this unstructured evidence actionable. The task entails inferring reported findings from a full-text article describing a randomized controlled trial (RCT) with respect to a given intervention, comparator, and outcome of interest, e.g., inferring if an article provides evidence supporting the use of aspirin to reduce risk of stroke, as compared to placebo. We present a new corpus for this task comprising 10,000+ prompts coupled with full-text articles describing RCTs. Results using a suite of models --- ranging from heuristic (rule-based) approaches to attentive neural architectures --- demonstrate the difficulty of the task, which we believe largely owes to the lengthy, technical input texts. To facilitate further work on this important, challenging problem we make the corpus, documentation, a website and leaderboard, and code for baselines and evaluation available at http://evidence-inference.ebm-nlp.com/.
Cross-Lingual Alignment of Contextual Word Embeddings, with Applications to Zero-shot Dependency Parsing
Apr 04, 2019
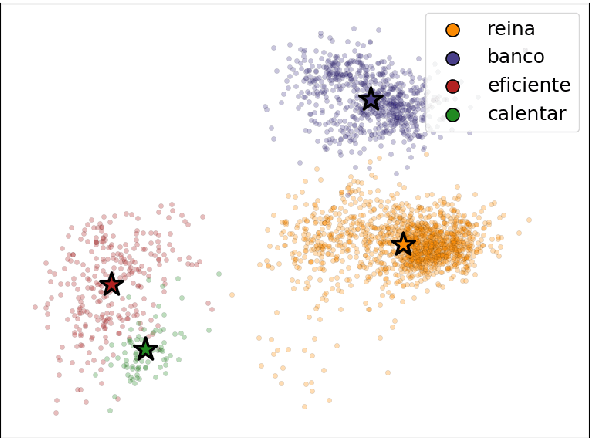
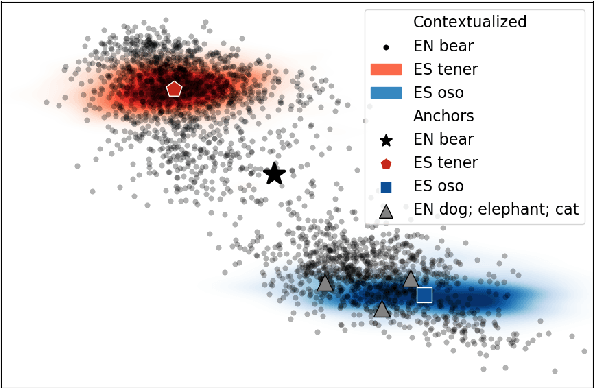

Abstract:We introduce a novel method for multilingual transfer that utilizes deep contextual embeddings, pretrained in an unsupervised fashion. While contextual embeddings have been shown to yield richer representations of meaning compared to their static counterparts, aligning them poses a challenge due to their dynamic nature. To this end, we construct context-independent variants of the original monolingual spaces and utilize their mapping to derive an alignment for the context-dependent spaces. This mapping readily supports processing of a target language, improving transfer by context-aware embeddings. Our experimental results demonstrate the effectiveness of this approach for zero-shot and few-shot learning of dependency parsing. Specifically, our method consistently outperforms the previous state-of-the-art on 6 tested languages, yielding an improvement of 6.8 LAS points on average.
Are Learned Molecular Representations Ready For Prime Time?
Apr 02, 2019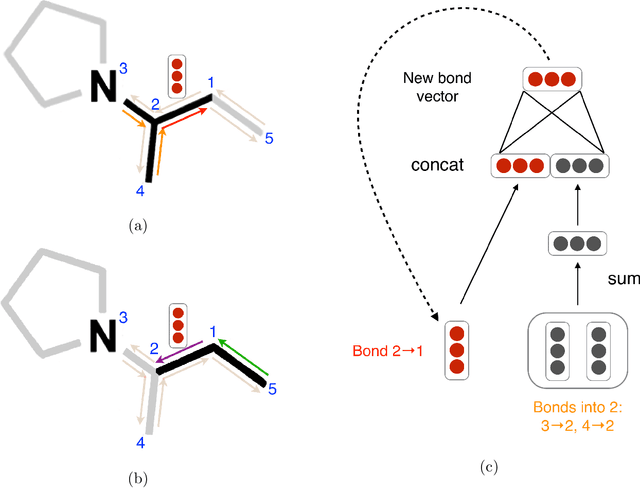
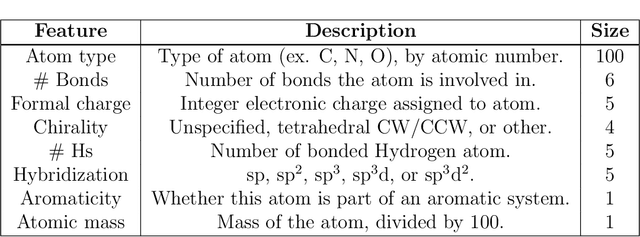
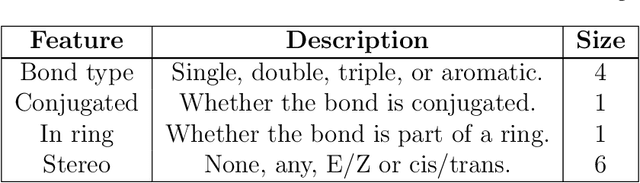


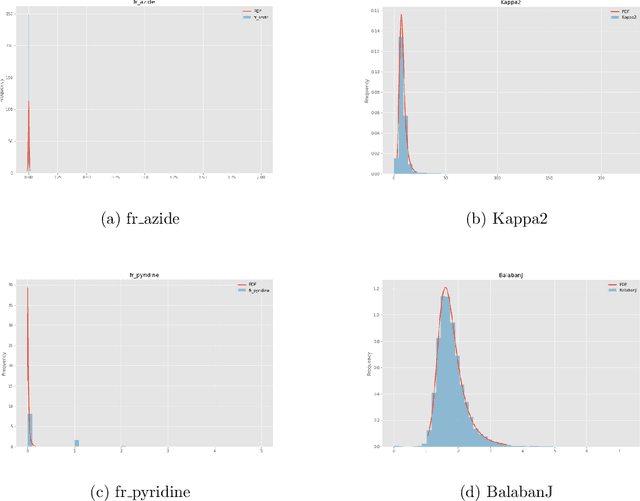
Abstract:Advancements in neural machinery have led to a wide range of algorithmic solutions for molecular property prediction. Two classes of models in particular have yielded promising results: neural networks applied to computed molecular fingerprints or expert-crafted descriptors, and graph convolutional neural networks that construct a learned molecular representation by operating on the graph structure of the molecule. However, recent literature has yet to clearly determine which of these two methods is superior when generalizing to new chemical space. Furthermore, prior research has rarely examined these new models in industry research settings in comparison to existing employed models. In this paper, we benchmark models extensively on 19 public and 15 proprietary industrial datasets spanning a wide variety of chemical endpoints. In addition, we introduce a graph convolutional model that consistently outperforms models using fixed molecular descriptors as well as previous graph neural architectures on both public and proprietary datasets. Our empirical findings indicate that while approaches based on these representations have yet to reach the level of experimental reproducibility, our proposed model nevertheless offers significant improvements over models currently used in industrial workflows.
Learning Multimodal Graph-to-Graph Translation for Molecular Optimization
Jan 07, 2019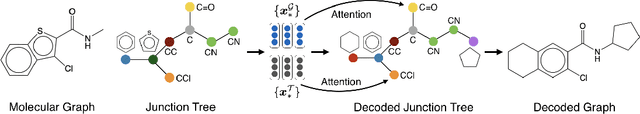
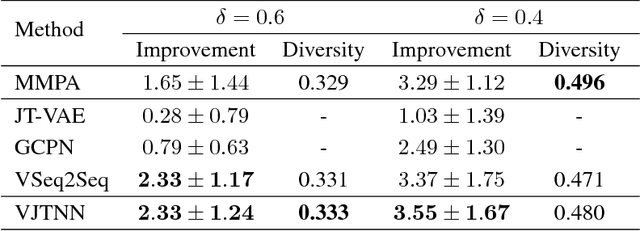
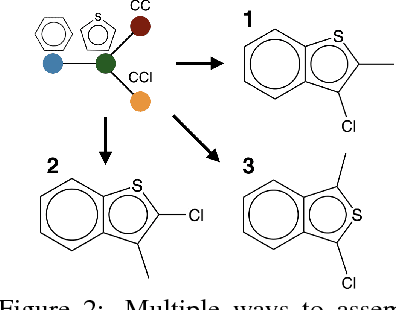
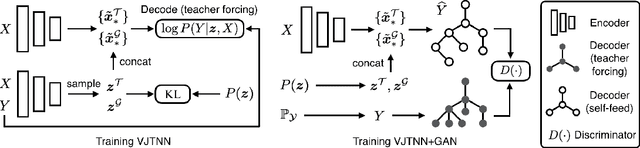
Abstract:We view molecular optimization as a graph-to-graph translation problem. The goal is to learn to map from one molecular graph to another with better properties based on an available corpus of paired molecules. Since molecules can be optimized in different ways, there are multiple viable translations for each input graph. A key challenge is therefore to model diverse translation outputs. Our primary contributions include a junction tree encoder-decoder for learning diverse graph translations along with a novel adversarial training method for aligning distributions of molecules. Diverse output distributions in our model are explicitly realized by low-dimensional latent vectors that modulate the translation process. We evaluate our model on multiple molecular optimization tasks and show that our model outperforms previous state-of-the-art baselines.
GraphIE: A Graph-Based Framework for Information Extraction
Oct 31, 2018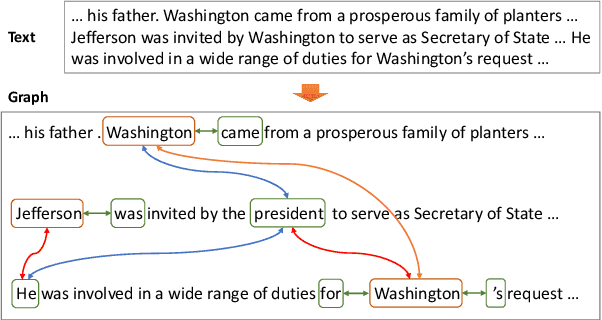
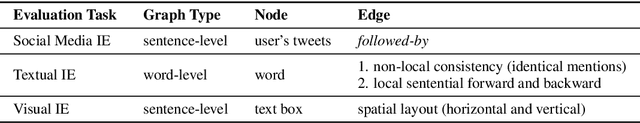

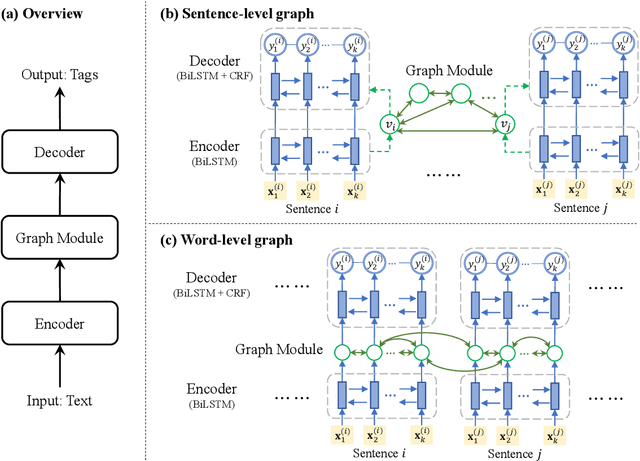
Abstract:Most modern Information Extraction (IE) systems are implemented as sequential taggers and focus on modelling local dependencies. Non-local and non-sequential context is, however, a valuable source of information to improve predictions. In this paper, we introduce GraphIE, a framework that operates over a graph representing both local and non-local dependencies between textual units (i.e. words or sentences). The algorithm propagates information between connected nodes through graph convolutions and exploits the richer representation to improve word level predictions. The framework is evaluated on three different tasks, namely social media, textual and visual information extraction. Results show that GraphIE outperforms a competitive baseline (BiLSTM+CRF) in all tasks by a significant margin.
Multi-Source Domain Adaptation with Mixture of Experts
Oct 16, 2018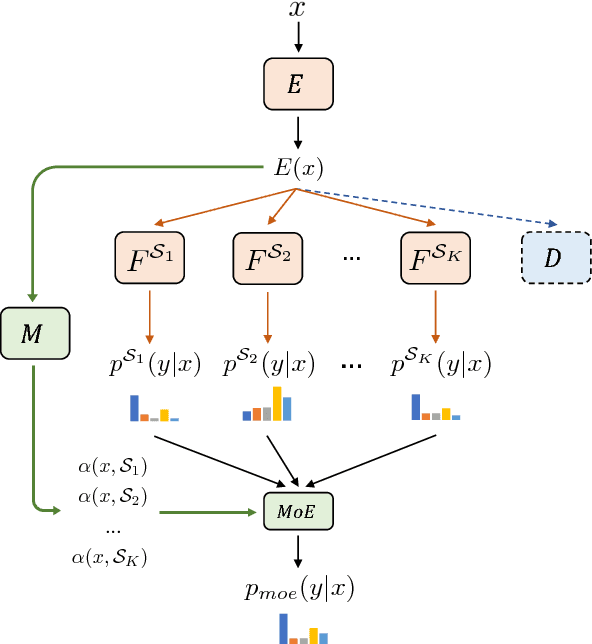
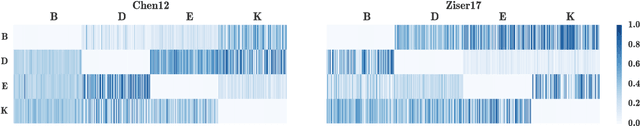
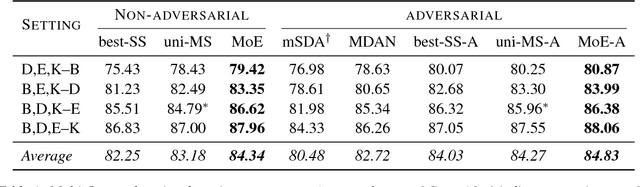
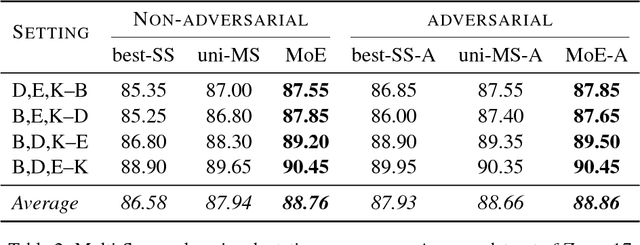

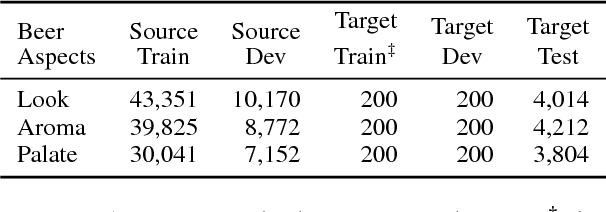
Abstract:We propose a mixture-of-experts approach for unsupervised domain adaptation from multiple sources. The key idea is to explicitly capture the relationship between a target example and different source domains. This relationship, expressed by a point-to-set metric, determines how to combine predictors trained on various domains. The metric is learned in an unsupervised fashion using meta-training. Experimental results on sentiment analysis and part-of-speech tagging demonstrate that our approach consistently outperforms multiple baselines and can robustly handle negative transfer.
Deriving Machine Attention from Human Rationales
Aug 28, 2018
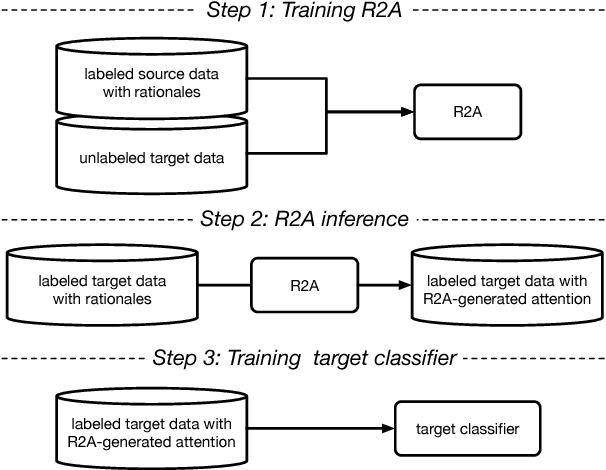
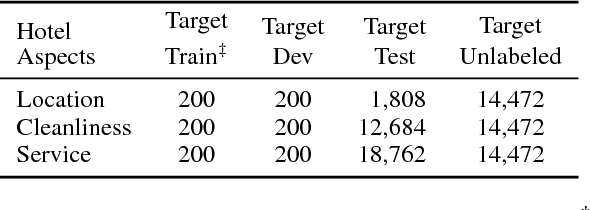
Abstract:Attention-based models are successful when trained on large amounts of data. In this paper, we demonstrate that even in the low-resource scenario, attention can be learned effectively. To this end, we start with discrete human-annotated rationales and map them into continuous attention. Our central hypothesis is that this mapping is general across domains, and thus can be transferred from resource-rich domains to low-resource ones. Our model jointly learns a domain-invariant representation and induces the desired mapping between rationales and attention. Our empirical results validate this hypothesis and show that our approach delivers significant gains over state-of-the-art baselines, yielding over 15% average error reduction on benchmark datasets.
The Three Pillars of Machine Programming
May 08, 2018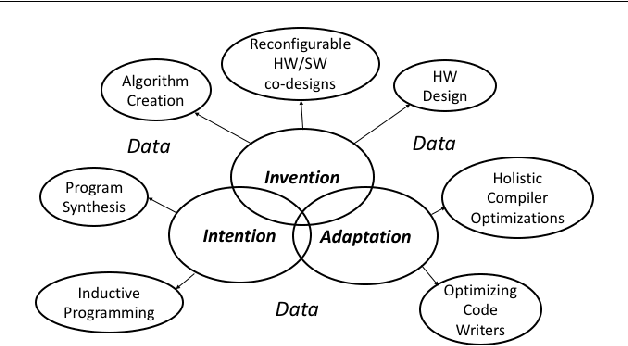
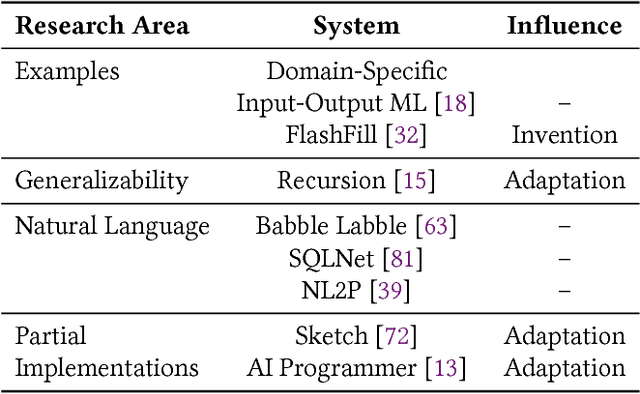
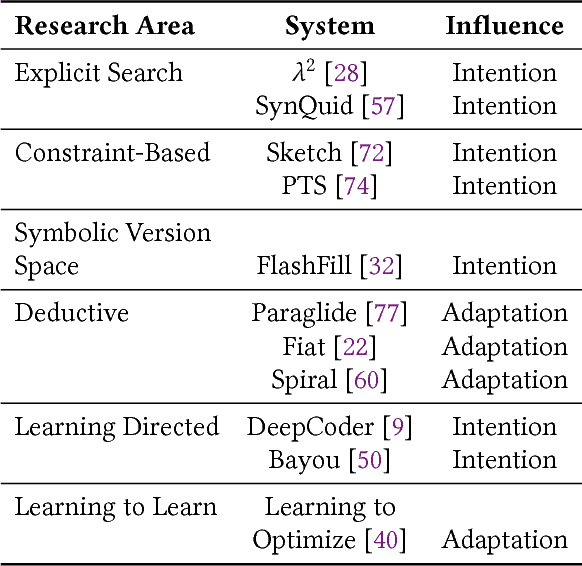
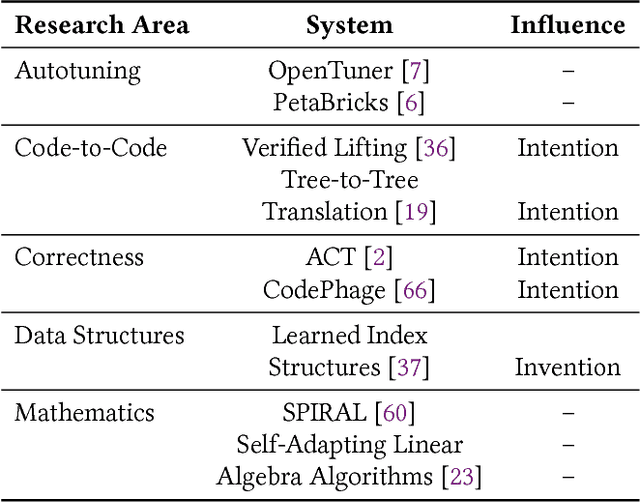
Abstract:In this position paper, we describe our vision of the future of machine programming through a categorical examination of three pillars of research. Those pillars are: (i) intention, (ii) invention, and(iii) adaptation. Intention emphasizes advancements in the human-to-computer and computer-to-machine-learning interfaces. Invention emphasizes the creation or refinement of algorithms or core hardware and software building blocks through machine learning (ML). Adaptation emphasizes advances in the use of ML-based constructs to autonomously evolve software.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge