Yujia Bao
Ares: Adaptive Reasoning Effort Selection for Efficient LLM Agents
Mar 09, 2026Abstract:Modern agents powered by thinking LLMs achieve high accuracy through long chain-of-thought reasoning but incur substantial inference costs. While many LLMs now support configurable reasoning levels (e.g., high/medium/low), static strategies are often ineffective: using low-effort modes at every step leads to significant performance degradation, while random selection fails to preserve accuracy or provide meaningful cost reduction. However, agents should reserve high reasoning effort for difficult steps like navigating complex website structures, while using lower-effort modes for simpler steps like opening a target URL. In this paper, we propose Ares, a framework for per-step dynamic reasoning effort selection tailored for multi-step agent tasks. Ares employs a lightweight router to predict the lowest appropriate reasoning level for each step based on the interaction history. To train this router, we develop a data generation pipeline that identifies the minimum reasoning effort required for successful step completion. We then fine-tune the router to predict these levels, enabling plug-and-play integration for any LLM agents. We evaluate Ares on a diverse set of agent tasks, including TAU-Bench for tool use agents, BrowseComp-Plus for deep-research agents, and WebArena for web agents. Experimental results show that Ares reduces reasoning token usage by up to 52.7% compared to fixed high-effort reasoning, while introducing minimal degradation in task success rates.
Observations and Remedies for Large Language Model Bias in Self-Consuming Performative Loop
Jan 08, 2026Abstract:The rapid advancement of large language models (LLMs) has led to growing interest in using synthetic data to train future models. However, this creates a self-consuming retraining loop, where models are trained on their own outputs and may cause performance drops and induce emerging biases. In real-world applications, previously deployed LLMs may influence the data they generate, leading to a dynamic system driven by user feedback. For example, if a model continues to underserve users from a group, less query data will be collected from this particular demographic of users. In this study, we introduce the concept of \textbf{S}elf-\textbf{C}onsuming \textbf{P}erformative \textbf{L}oop (\textbf{SCPL}) and investigate the role of synthetic data in shaping bias during these dynamic iterative training processes under controlled performative feedback. This controlled setting is motivated by the inaccessibility of real-world user preference data from dynamic production systems, and enables us to isolate and analyze feedback-driven bias evolution in a principled manner. We focus on two types of loops, including the typical retraining setting and the incremental fine-tuning setting, which is largely underexplored. Through experiments on three real-world tasks, we find that the performative loop increases preference bias and decreases disparate bias. We design a reward-based rejection sampling strategy to mitigate the bias, moving towards more trustworthy self-improving systems.
DRAGON: Guard LLM Unlearning in Context via Negative Detection and Reasoning
Nov 11, 2025



Abstract:Unlearning in Large Language Models (LLMs) is crucial for protecting private data and removing harmful knowledge. Most existing approaches rely on fine-tuning to balance unlearning efficiency with general language capabilities. However, these methods typically require training or access to retain data, which is often unavailable in real world scenarios. Although these methods can perform well when both forget and retain data are available, few works have demonstrated equivalent capability in more practical, data-limited scenarios. To overcome these limitations, we propose Detect-Reasoning Augmented GeneratiON (DRAGON), a systematic, reasoning-based framework that utilizes in-context chain-of-thought (CoT) instructions to guard deployed LLMs before inference. Instead of modifying the base model, DRAGON leverages the inherent instruction-following ability of LLMs and introduces a lightweight detection module to identify forget-worthy prompts without any retain data. These are then routed through a dedicated CoT guard model to enforce safe and accurate in-context intervention. To robustly evaluate unlearning performance, we introduce novel metrics for unlearning performance and the continual unlearning setting. Extensive experiments across three representative unlearning tasks validate the effectiveness of DRAGON, demonstrating its strong unlearning capability, scalability, and applicability in practical scenarios.
SFT-GO: Supervised Fine-Tuning with Group Optimization for Large Language Models
Jun 17, 2025Abstract:Supervised fine-tuning (SFT) has become an essential step in tailoring large language models (LLMs) to align with human expectations and specific downstream tasks. However, existing SFT methods typically treat each training instance as a uniform sequence, giving equal importance to all tokens regardless of their relevance. This overlooks the fact that only a subset of tokens often contains critical, task-specific information. To address this limitation, we introduce Supervised Fine-Tuning with Group Optimization (SFT-GO), a novel approach that treats groups of tokens differently based on their importance.SFT-GO groups tokens in each sample based on their importance values and optimizes the LLM using a weighted combination of the worst-group loss and the standard cross-entropy loss. This mechanism adaptively emphasizes the most challenging token groups and guides the model to better handle different group distributions, thereby improving overall learning dynamics. We provide a theoretical analysis of SFT-GO's convergence rate, demonstrating its efficiency. Empirically, we apply SFT-GO with three different token grouping strategies and show that models trained with SFT-GO consistently outperform baseline approaches across popular LLM benchmarks. These improvements hold across various datasets and base models, demonstrating the robustness and the effectiveness of our method.
Collaborative Memory: Multi-User Memory Sharing in LLM Agents with Dynamic Access Control
May 23, 2025Abstract:Complex tasks are increasingly delegated to ensembles of specialized LLM-based agents that reason, communicate, and coordinate actions-both among themselves and through interactions with external tools, APIs, and databases. While persistent memory has been shown to enhance single-agent performance, most approaches assume a monolithic, single-user context-overlooking the benefits and challenges of knowledge transfer across users under dynamic, asymmetric permissions. We introduce Collaborative Memory, a framework for multi-user, multi-agent environments with asymmetric, time-evolving access controls encoded as bipartite graphs linking users, agents, and resources. Our system maintains two memory tiers: (1) private memory-private fragments visible only to their originating user; and (2) shared memory-selectively shared fragments. Each fragment carries immutable provenance attributes (contributing agents, accessed resources, and timestamps) to support retrospective permission checks. Granular read policies enforce current user-agent-resource constraints and project existing memory fragments into filtered transformed views. Write policies determine fragment retention and sharing, applying context-aware transformations to update the memory. Both policies may be designed conditioned on system, agent, and user-level information. Our framework enables safe, efficient, and interpretable cross-user knowledge sharing, with provable adherence to asymmetric, time-varying policies and full auditability of memory operations.
Advertising in AI systems: Society must be vigilant
May 23, 2025



Abstract:AI systems have increasingly become our gateways to the Internet. We argue that just as advertising has driven the monetization of web search and social media, so too will commercial incentives shape the content served by AI. Unlike traditional media, however, the outputs of these systems are dynamic, personalized, and lack clear provenance -- raising concerns for transparency and regulation. In this paper, we envision how commercial content could be delivered through generative AI-based systems. Based on the requirements of key stakeholders -- advertisers, consumers, and platforms -- we propose design principles for commercially-influenced AI systems. We then outline high-level strategies for end users to identify and mitigate commercial biases from model outputs. Finally, we conclude with open questions and a call to action towards these goals.
KVLink: Accelerating Large Language Models via Efficient KV Cache Reuse
Feb 21, 2025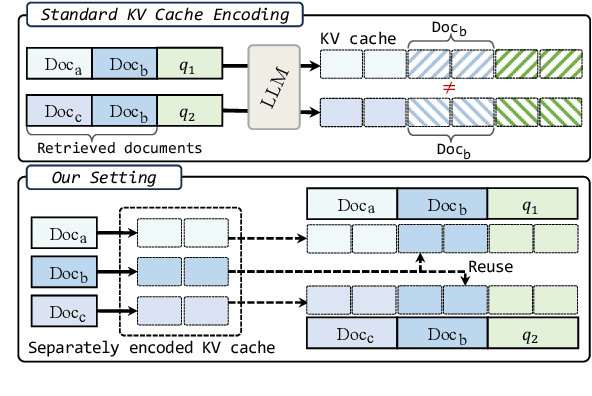
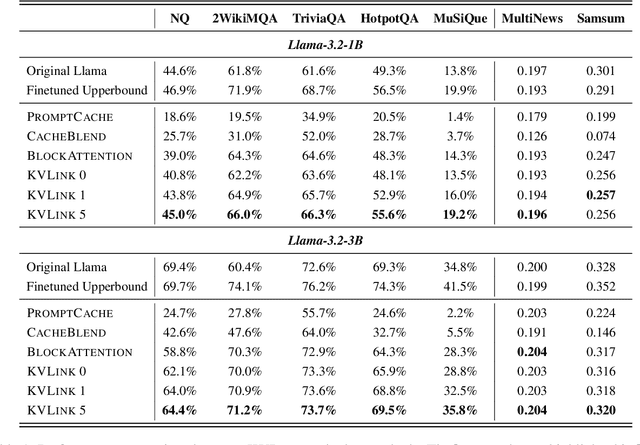
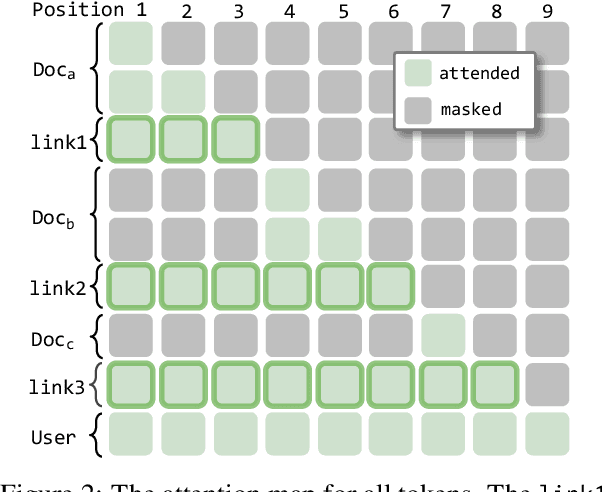
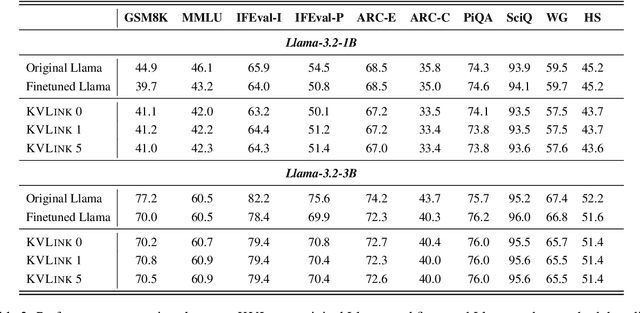
Abstract:We describe KVLink, an approach for efficient key-value (KV) cache reuse in large language models (LLMs). In many LLM applications, different inputs can share overlapping context, such as the same retrieved document appearing in multiple queries. However, the LLMs still need to encode the entire context for each query, leading to redundant computation. In this paper, we propose a new strategy to eliminate such inefficiency, where the KV cache of each document is precomputed independently. During inference, the KV caches of retrieved documents are concatenated, allowing the model to reuse cached representations instead of recomputing them. To mitigate the performance degradation of LLMs when using KV caches computed independently for each document, KVLink introduces three key components: adjusting positional embeddings of the KV cache at inference to match the global position after concatenation, using trainable special tokens to restore self-attention across independently encoded documents, and applying mixed-data fine-tuning to enhance performance while preserving the model's original capabilities. Experiments across 7 datasets demonstrate that KVLink improves question answering accuracy by an average of 4% over state-of-the-art methods. Furthermore, by leveraging precomputed KV caches, our approach reduces time-to-first-token by up to 90% compared to standard LLM inference, making it a scalable and efficient solution for context reuse.
H-CoT: Hijacking the Chain-of-Thought Safety Reasoning Mechanism to Jailbreak Large Reasoning Models, Including OpenAI o1/o3, DeepSeek-R1, and Gemini 2.0 Flash Thinking
Feb 18, 2025



Abstract:Large Reasoning Models (LRMs) have recently extended their powerful reasoning capabilities to safety checks-using chain-of-thought reasoning to decide whether a request should be answered. While this new approach offers a promising route for balancing model utility and safety, its robustness remains underexplored. To address this gap, we introduce Malicious-Educator, a benchmark that disguises extremely dangerous or malicious requests beneath seemingly legitimate educational prompts. Our experiments reveal severe security flaws in popular commercial-grade LRMs, including OpenAI o1/o3, DeepSeek-R1, and Gemini 2.0 Flash Thinking. For instance, although OpenAI's o1 model initially maintains a high refusal rate of about 98%, subsequent model updates significantly compromise its safety; and attackers can easily extract criminal strategies from DeepSeek-R1 and Gemini 2.0 Flash Thinking without any additional tricks. To further highlight these vulnerabilities, we propose Hijacking Chain-of-Thought (H-CoT), a universal and transferable attack method that leverages the model's own displayed intermediate reasoning to jailbreak its safety reasoning mechanism. Under H-CoT, refusal rates sharply decline-dropping from 98% to below 2%-and, in some instances, even transform initially cautious tones into ones that are willing to provide harmful content. We hope these findings underscore the urgent need for more robust safety mechanisms to preserve the benefits of advanced reasoning capabilities without compromising ethical standards.
From Isolated Conversations to Hierarchical Schemas: Dynamic Tree Memory Representation for LLMs
Oct 17, 2024



Abstract:Recent advancements in large language models have significantly improved their context windows, yet challenges in effective long-term memory management remain. We introduce MemTree, an algorithm that leverages a dynamic, tree-structured memory representation to optimize the organization, retrieval, and integration of information, akin to human cognitive schemas. MemTree organizes memory hierarchically, with each node encapsulating aggregated textual content, corresponding semantic embeddings, and varying abstraction levels across the tree's depths. Our algorithm dynamically adapts this memory structure by computing and comparing semantic embeddings of new and existing information to enrich the model's context-awareness. This approach allows MemTree to handle complex reasoning and extended interactions more effectively than traditional memory augmentation methods, which often rely on flat lookup tables. Evaluations on benchmarks for multi-turn dialogue understanding and document question answering show that MemTree significantly enhances performance in scenarios that demand structured memory management.
LLM Unlearning via Loss Adjustment with Only Forget Data
Oct 14, 2024



Abstract:Unlearning in Large Language Models (LLMs) is essential for ensuring ethical and responsible AI use, especially in addressing privacy leak, bias, safety, and evolving regulations. Existing approaches to LLM unlearning often rely on retain data or a reference LLM, yet they struggle to adequately balance unlearning performance with overall model utility. This challenge arises because leveraging explicit retain data or implicit knowledge of retain data from a reference LLM to fine-tune the model tends to blur the boundaries between the forgotten and retain data, as different queries often elicit similar responses. In this work, we propose eliminating the need to retain data or the reference LLM for response calibration in LLM unlearning. Recognizing that directly applying gradient ascent on the forget data often leads to optimization instability and poor performance, our method guides the LLM on what not to respond to, and importantly, how to respond, based on the forget data. Hence, we introduce Forget data only Loss AjustmenT (FLAT), a "flat" loss adjustment approach which addresses these issues by maximizing f-divergence between the available template answer and the forget answer only w.r.t. the forget data. The variational form of the defined f-divergence theoretically provides a way of loss adjustment by assigning different importance weights for the learning w.r.t. template responses and the forgetting of responses subject to unlearning. Empirical results demonstrate that our approach not only achieves superior unlearning performance compared to existing methods but also minimizes the impact on the model's retained capabilities, ensuring high utility across diverse tasks, including copyrighted content unlearning on Harry Potter dataset and MUSE Benchmark, and entity unlearning on the TOFU dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge