Luyao Chen
Language Hierarchization Provides the Optimal Solution to Human Working Memory Limits
Jan 06, 2026Abstract:Language is a uniquely human trait, conveying information efficiently by organizing word sequences in sentences into hierarchical structures. A central question persists: Why is human language hierarchical? In this study, we show that hierarchization optimally solves the challenge of our limited working memory capacity. We established a likelihood function that quantifies how well the average number of units according to the language processing mechanisms aligns with human working memory capacity (WMC) in a direct fashion. The maximum likelihood estimate (MLE) of this function, tehta_MLE, turns out to be the mean of units. Through computational simulations of symbol sequences and validation analyses of natural language sentences, we uncover that compared to linear processing, hierarchical processing far surpasses it in constraining the tehta_MLE values under the human WMC limit, along with the increase of sequence/sentence length successfully. It also shows a converging pattern related to children's WMC development. These results suggest that constructing hierarchical structures optimizes the processing efficiency of sequential language input while staying within memory constraints, genuinely explaining the universal hierarchical nature of human language.
Ascend HiFloat8 Format for Deep Learning
Sep 26, 2024



Abstract:This preliminary white paper proposes a novel 8-bit floating-point data format HiFloat8 (abbreviated as HiF8) for deep learning. HiF8 features tapered precision. For normal value encoding, it provides 7 exponent values with 3-bit mantissa, 8 exponent values with 2-bit mantissa, and 16 exponent values with 1-bit mantissa. For denormal value encoding, it extends the dynamic range by 7 extra powers of 2, from 31 to 38 binades (notice that FP16 covers 40 binades). Meanwhile, HiF8 encodes all the special values except that positive zero and negative zero are represented by only one bit-pattern. Thanks to the better balance between precision and dynamic range, HiF8 can be simultaneously used in both forward and backward passes of AI training. In this paper, we will describe the definition and rounding methods of HiF8, as well as the tentative training and inference solutions. To demonstrate the efficacy of HiF8, massive simulation results on various neural networks, including traditional neural networks and large language models (LLMs), will also be presented.
Evaluating Large Language Models on Spatial Tasks: A Multi-Task Benchmarking Study
Aug 26, 2024



Abstract:The advent of large language models such as ChatGPT, Gemini, and others has underscored the importance of evaluating their diverse capabilities, ranging from natural language understanding to code generation. However, their performance on spatial tasks has not been comprehensively assessed. This study addresses this gap by introducing a novel multi-task spatial evaluation dataset, designed to systematically explore and compare the performance of several advanced models on spatial tasks. The dataset encompasses twelve distinct task types, including spatial understanding and path planning, each with verified, accurate answers. We evaluated multiple models, including OpenAI's gpt-3.5-turbo, gpt-4o, and ZhipuAI's glm-4, through a two-phase testing approach. Initially, we conducted zero-shot testing, followed by categorizing the dataset by difficulty and performing prompt tuning tests. Results indicate that gpt-4o achieved the highest overall accuracy in the first phase, with an average of 71.3%. Although moonshot-v1-8k slightly underperformed overall, it surpassed gpt-4o in place name recognition tasks. The study also highlights the impact of prompt strategies on model performance in specific tasks. For example, the Chain-of-Thought (COT) strategy increased gpt-4o's accuracy in path planning from 12.4% to 87.5%, while a one-shot strategy enhanced moonshot-v1-8k's accuracy in mapping tasks from 10.1% to 76.3%.
Dental CLAIRES: Contrastive LAnguage Image REtrieval Search for Dental Research
Jun 27, 2023



Abstract:Learning about diagnostic features and related clinical information from dental radiographs is important for dental research. However, the lack of expert-annotated data and convenient search tools poses challenges. Our primary objective is to design a search tool that uses a user's query for oral-related research. The proposed framework, Contrastive LAnguage Image REtrieval Search for dental research, Dental CLAIRES, utilizes periapical radiographs and associated clinical details such as periodontal diagnosis, demographic information to retrieve the best-matched images based on the text query. We applied a contrastive representation learning method to find images described by the user's text by maximizing the similarity score of positive pairs (true pairs) and minimizing the score of negative pairs (random pairs). Our model achieved a hit@3 ratio of 96% and a Mean Reciprocal Rank (MRR) of 0.82. We also designed a graphical user interface that allows researchers to verify the model's performance with interactions.
* 10 pages, 7 figures, 4 tables
An End-to-end Entangled Segmentation and Classification Convolutional Neural Network for Periodontitis Stage Grading from Periapical Radiographic Images
Sep 27, 2021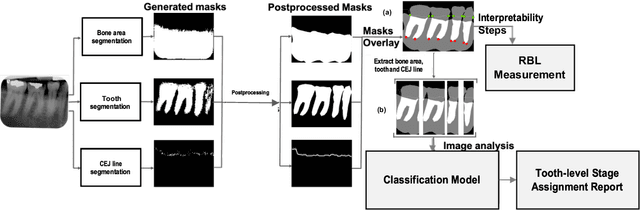
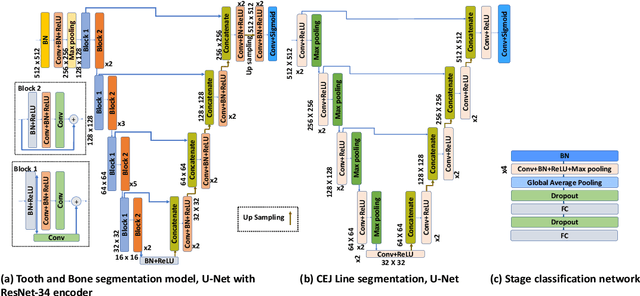
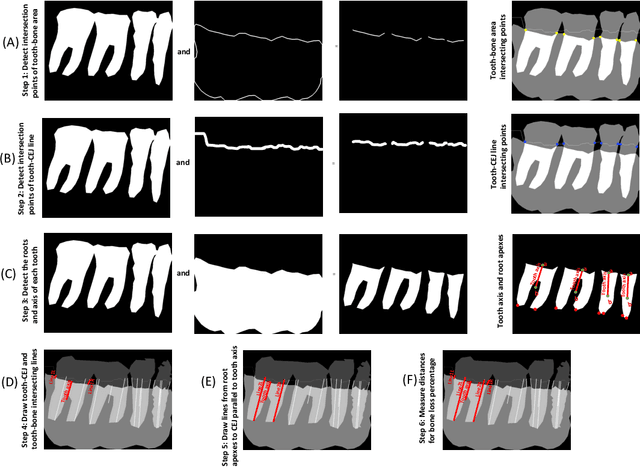
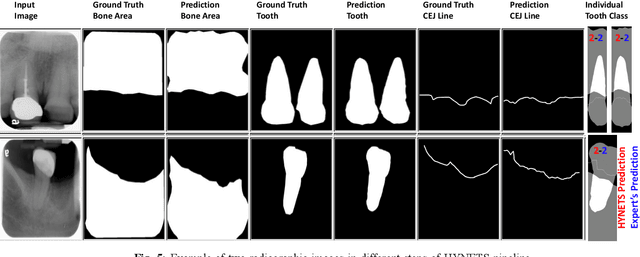
Abstract:Periodontitis is a biofilm-related chronic inflammatory disease characterized by gingivitis and bone loss in the teeth area. Approximately 61 million adults over 30 suffer from periodontitis (42.2%), with 7.8% having severe periodontitis in the United States. The measurement of radiographic bone loss (RBL) is necessary to make a correct periodontal diagnosis, especially if the comprehensive and longitudinal periodontal mapping is unavailable. However, doctors can interpret X-rays differently depending on their experience and knowledge. Computerized diagnosis support for doctors sheds light on making the diagnosis with high accuracy and consistency and drawing up an appropriate treatment plan for preventing or controlling periodontitis. We developed an end-to-end deep learning network HYNETS (Hybrid NETwork for pEriodoNTiTiS STagES from radiograpH) by integrating segmentation and classification tasks for grading periodontitis from periapical radiographic images. HYNETS leverages a multi-task learning strategy by combining a set of segmentation networks and a classification network to provide an end-to-end interpretable solution and highly accurate and consistent results. HYNETS achieved the average dice coefficient of 0.96 and 0.94 for the bone area and tooth segmentation and the average AUC of 0.97 for periodontitis stage assignment. Additionally, conventional image processing techniques provide RBL measurements and build transparency and trust in the model's prediction. HYNETS will potentially transform clinical diagnosis from a manual time-consuming, and error-prone task to an efficient and automated periodontitis stage assignment based on periapical radiographic images.
Heterogeneous Treatment Effect Estimation using machine learning for Healthcare application: tutorial and benchmark
Sep 27, 2021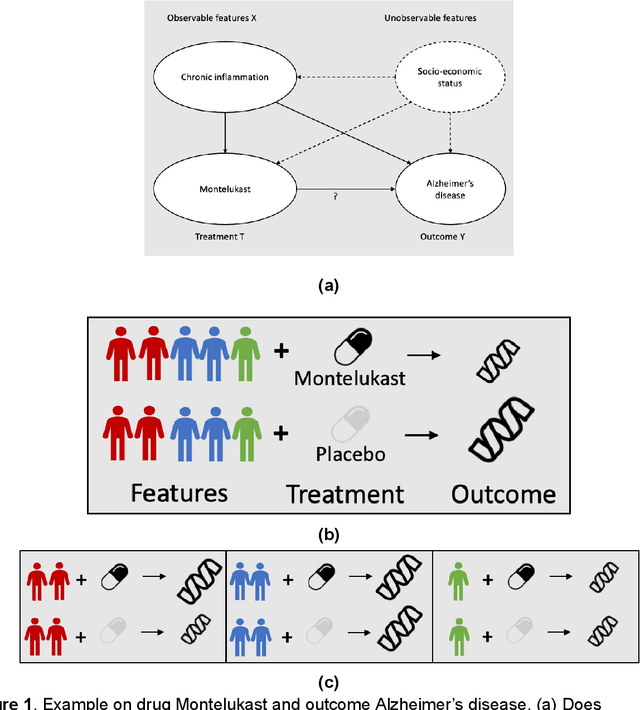
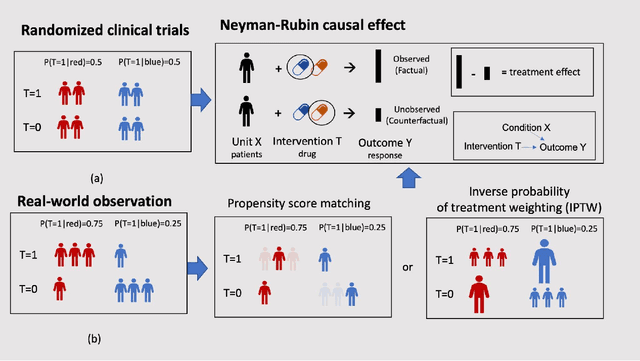
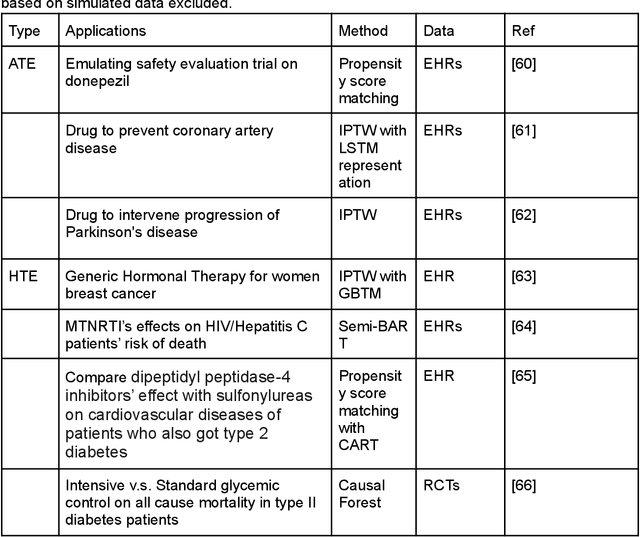
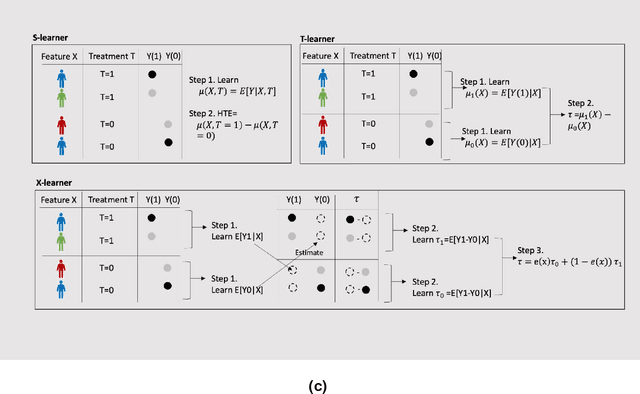
Abstract:Developing new drugs for target diseases is a time-consuming and expensive task, drug repurposing has become a popular topic in the drug development field. As much health claim data become available, many studies have been conducted on the data. The real-world data is noisy, sparse, and has many confounding factors. In addition, many studies have shown that drugs effects are heterogeneous among the population. Lots of advanced machine learning models about estimating heterogeneous treatment effects (HTE) have emerged in recent years, and have been applied to in econometrics and machine learning communities. These studies acknowledge medicine and drug development as the main application area, but there has been limited translational research from the HTE methodology to drug development. We aim to introduce the HTE methodology to the healthcare area and provide feasibility consideration when translating the methodology with benchmark experiments on healthcare administrative claim data. Also, we want to use benchmark experiments to show how to interpret and evaluate the model when it is applied to healthcare research. By introducing the recent HTE techniques to a broad readership in biomedical informatics communities, we expect to promote the wide adoption of causal inference using machine learning. We also expect to provide the feasibility of HTE for personalized drug effectiveness.
Drug Repurposing for COVID-19 using Graph Neural Network with Genetic, Mechanistic, and Epidemiological Validation
Sep 23, 2020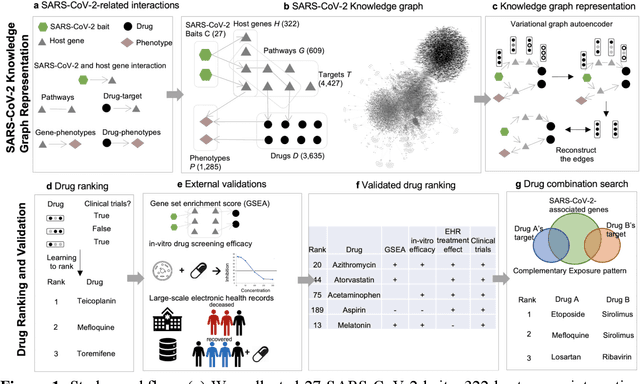
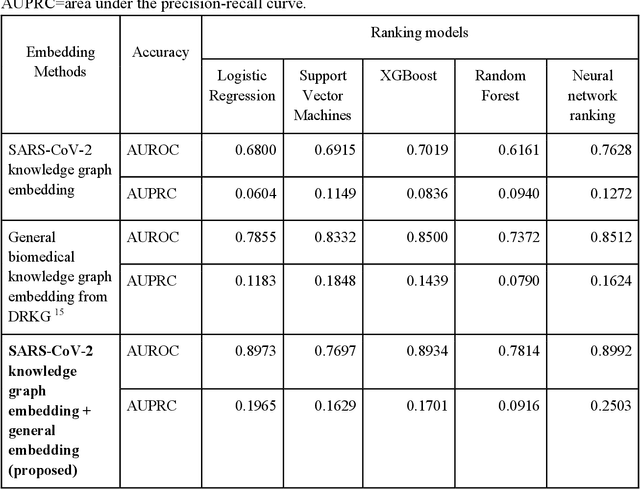
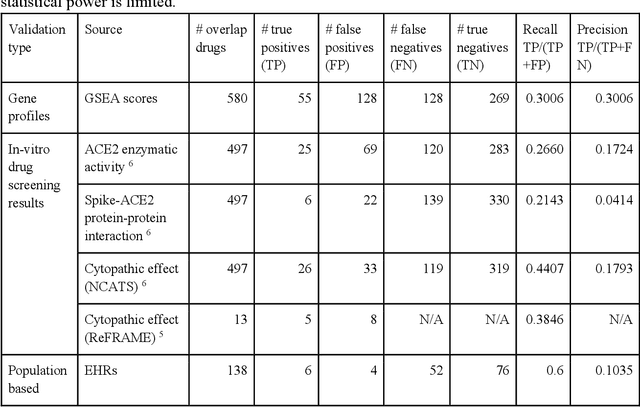
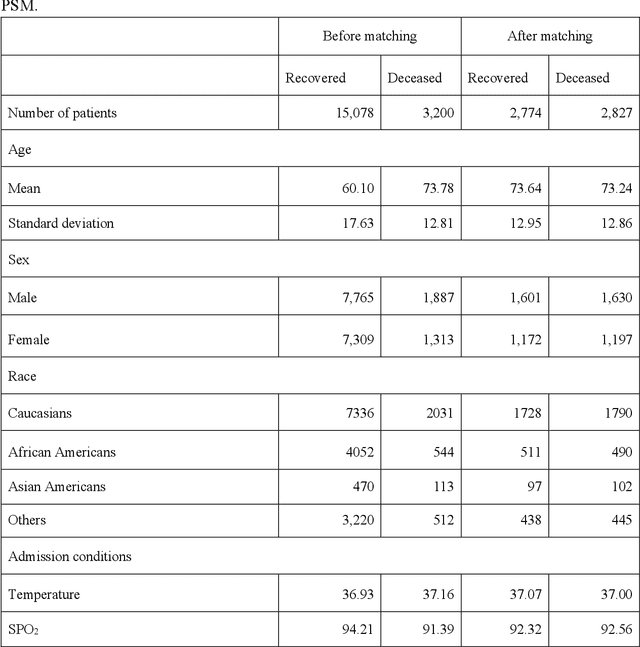
Abstract:Amid the pandemic of 2019 novel coronavirus disease (COVID-19) infected by SARS-CoV-2, a vast amount of drug research for prevention and treatment has been quickly conducted, but these efforts have been unsuccessful thus far. Our objective is to prioritize repurposable drugs using a drug repurposing pipeline that systematically integrates multiple SARS-CoV-2 and drug interactions, deep graph neural networks, and in-vitro/population-based validations. We first collected all the available drugs (n= 3,635) involved in COVID-19 patient treatment through CTDbase. We built a SARS-CoV-2 knowledge graph based on the interactions among virus baits, host genes, pathways, drugs, and phenotypes. A deep graph neural network approach was used to derive the candidate representation based on the biological interactions. We prioritized the candidate drugs using clinical trial history, and then validated them with their genetic profiles, in vitro experimental efficacy, and electronic health records. We highlight the top 22 drugs including Azithromycin, Atorvastatin, Aspirin, Acetaminophen, and Albuterol. We further pinpointed drug combinations that may synergistically target COVID-19. In summary, we demonstrated that the integration of extensive interactions, deep neural networks, and rigorous validation can facilitate the rapid identification of candidate drugs for COVID-19 treatment.
CoSQL: A Conversational Text-to-SQL Challenge Towards Cross-Domain Natural Language Interfaces to Databases
Sep 11, 2019
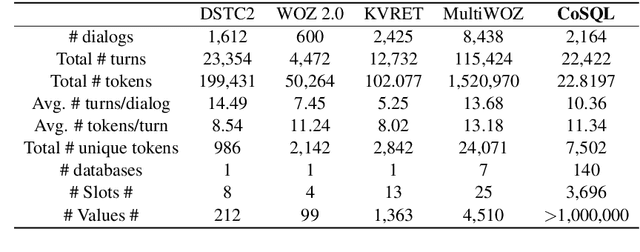
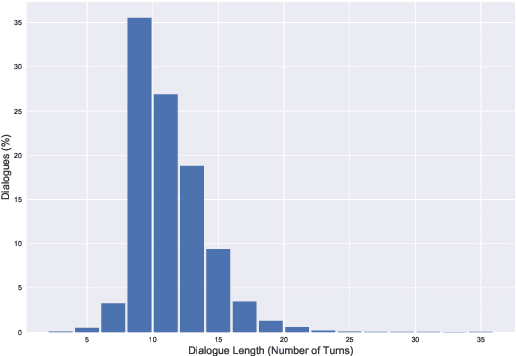
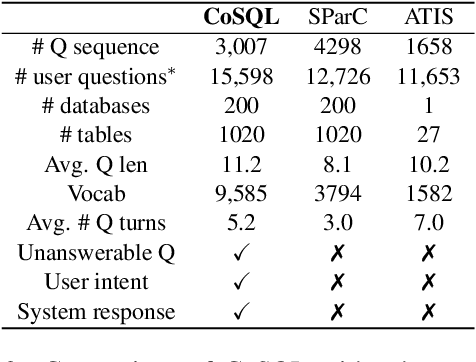
Abstract:We present CoSQL, a corpus for building cross-domain, general-purpose database (DB) querying dialogue systems. It consists of 30k+ turns plus 10k+ annotated SQL queries, obtained from a Wizard-of-Oz (WOZ) collection of 3k dialogues querying 200 complex DBs spanning 138 domains. Each dialogue simulates a real-world DB query scenario with a crowd worker as a user exploring the DB and a SQL expert retrieving answers with SQL, clarifying ambiguous questions, or otherwise informing of unanswerable questions. When user questions are answerable by SQL, the expert describes the SQL and execution results to the user, hence maintaining a natural interaction flow. CoSQL introduces new challenges compared to existing task-oriented dialogue datasets:(1) the dialogue states are grounded in SQL, a domain-independent executable representation, instead of domain-specific slot-value pairs, and (2) because testing is done on unseen databases, success requires generalizing to new domains. CoSQL includes three tasks: SQL-grounded dialogue state tracking, response generation from query results, and user dialogue act prediction. We evaluate a set of strong baselines for each task and show that CoSQL presents significant challenges for future research. The dataset, baselines, and leaderboard will be released at https://yale-lily.github.io/cosql.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge