Mattias P. Heinrich
Self-supervised Learning of Dense Hierarchical Representations for Medical Image Segmentation
Jan 12, 2024



Abstract:This paper demonstrates a self-supervised framework for learning voxel-wise coarse-to-fine representations tailored for dense downstream tasks. Our approach stems from the observation that existing methods for hierarchical representation learning tend to prioritize global features over local features due to inherent architectural bias. To address this challenge, we devise a training strategy that balances the contributions of features from multiple scales, ensuring that the learned representations capture both coarse and fine-grained details. Our strategy incorporates 3-fold improvements: (1) local data augmentations, (2) a hierarchically balanced architecture, and (3) a hybrid contrastive-restorative loss function. We evaluate our method on CT and MRI data and demonstrate that our new approach particularly beneficial for fine-tuning with limited annotated data and consistently outperforms the baseline counterpart in linear evaluation settings.
DG-TTA: Out-of-domain medical image segmentation through Domain Generalization and Test-Time Adaptation
Dec 22, 2023



Abstract:Applying pre-trained medical segmentation models on out-of-domain images often yields predictions of insufficient quality. Several strategies have been proposed to maintain model performance, such as finetuning or unsupervised- and source-free domain adaptation. These strategies set restrictive requirements for data availability. In this study, we propose to combine domain generalization and test-time adaptation to create a highly effective approach for reusing pre-trained models in unseen target domains. Domain-generalized pre-training on source data is used to obtain the best initial performance in the target domain. We introduce the MIND descriptor previously used in image registration tasks as a further technique to achieve generalization and present superior performance for small-scale datasets compared to existing approaches. At test-time, high-quality segmentation for every single unseen scan is ensured by optimizing the model weights for consistency given different image augmentations. That way, our method enables separate use of source and target data and thus removes current data availability barriers. Moreover, the presented method is highly modular as it does not require specific model architectures or prior knowledge of involved domains and labels. We demonstrate this by integrating it into the nnUNet, which is currently the most popular and accurate framework for medical image segmentation. We employ multiple datasets covering abdominal, cardiac, and lumbar spine scans and compose several out-of-domain scenarios in this study. We demonstrate that our method, combined with pre-trained whole-body CT models, can effectively segment MR images with high accuracy in all of the aforementioned scenarios. Open-source code can be found here: https://github.com/multimodallearning/DG-TTA
Shape Matters: Detecting Vertebral Fractures Using Differentiable Point-Based Shape Decoding
Dec 08, 2023



Abstract:Degenerative spinal pathologies are highly prevalent among the elderly population. Timely diagnosis of osteoporotic fractures and other degenerative deformities facilitates proactive measures to mitigate the risk of severe back pain and disability. In this study, we specifically explore the use of shape auto-encoders for vertebrae, taking advantage of advancements in automated multi-label segmentation and the availability of large datasets for unsupervised learning. Our shape auto-encoders are trained on a large set of vertebrae surface patches, leveraging the vast amount of available data for vertebra segmentation. This addresses the label scarcity problem faced when learning shape information of vertebrae from image intensities. Based on the learned shape features we train an MLP to detect vertebral body fractures. Using segmentation masks that were automatically generated using the TotalSegmentator, our proposed method achieves an AUC of 0.901 on the VerSe19 testset. This outperforms image-based and surface-based end-to-end trained models. Additionally, our results demonstrate that pre-training the models in an unsupervised manner enhances geometric methods like PointNet and DGCNN. Our findings emphasise the advantages of explicitly learning shape features for diagnosing osteoporotic vertebrae fractures. This approach improves the reliability of classification results and reduces the need for annotated labels. This study provides novel insights into the effectiveness of various encoder-decoder models for shape analysis of vertebrae and proposes a new decoder architecture: the point-based shape decoder.
A denoised Mean Teacher for domain adaptive point cloud registration
Jul 04, 2023



Abstract:Point cloud-based medical registration promises increased computational efficiency, robustness to intensity shifts, and anonymity preservation but is limited by the inefficacy of unsupervised learning with similarity metrics. Supervised training on synthetic deformations is an alternative but, in turn, suffers from the domain gap to the real domain. In this work, we aim to tackle this gap through domain adaptation. Self-training with the Mean Teacher is an established approach to this problem but is impaired by the inherent noise of the pseudo labels from the teacher. As a remedy, we present a denoised teacher-student paradigm for point cloud registration, comprising two complementary denoising strategies. First, we propose to filter pseudo labels based on the Chamfer distances of teacher and student registrations, thus preventing detrimental supervision by the teacher. Second, we make the teacher dynamically synthesize novel training pairs with noise-free labels by warping its moving inputs with the predicted deformations. Evaluation is performed for inhale-to-exhale registration of lung vessel trees on the public PVT dataset under two domain shifts. Our method surpasses the baseline Mean Teacher by 13.5/62.8%, consistently outperforms diverse competitors, and sets a new state-of-the-art accuracy (TRE=2.31mm). Code is available at https://github.com/multimodallearning/denoised_mt_pcd_reg.
Unsupervised 3D registration through optimization-guided cyclical self-training
Jun 29, 2023



Abstract:State-of-the-art deep learning-based registration methods employ three different learning strategies: supervised learning, which requires costly manual annotations, unsupervised learning, which heavily relies on hand-crafted similarity metrics designed by domain experts, or learning from synthetic data, which introduces a domain shift. To overcome the limitations of these strategies, we propose a novel self-supervised learning paradigm for unsupervised registration, relying on self-training. Our idea is based on two key insights. Feature-based differentiable optimizers 1) perform reasonable registration even from random features and 2) stabilize the training of the preceding feature extraction network on noisy labels. Consequently, we propose cyclical self-training, where pseudo labels are initialized as the displacement fields inferred from random features and cyclically updated based on more and more expressive features from the learning feature extractor, yielding a self-reinforcement effect. We evaluate the method for abdomen and lung registration, consistently surpassing metric-based supervision and outperforming diverse state-of-the-art competitors. Source code is available at https://github.com/multimodallearning/reg-cyclical-self-train.
Anatomy-guided domain adaptation for 3D in-bed human pose estimation
Nov 22, 2022Abstract:3D human pose estimation is a key component of clinical monitoring systems. The clinical applicability of deep pose estimation models, however, is limited by their poor generalization under domain shifts along with their need for sufficient labeled training data. As a remedy, we present a novel domain adaptation method, adapting a model from a labeled source to a shifted unlabeled target domain. Our method comprises two complementary adaptation strategies based on prior knowledge about human anatomy. First, we guide the learning process in the target domain by constraining predictions to the space of anatomically plausible poses. To this end, we embed the prior knowledge into an anatomical loss function that penalizes asymmetric limb lengths, implausible bone lengths, and implausible joint angles. Second, we propose to filter pseudo labels for self-training according to their anatomical plausibility and incorporate the concept into the Mean Teacher paradigm. We unify both strategies in a point cloud-based framework applicable to unsupervised and source-free domain adaptation. Evaluation is performed for in-bed pose estimation under two adaptation scenarios, using the public SLP dataset and a newly created dataset. Our method consistently outperforms various state-of-the-art domain adaptation methods, surpasses the baseline model by 31%/66%, and reduces the domain gap by 65%/82%. Source code is available at https://github.com/multimodallearning/da-3dhpe-anatomy.
Adapting the Mean Teacher for keypoint-based lung registration under geometric domain shifts
Jul 01, 2022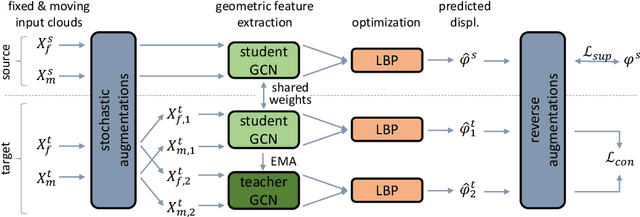
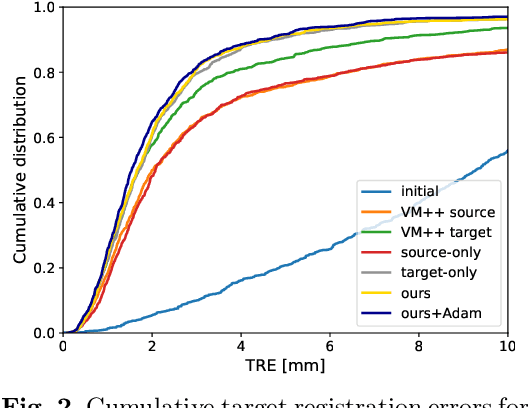
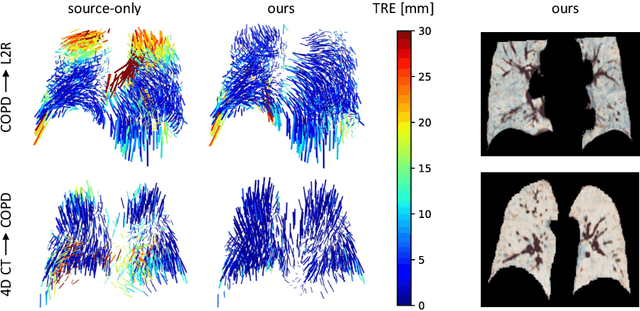
Abstract:Recent deep learning-based methods for medical image registration achieve results that are competitive with conventional optimization algorithms at reduced run times. However, deep neural networks generally require plenty of labeled training data and are vulnerable to domain shifts between training and test data. While typical intensity shifts can be mitigated by keypoint-based registration, these methods still suffer from geometric domain shifts, for instance, due to different fields of view. As a remedy, in this work, we present a novel approach to geometric domain adaptation for image registration, adapting a model from a labeled source to an unlabeled target domain. We build on a keypoint-based registration model, combining graph convolutions for geometric feature learning with loopy belief optimization, and propose to reduce the domain shift through self-ensembling. To this end, we embed the model into the Mean Teacher paradigm. We extend the Mean Teacher to this context by 1) adapting the stochastic augmentation scheme and 2) combining learned feature extraction with differentiable optimization. This enables us to guide the learning process in the unlabeled target domain by enforcing consistent predictions of the learning student and the temporally averaged teacher model. We evaluate the method for exhale-to-inhale lung CT registration under two challenging adaptation scenarios (DIR-Lab 4D CT to COPD, COPD to Learn2Reg). Our method consistently improves on the baseline model by 50%/47% while even matching the accuracy of models trained on target data. Source code is available at https://github.com/multimodallearning/registration-da-mean-teacher.
Voxelmorph++ Going beyond the cranial vault with keypoint supervision and multi-channel instance optimisation
Feb 28, 2022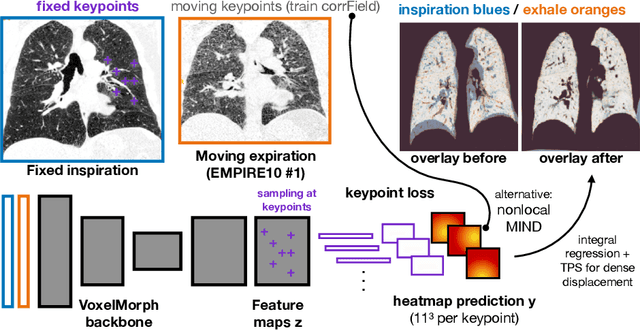
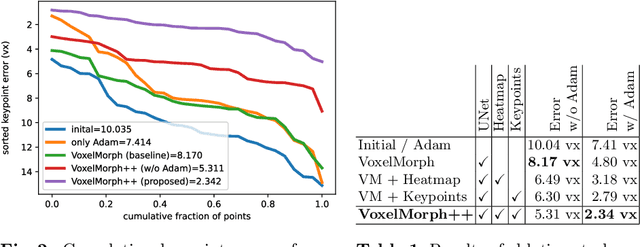
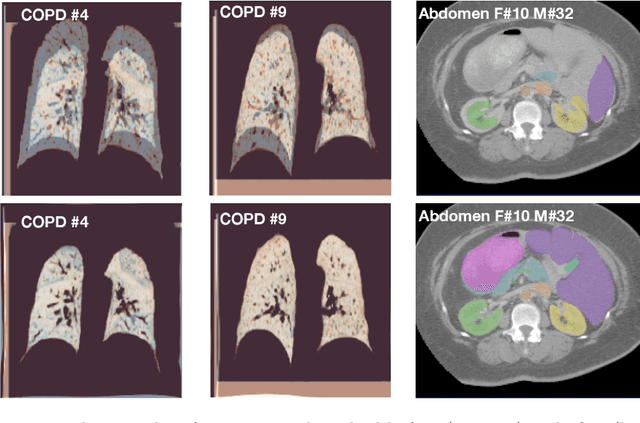
Abstract:The majority of current research in deep learning based image registration addresses inter-patient brain registration with moderate deformation magnitudes. The recent Learn2Reg medical registration benchmark has demonstrated that single-scale U-Net architectures, such as VoxelMorph that directly employ a spatial transformer loss, often do not generalise well beyond the cranial vault and fall short of state-of-the-art performance for abdominal or intra-patient lung registration. Here, we propose two straightforward steps that greatly reduce this gap in accuracy. First, we employ keypoint self-supervision with a novel network head that predicts a discretised heatmap and robustly reduces large deformations for better robustness. Second, we replace multiple learned fine-tuning steps by a single instance optimisation with hand-crafted features and the Adam optimiser. Different to other related work, including FlowNet or PDD-Net, our approach does not require a fully discretised architecture with correlation layer. Our ablation study demonstrates the importance of keypoints in both self-supervised and unsupervised (using only a MIND metric) settings. On a multi-centric inspiration-exhale lung CT dataset, including very challenging COPD scans, our method outperforms VoxelMorph by improving nonlinear alignment by 77% compared to 19% - reaching target registration errors of 2 mm that outperform all but one learning methods published to date. Extending the method to semantic features sets new stat-of-the-art performance on inter-subject abdominal CT registration.
CrossMoDA 2021 challenge: Benchmark of Cross-Modality Domain Adaptation techniques for Vestibular Schwnannoma and Cochlea Segmentation
Jan 08, 2022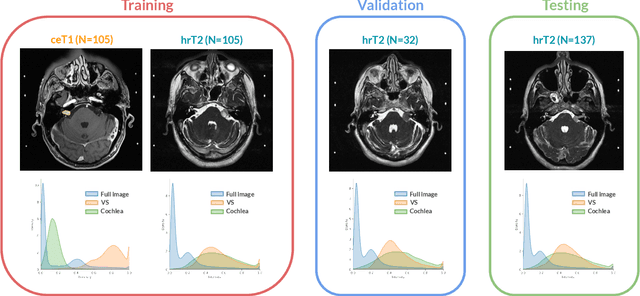
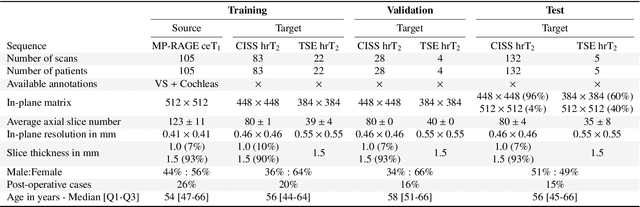
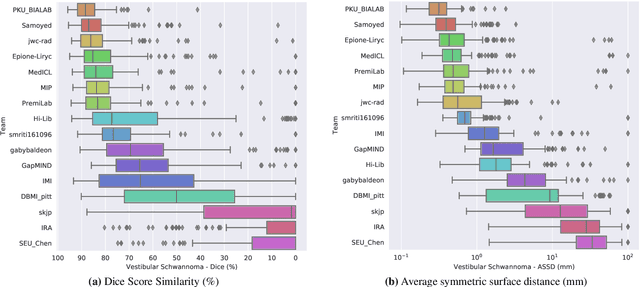
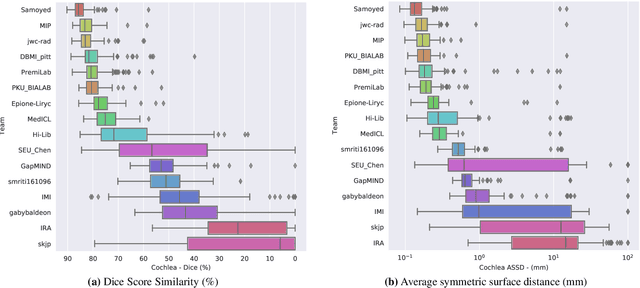
Abstract:Domain Adaptation (DA) has recently raised strong interests in the medical imaging community. While a large variety of DA techniques has been proposed for image segmentation, most of these techniques have been validated either on private datasets or on small publicly available datasets. Moreover, these datasets mostly addressed single-class problems. To tackle these limitations, the Cross-Modality Domain Adaptation (crossMoDA) challenge was organised in conjunction with the 24th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2021). CrossMoDA is the first large and multi-class benchmark for unsupervised cross-modality DA. The challenge's goal is to segment two key brain structures involved in the follow-up and treatment planning of vestibular schwannoma (VS): the VS and the cochleas. Currently, the diagnosis and surveillance in patients with VS are performed using contrast-enhanced T1 (ceT1) MRI. However, there is growing interest in using non-contrast sequences such as high-resolution T2 (hrT2) MRI. Therefore, we created an unsupervised cross-modality segmentation benchmark. The training set provides annotated ceT1 (N=105) and unpaired non-annotated hrT2 (N=105). The aim was to automatically perform unilateral VS and bilateral cochlea segmentation on hrT2 as provided in the testing set (N=137). A total of 16 teams submitted their algorithm for the evaluation phase. The level of performance reached by the top-performing teams is strikingly high (best median Dice - VS:88.4%; Cochleas:85.7%) and close to full supervision (median Dice - VS:92.5%; Cochleas:87.7%). All top-performing methods made use of an image-to-image translation approach to transform the source-domain images into pseudo-target-domain images. A segmentation network was then trained using these generated images and the manual annotations provided for the source image.
Learn2Reg: comprehensive multi-task medical image registration challenge, dataset and evaluation in the era of deep learning
Dec 23, 2021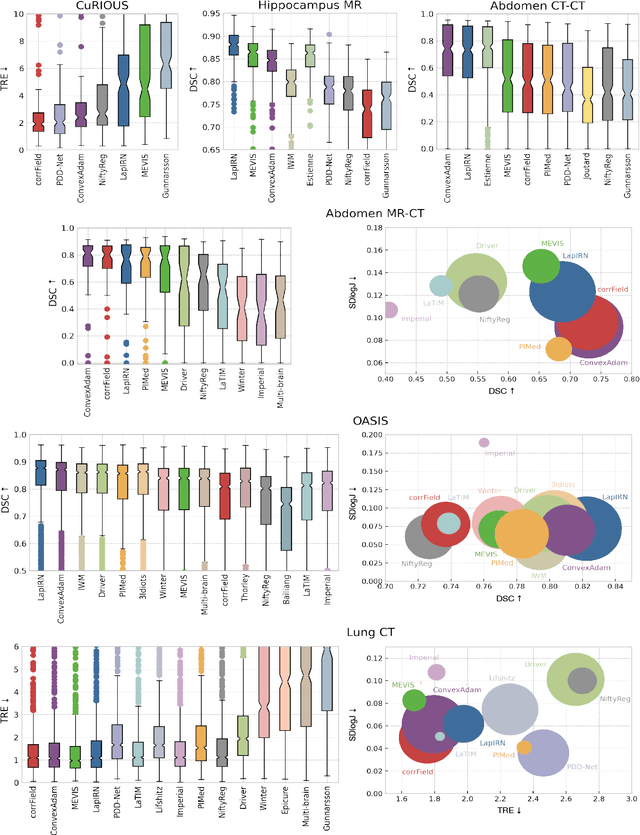
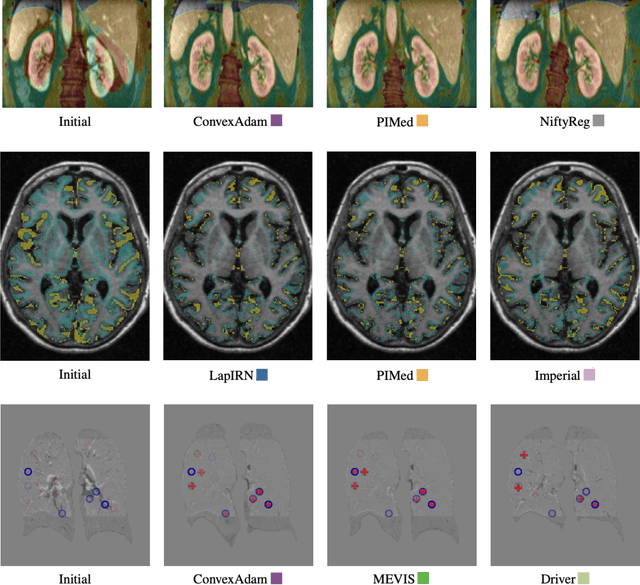
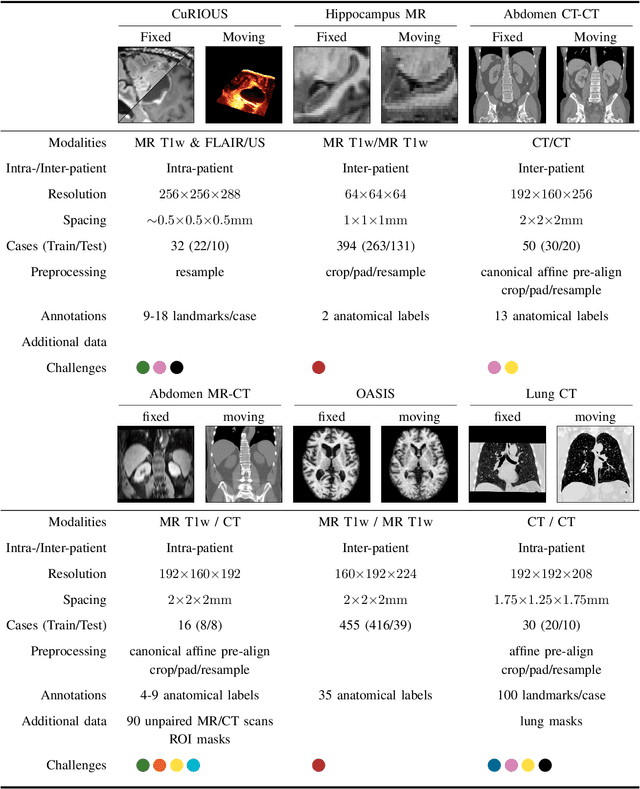
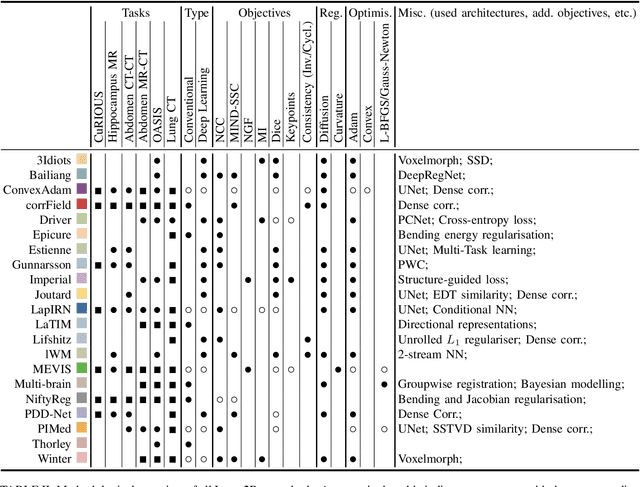
Abstract:Image registration is a fundamental medical image analysis task, and a wide variety of approaches have been proposed. However, only a few studies have comprehensively compared medical image registration approaches on a wide range of clinically relevant tasks, in part because of the lack of availability of such diverse data. This limits the development of registration methods, the adoption of research advances into practice, and a fair benchmark across competing approaches. The Learn2Reg challenge addresses these limitations by providing a multi-task medical image registration benchmark for comprehensive characterisation of deformable registration algorithms. A continuous evaluation will be possible at https://learn2reg.grand-challenge.org. Learn2Reg covers a wide range of anatomies (brain, abdomen, and thorax), modalities (ultrasound, CT, MR), availability of annotations, as well as intra- and inter-patient registration evaluation. We established an easily accessible framework for training and validation of 3D registration methods, which enabled the compilation of results of over 65 individual method submissions from more than 20 unique teams. We used a complementary set of metrics, including robustness, accuracy, plausibility, and runtime, enabling unique insight into the current state-of-the-art of medical image registration. This paper describes datasets, tasks, evaluation methods and results of the challenge, and the results of further analysis of transferability to new datasets, the importance of label supervision, and resulting bias.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge