Pengtao Xie
University of California San Diego
Pathological Visual Question Answering
Oct 06, 2020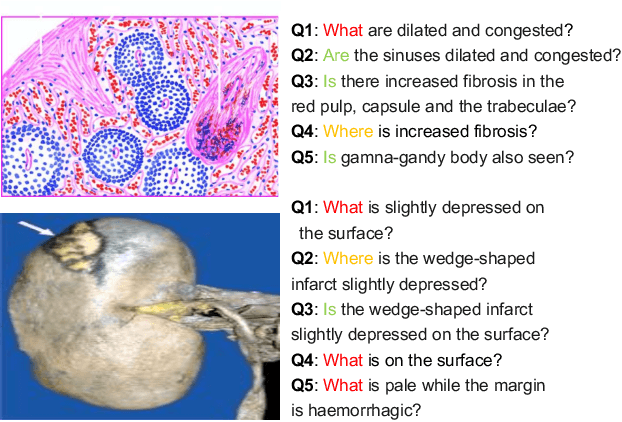
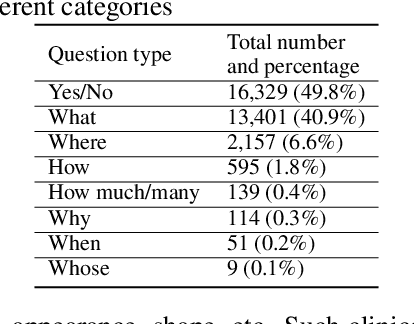
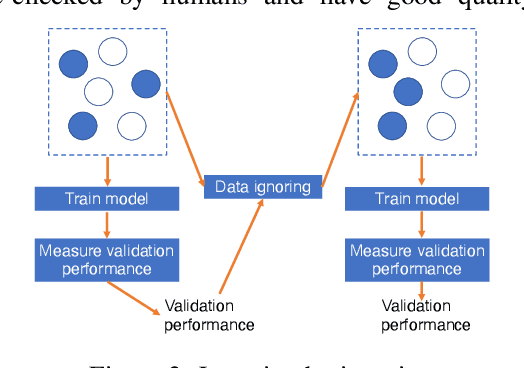
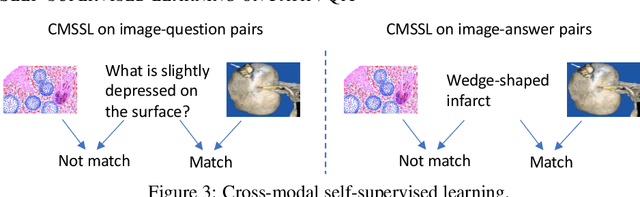
Abstract:Is it possible to develop an "AI Pathologist" to pass the board-certified examination of the American Board of Pathology (ABP)? To build such a system, three challenges need to be addressed. First, we need to create a visual question answering (VQA) dataset where the AI agent is presented with a pathology image together with a question and is asked to give the correct answer. Due to privacy concerns, pathology images are usually not publicly available. Besides, only well-trained pathologists can understand pathology images, but they barely have time to help create datasets for AI research. The second challenge is: since it is difficult to hire highly experienced pathologists to create pathology visual questions and answers, the resulting pathology VQA dataset may contain errors. Training pathology VQA models using these noisy or even erroneous data will lead to problematic models that cannot generalize well on unseen images. The third challenge is: the medical concepts and knowledge covered in pathology question-answer (QA) pairs are very diverse while the number of QA pairs available for modeling training is limited. How to learn effective representations of diverse medical concepts based on limited data is technically demanding. In this paper, we aim to address these three challenges. To our best knowledge, our work represents the first one addressing the pathology VQA problem. To deal with the issue that a publicly available pathology VQA dataset is lacking, we create PathVQA dataset. To address the second challenge, we propose a learning-by-ignoring approach. To address the third challenge, we propose to use cross-modal self-supervised learning. We perform experiments on our created PathVQA dataset and the results demonstrate the effectiveness of our proposed learning-by-ignoring method and cross-modal self-supervised learning methods.
TreeGAN: Incorporating Class Hierarchy into Image Generation
Sep 16, 2020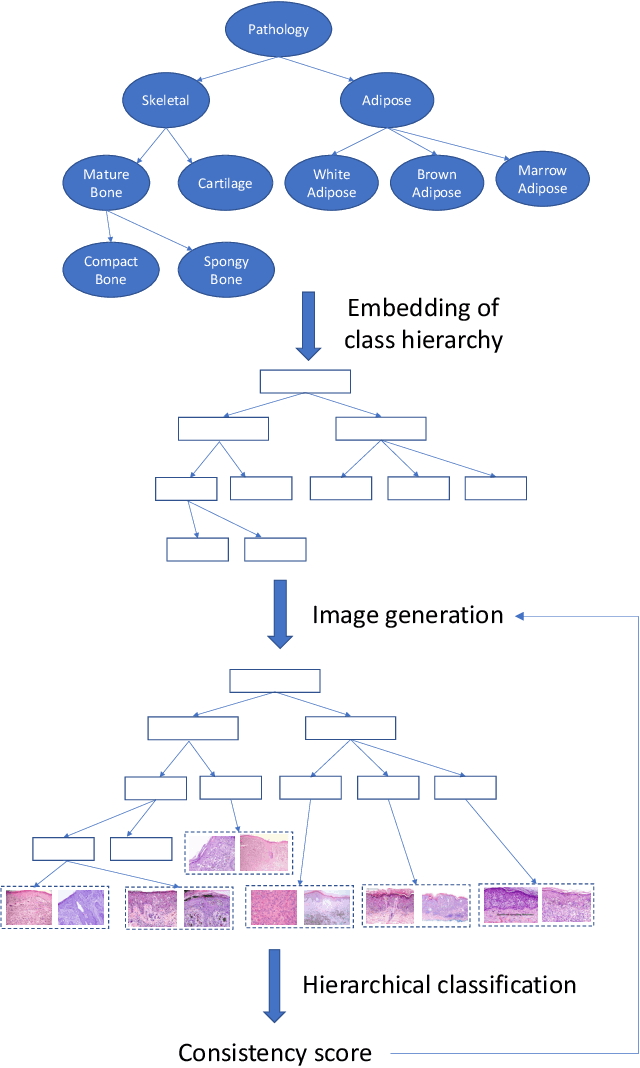
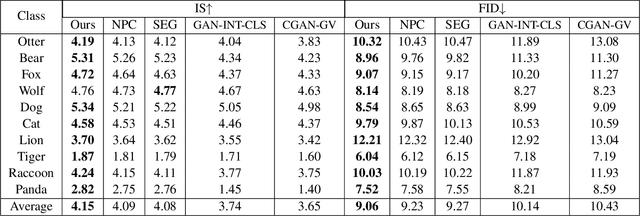
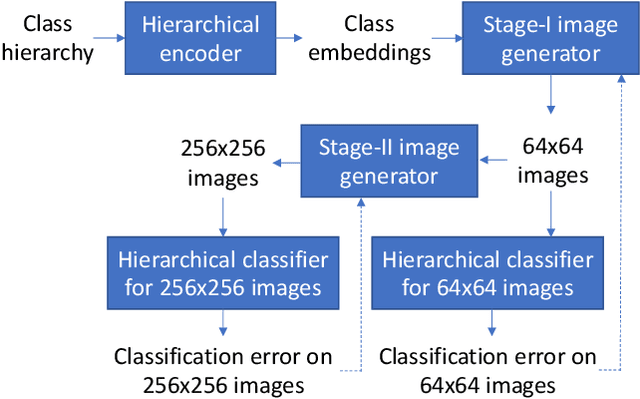
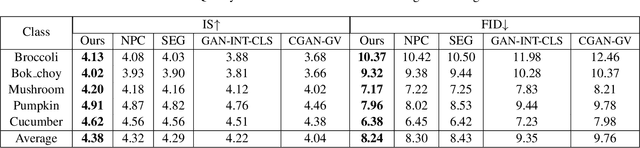
Abstract:Conditional image generation (CIG) is a widely studied problem in computer vision and machine learning. Given a class, CIG takes the name of this class as input and generates a set of images that belong to this class. In existing CIG works, for different classes, their corresponding images are generated independently, without considering the relationship among classes. In real-world applications, the classes are organized into a hierarchy and their hierarchical relationships are informative for generating high-fidelity images. In this paper, we aim to leverage the class hierarchy for conditional image generation. We propose two ways of incorporating class hierarchy: prior control and post constraint. In prior control, we first encode the class hierarchy, then feed it as a prior into the conditional generator to generate images. In post constraint, after the images are generated, we measure their consistency with the class hierarchy and use the consistency score to guide the training of the generator. Based on these two ideas, we propose a TreeGAN model which consists of three modules: (1) a class hierarchy encoder (CHE) which takes the hierarchical structure of classes and their textual names as inputs and learns an embedding for each class; the embedding captures the hierarchical relationship among classes; (2) a conditional image generator (CIG) which takes the CHE-generated embedding of a class as input and generates a set of images belonging to this class; (3) a consistency checker which performs hierarchical classification on the generated images and checks whether the generated images are compatible with the class hierarchy; the consistency score is used to guide the CIG to generate hierarchy-compatible images. Experiments on various datasets demonstrate the effectiveness of our method.
Contrastive Self-supervised Learning for Graph Classification
Sep 13, 2020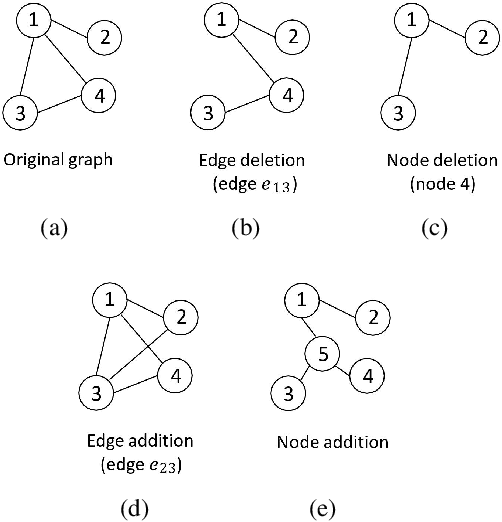
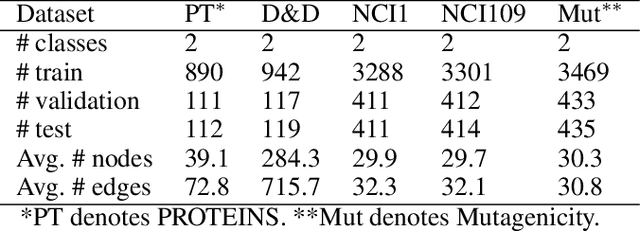
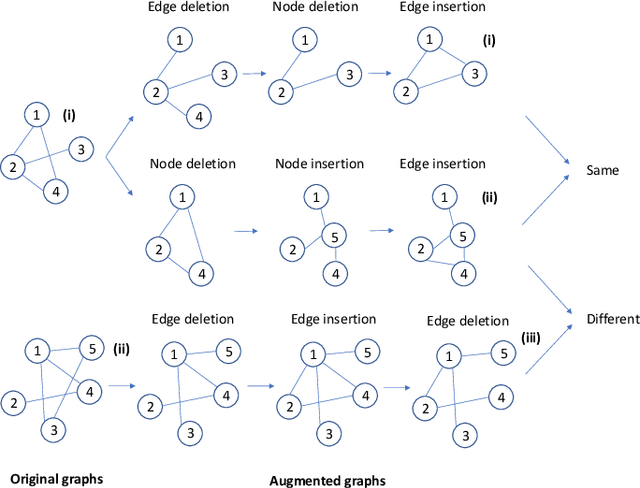
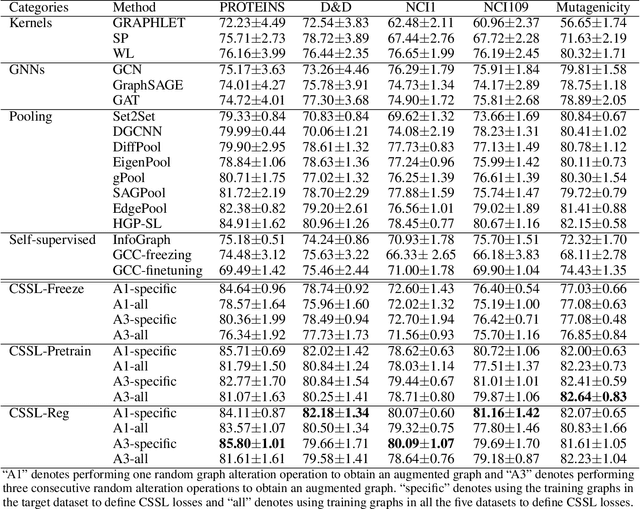
Abstract:Graph classification is a widely studied problem and has broad applications. In many real-world problems, the number of labeled graphs available for training classification models is limited, which renders these models prone to overfitting. To address this problem, we propose two approaches based on contrastive self-supervised learning (CSSL) to alleviate overfitting. In the first approach, we use CSSL to pretrain graph encoders on widely-available unlabeled graphs without relying on human-provided labels, then finetune the pretrained encoders on labeled graphs. In the second approach, we develop a regularizer based on CSSL, and solve the supervised classification task and the unsupervised CSSL task simultaneously. To perform CSSL on graphs, given a collection of original graphs, we perform data augmentation to create augmented graphs out of the original graphs. An augmented graph is created by consecutively applying a sequence of graph alteration operations. A contrastive loss is defined to learn graph encoders by judging whether two augmented graphs are from the same original graph. Experiments on various graph classification datasets demonstrate the effectiveness of our proposed methods.
Differentially-private Federated Neural Architecture Search
Jun 22, 2020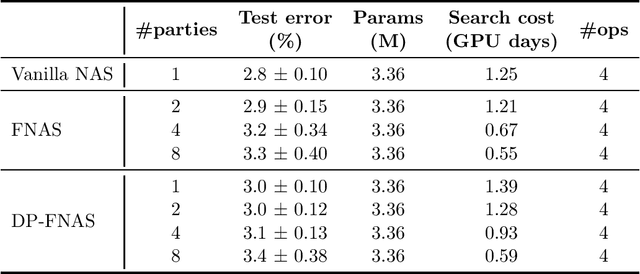
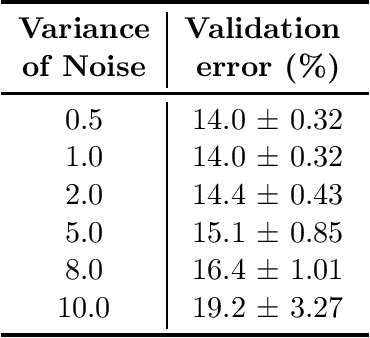
Abstract:Neural architecture search, which aims to automatically search for architectures (e.g., convolution, max pooling) of neural networks that maximize validation performance, has achieved remarkable progress recently. In many application scenarios, several parties would like to collaboratively search for a shared neural architecture by leveraging data from all parties. However, due to privacy concerns, no party wants its data to be seen by other parties. To address this problem, we propose federated neural architecture search (FNAS), where different parties collectively search for a differentiable architecture by exchanging gradients of architecture variables without exposing their data to other parties. To further preserve privacy, we study differentially-private FNAS (DP-FNAS), which adds random noise to the gradients of architecture variables. We provide theoretical guarantees of DP-FNAS in achieving differential privacy. Experiments show that DP-FNAS can search highly-performant neural architectures while protecting the privacy of individual parties. The code is available at https://github.com/UCSD-AI4H/DP-FNAS
Transfer Learning or Self-supervised Learning? A Tale of Two Pretraining Paradigms
Jun 19, 2020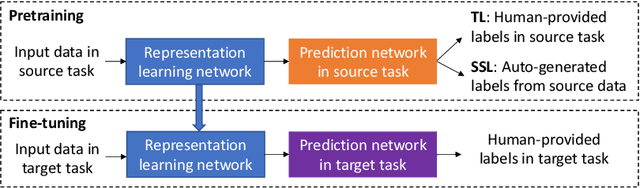
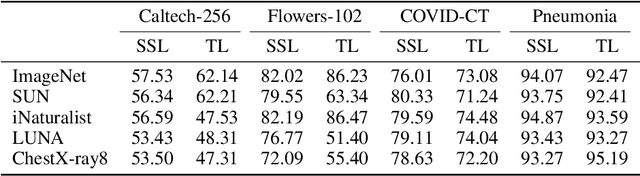
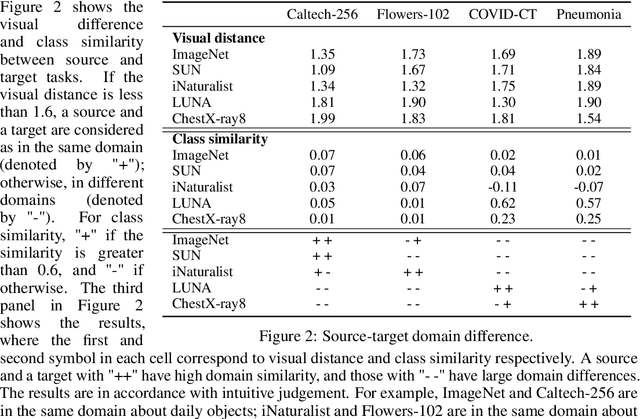
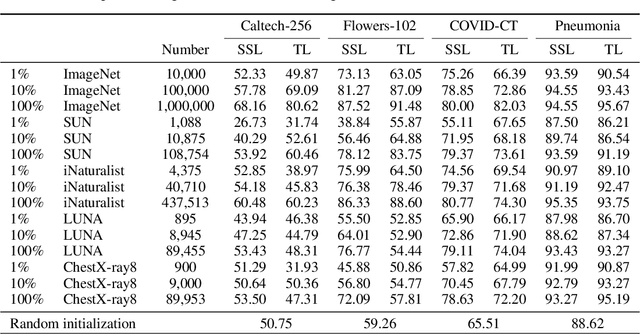
Abstract:Pretraining has become a standard technique in computer vision and natural language processing, which usually helps to improve performance substantially. Previously, the most dominant pretraining method is transfer learning (TL), which uses labeled data to learn a good representation network. Recently, a new pretraining approach -- self-supervised learning (SSL) -- has demonstrated promising results on a wide range of applications. SSL does not require annotated labels. It is purely conducted on input data by solving auxiliary tasks defined on the input data examples. The current reported results show that in certain applications, SSL outperforms TL and the other way around in other applications. There has not been a clear understanding on what properties of data and tasks render one approach outperforms the other. Without an informed guideline, ML researchers have to try both methods to find out which one is better empirically. It is usually time-consuming to do so. In this work, we aim to address this problem. We perform a comprehensive comparative study between SSL and TL regarding which one works better under different properties of data and tasks, including domain difference between source and target tasks, the amount of pretraining data, class imbalance in source data, and usage of target data for additional pretraining, etc. The insights distilled from our comparative studies can help ML researchers decide which method to use based on the properties of their applications.
XRayGAN: Consistency-preserving Generation of X-ray Images from Radiology Reports
Jun 17, 2020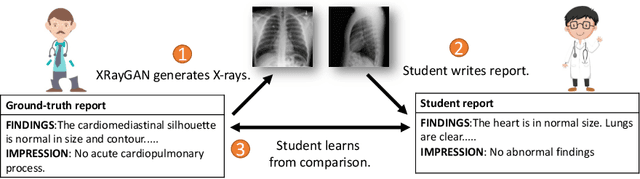
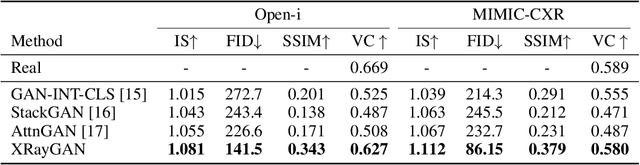
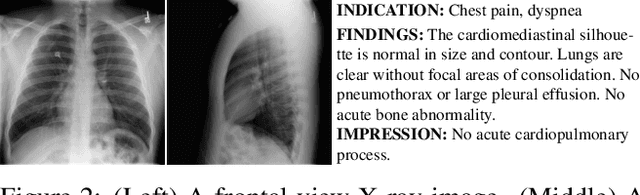
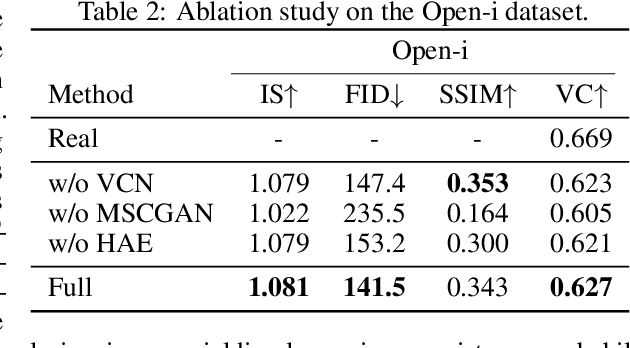
Abstract:To effectively train medical students to become qualified radiologists, a large number of X-ray images collected from patients with diverse medical conditions are needed. However, due to data privacy concerns, such images are typically difficult to obtain. To address this problem, we develop methods to generate view-consistent, high-fidelity, and high-resolution X-ray images from radiology reports to facilitate radiology training of medical students. This task is presented with several challenges. First, from a single report, images with different views (e.g., frontal, lateral) need to be generated. How to ensure consistency of these images (i.e., make sure they are about the same patient)? Second, X-ray images are required to have high resolution. Otherwise, many details of diseases would be lost. How to generate high-resolutions images? Third, radiology reports are long and have complicated structure. How to effectively understand their semantics to generate high-fidelity images that accurately reflect the contents of the reports? To address these three challenges, we propose an XRayGAN composed of three modules: (1) a view consistency network that maximizes the consistency between generated frontal-view and lateral-view images; (2) a multi-scale conditional GAN that progressively generates a cascade of images with increasing resolution; (3) a hierarchical attentional encoder that learns the latent semantics of a radiology report by capturing its hierarchical linguistic structure and various levels of clinical importance of words and sentences. Experiments on two radiology datasets demonstrate the effectiveness of our methods. To our best knowledge, this work represents the first one generating consistent and high-resolution X-ray images from radiology reports. The code is available at https://github.com/UCSD-AI4H/XRayGAN.
CERT: Contrastive Self-supervised Learning for Language Understanding
May 16, 2020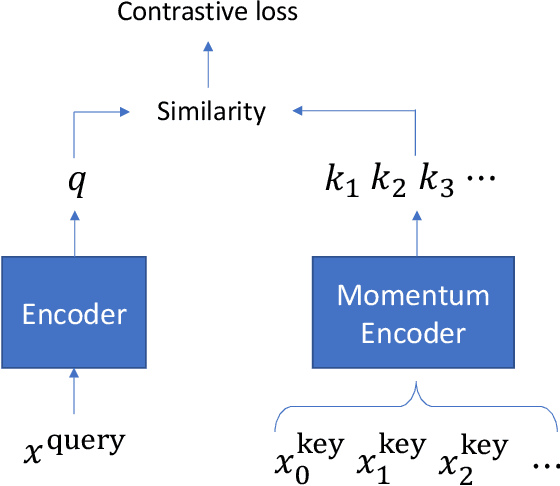
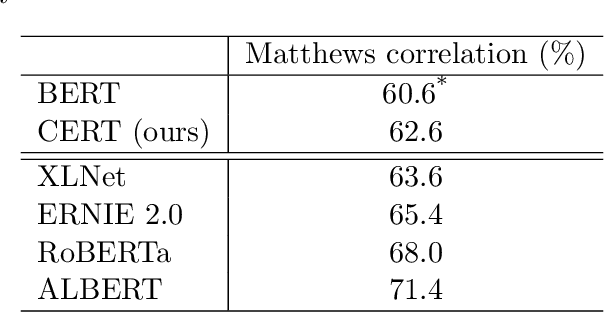
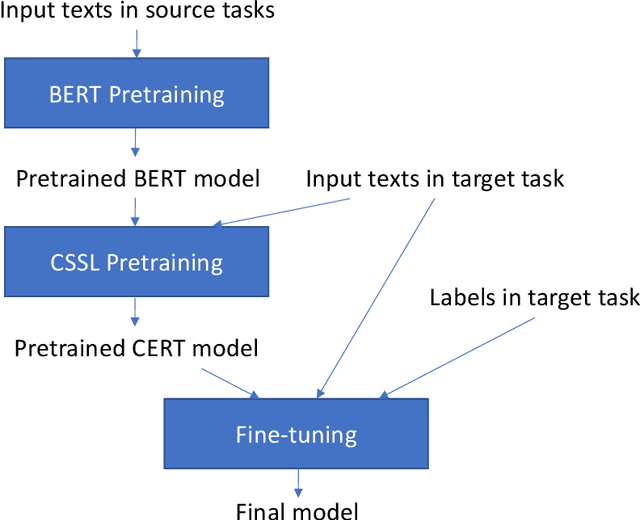
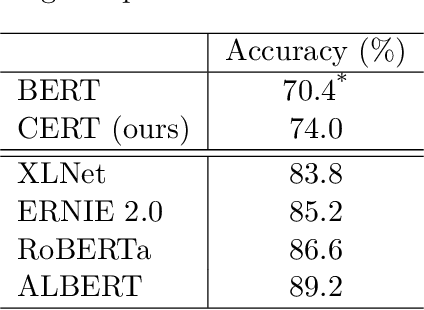
Abstract:Pretrained language models such as BERT, GPT have shown great effectiveness in language understanding. The auxiliary predictive tasks in existing pretraining approaches are mostly defined on tokens, thus may not be able to capture sentence-level semantics very well. To address this issue, we propose CERT: Contrastive self-supervised Encoder Representations from Transformers, which pretrains language representation models using contrastive self-supervised learning at the sentence level. CERT creates augmentations of original sentences using back-translation. Then it finetunes a pretrained language encoder (e.g., BERT) by predicting whether two augmented sentences originate from the same sentence. CERT is simple to use and can be flexibly plugged into any pretraining-finetuning NLP pipeline. We evaluate CERT on three language understanding tasks: CoLA, RTE, and QNLI. CERT outperforms BERT significantly.
On the Generation of Medical Dialogues for COVID-19
May 11, 2020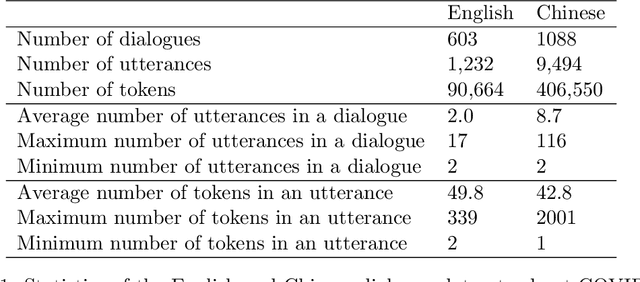

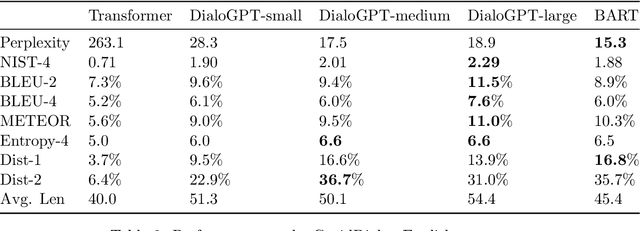

Abstract:Under the pandemic of COVID-19, people experiencing COVID19-related symptoms or exposed to risk factors have a pressing need to consult doctors. Due to hospital closure, a lot of consulting services have been moved online. Because of the shortage of medical professionals, many people cannot receive online consultations timely. To address this problem, we aim to develop a medical dialogue system that can provide COVID19-related consultations. We collected two dialogue datasets -CovidDialog- (in English and Chinese respectively) containing conversations between doctors and patients about COVID-19. On these two datasets, we train several dialogue generation models based on Transformer, GPT, and BERT-GPT. Since the two COVID-19 dialogue datasets are small in size, which bears high risk of overfitting, we leverage transfer learning to mitigate data deficiency. Specifically, we take the pretrained models of Transformer, GPT, and BERT-GPT on dialog datasets and other large-scale texts, then finetune them on our CovidDialog datasets. Experiments demonstrate that these approaches are promising in generating meaningful medical dialogues about COVID-19. But more advanced approaches are needed to build a fully useful dialogue system that can offer accurate COVID-related consultations. The data and code are available at https://github.com/UCSD-AI4H/COVID-Dialogue
MedDialog: A Large-scale Medical Dialogue Dataset
Apr 07, 2020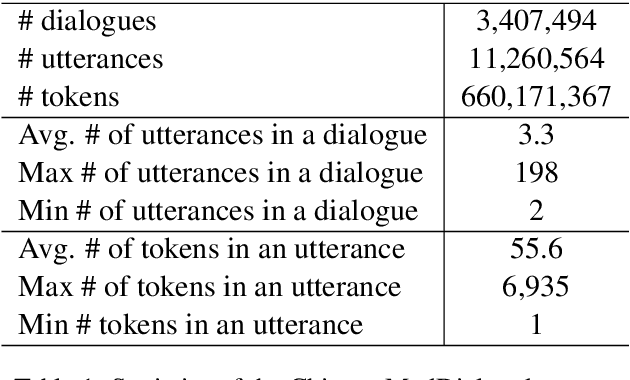

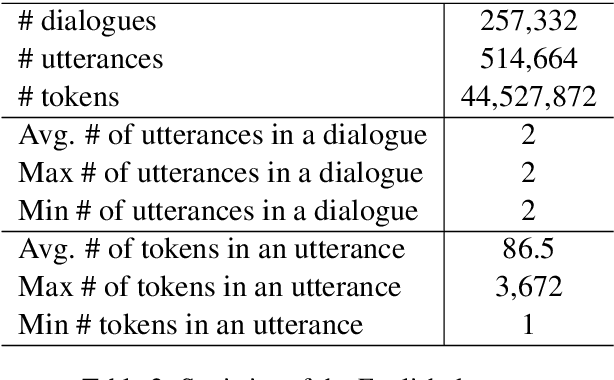
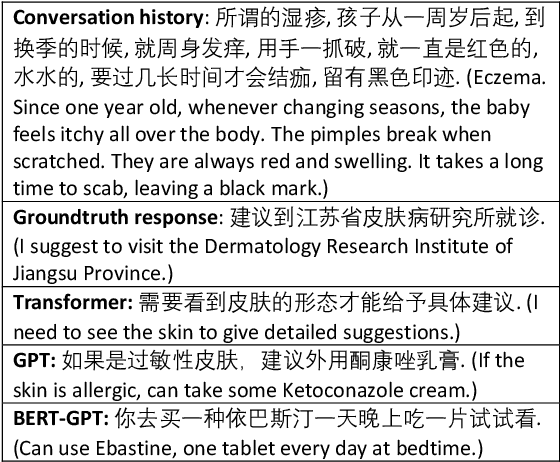
Abstract:Medical dialogue systems are promising in assisting in telemedicine to increase access to healthcare services, improve the quality of patient care, and reduce medical costs. To facilitate the research and development of medical dialogue systems, we build a large-scale medical dialogue dataset -- MedDialog -- that contains 1.1 million conversations between patients and doctors and 4 million utterances. To our best knowledge, MedDialog is the largest medical dialogue dataset to date. The dataset is available at https://github.com/UCSD-AI4H/Medical-Dialogue-System
Identifying Radiological Findings Related to COVID-19 from Medical Literature
Apr 04, 2020
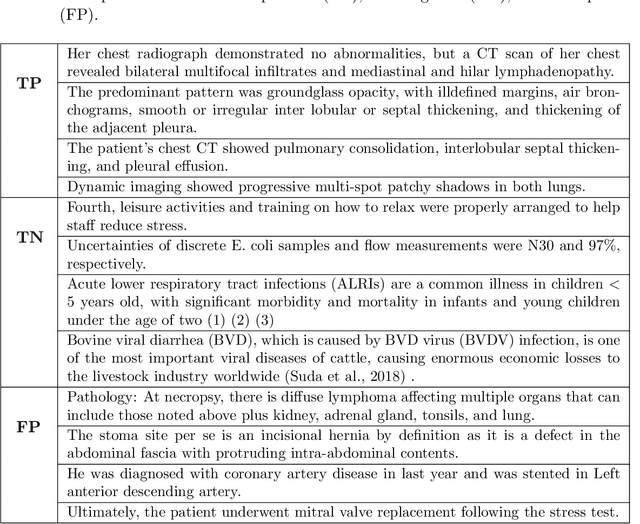
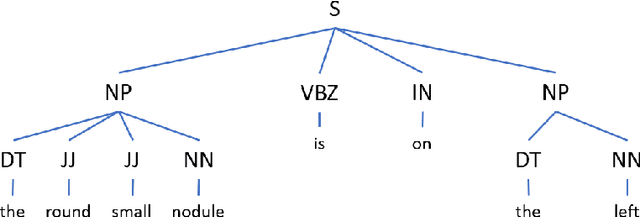

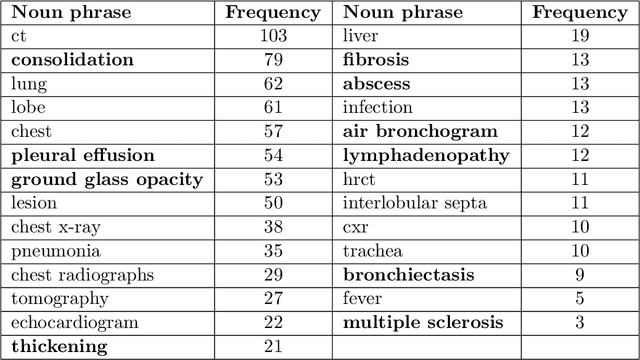
Abstract:Coronavirus disease 2019 (COVID-19) has infected more than one million individuals all over the world and caused more than 55,000 deaths, as of April 3 in 2020. Radiological findings are important sources of information in guiding the diagnosis and treatment of COVID-19. However, the existing studies on how radiological findings are correlated with COVID-19 are conducted separately by different hospitals, which may be inconsistent or even conflicting due to population bias. To address this problem, we develop natural language processing methods to analyze a large collection of COVID-19 literature containing study reports from hospitals all over the world, reconcile these results, and draw unbiased and universally-sensible conclusions about the correlation between radiological findings and COVID-19. We apply our method to the CORD-19 dataset and successfully extract a set of radiological findings that are closely tied to COVID-19.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge