Krzysztof J. Geras
Weakly-supervised High-resolution Segmentation of Mammography Images for Breast Cancer Diagnosis
Jun 15, 2021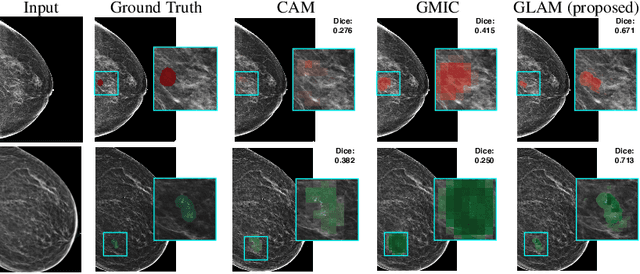
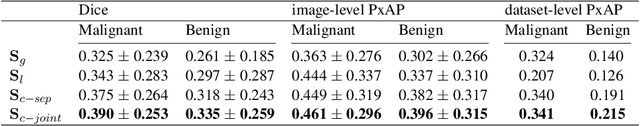
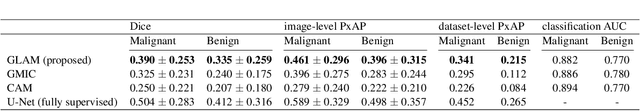
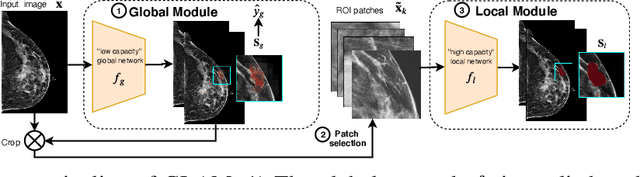

Abstract:In the last few years, deep learning classifiers have shown promising results in image-based medical diagnosis. However, interpreting the outputs of these models remains a challenge. In cancer diagnosis, interpretability can be achieved by localizing the region of the input image responsible for the output, i.e. the location of a lesion. Alternatively, segmentation or detection models can be trained with pixel-wise annotations indicating the locations of malignant lesions. Unfortunately, acquiring such labels is labor-intensive and requires medical expertise. To overcome this difficulty, weakly-supervised localization can be utilized. These methods allow neural network classifiers to output saliency maps highlighting the regions of the input most relevant to the classification task (e.g. malignant lesions in mammograms) using only image-level labels (e.g. whether the patient has cancer or not) during training. When applied to high-resolution images, existing methods produce low-resolution saliency maps. This is problematic in applications in which suspicious lesions are small in relation to the image size. In this work, we introduce a novel neural network architecture to perform weakly-supervised segmentation of high-resolution images. The proposed model selects regions of interest via coarse-level localization, and then performs fine-grained segmentation of those regions. We apply this model to breast cancer diagnosis with screening mammography, and validate it on a large clinically-realistic dataset. Measured by Dice similarity score, our approach outperforms existing methods by a large margin in terms of localization performance of benign and malignant lesions, relatively improving the performance by 39.6% and 20.0%, respectively. Code and the weights of some of the models are available at https://github.com/nyukat/GLAM
COVID-19 Prognosis via Self-Supervised Representation Learning and Multi-Image Prediction
Jan 25, 2021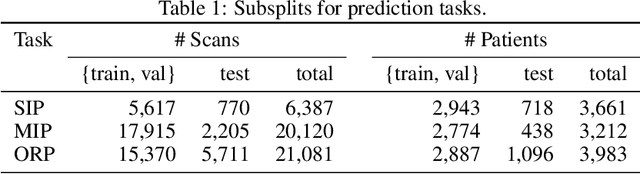
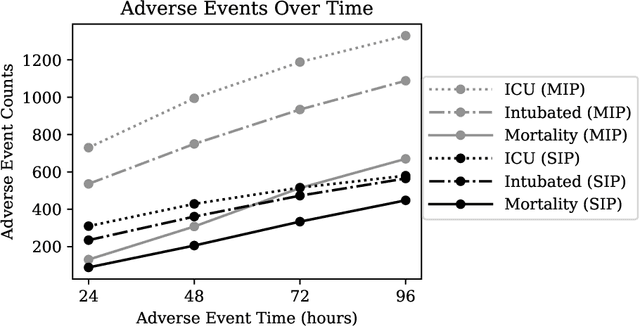
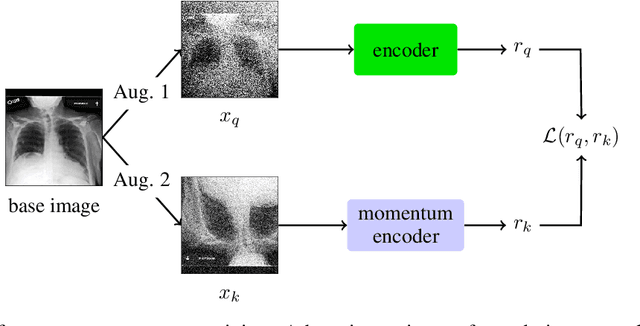
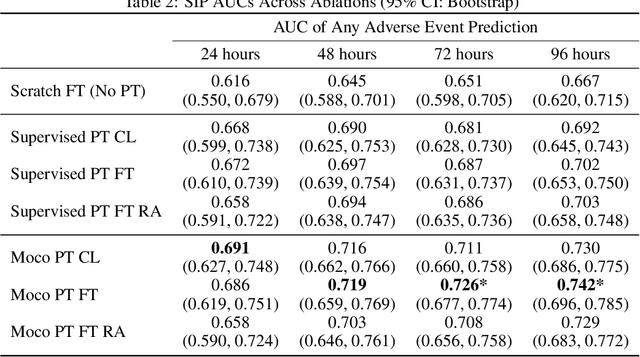
Abstract:The rapid spread of COVID-19 cases in recent months has strained hospital resources, making rapid and accurate triage of patients presenting to emergency departments a necessity. Machine learning techniques using clinical data such as chest X-rays have been used to predict which patients are most at risk of deterioration. We consider the task of predicting two types of patient deterioration based on chest X-rays: adverse event deterioration (i.e., transfer to the intensive care unit, intubation, or mortality) and increased oxygen requirements beyond 6 L per day. Due to the relative scarcity of COVID-19 patient data, existing solutions leverage supervised pretraining on related non-COVID images, but this is limited by the differences between the pretraining data and the target COVID-19 patient data. In this paper, we use self-supervised learning based on the momentum contrast (MoCo) method in the pretraining phase to learn more general image representations to use for downstream tasks. We present three results. The first is deterioration prediction from a single image, where our model achieves an area under receiver operating characteristic curve (AUC) of 0.742 for predicting an adverse event within 96 hours (compared to 0.703 with supervised pretraining) and an AUC of 0.765 for predicting oxygen requirements greater than 6 L a day at 24 hours (compared to 0.749 with supervised pretraining). We then propose a new transformer-based architecture that can process sequences of multiple images for prediction and show that this model can achieve an improved AUC of 0.786 for predicting an adverse event at 96 hours and an AUC of 0.848 for predicting mortalities at 96 hours. A small pilot clinical study suggested that the prediction accuracy of our model is comparable to that of experienced radiologists analyzing the same information.
Differences between human and machine perception in medical diagnosis
Nov 28, 2020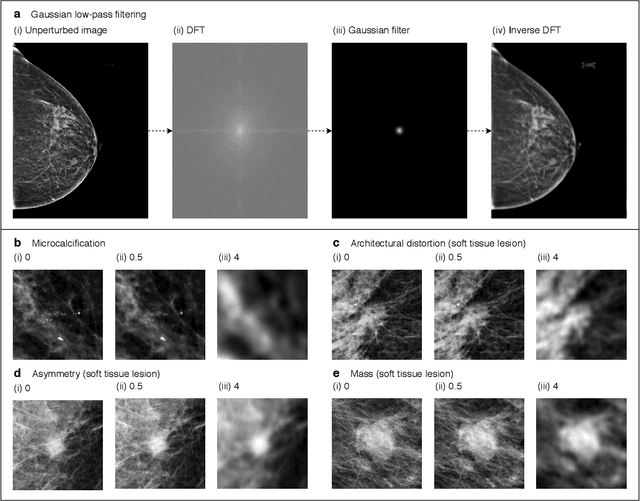
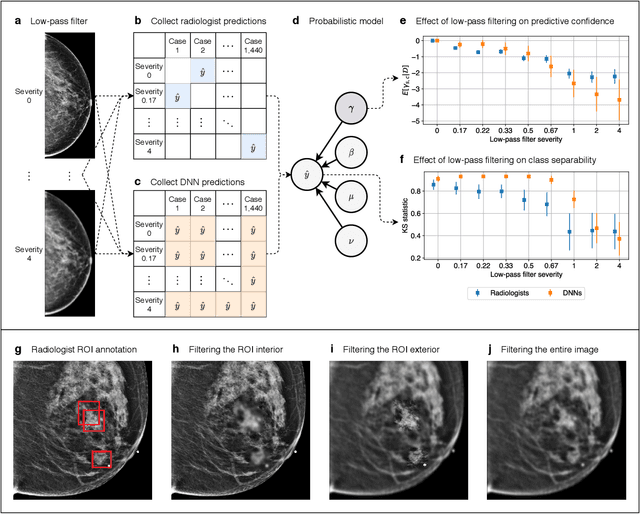
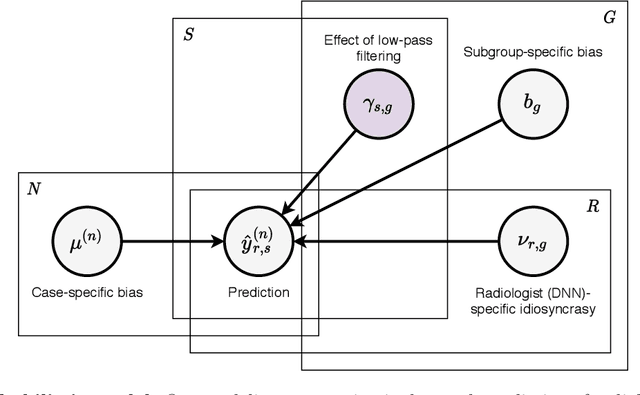
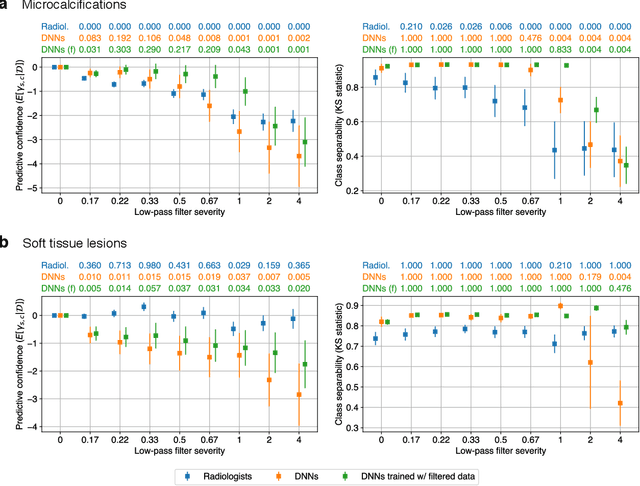
Abstract:Deep neural networks (DNNs) show promise in image-based medical diagnosis, but cannot be fully trusted since their performance can be severely degraded by dataset shifts to which human perception remains invariant. If we can better understand the differences between human and machine perception, we can potentially characterize and mitigate this effect. We therefore propose a framework for comparing human and machine perception in medical diagnosis. The two are compared with respect to their sensitivity to the removal of clinically meaningful information, and to the regions of an image deemed most suspicious. Drawing inspiration from the natural image domain, we frame both comparisons in terms of perturbation robustness. The novelty of our framework is that separate analyses are performed for subgroups with clinically meaningful differences. We argue that this is necessary in order to avert Simpson's paradox and draw correct conclusions. We demonstrate our framework with a case study in breast cancer screening, and reveal significant differences between radiologists and DNNs. We compare the two with respect to their robustness to Gaussian low-pass filtering, performing a subgroup analysis on microcalcifications and soft tissue lesions. For microcalcifications, DNNs use a separate set of high frequency components than radiologists, some of which lie outside the image regions considered most suspicious by radiologists. These features run the risk of being spurious, but if not, could represent potential new biomarkers. For soft tissue lesions, the divergence between radiologists and DNNs is even starker, with DNNs relying heavily on spurious high frequency components ignored by radiologists. Importantly, this deviation in soft tissue lesions was only observable through subgroup analysis, which highlights the importance of incorporating medical domain knowledge into our comparison framework.
Investigating and Simplifying Masking-based Saliency Methods for Model Interpretability
Oct 19, 2020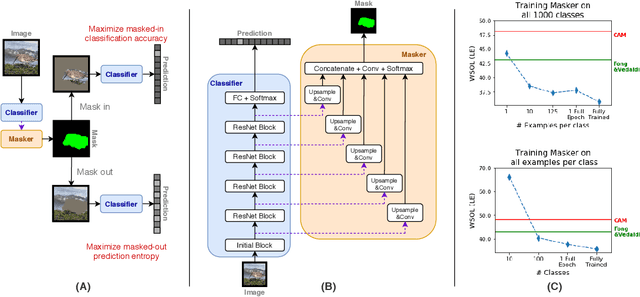
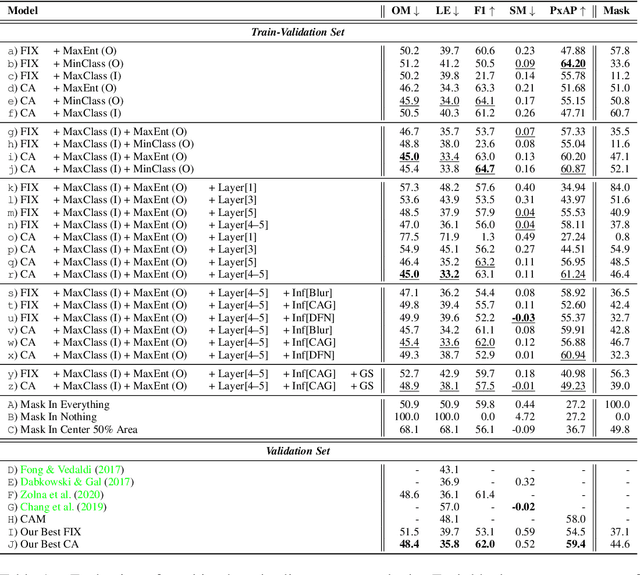
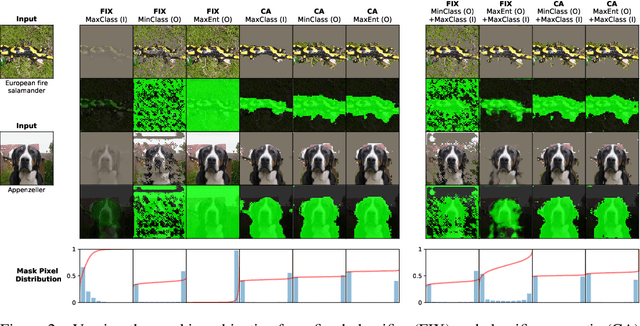
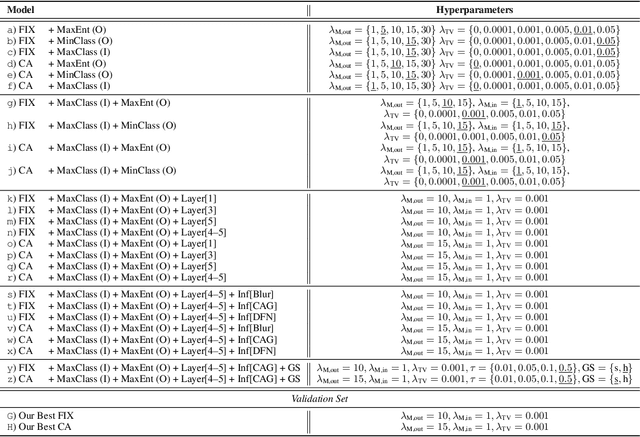
Abstract:Saliency maps that identify the most informative regions of an image for a classifier are valuable for model interpretability. A common approach to creating saliency maps involves generating input masks that mask out portions of an image to maximally deteriorate classification performance, or mask in an image to preserve classification performance. Many variants of this approach have been proposed in the literature, such as counterfactual generation and optimizing over a Gumbel-Softmax distribution. Using a general formulation of masking-based saliency methods, we conduct an extensive evaluation study of a number of recently proposed variants to understand which elements of these methods meaningfully improve performance. Surprisingly, we find that a well-tuned, relatively simple formulation of a masking-based saliency model outperforms many more complex approaches. We find that the most important ingredients for high quality saliency map generation are (1) using both masked-in and masked-out objectives and (2) training the classifier alongside the masking model. Strikingly, we show that a masking model can be trained with as few as 10 examples per class and still generate saliency maps with only a 0.7-point increase in localization error.
Reducing false-positive biopsies with deep neural networks that utilize local and global information in screening mammograms
Sep 19, 2020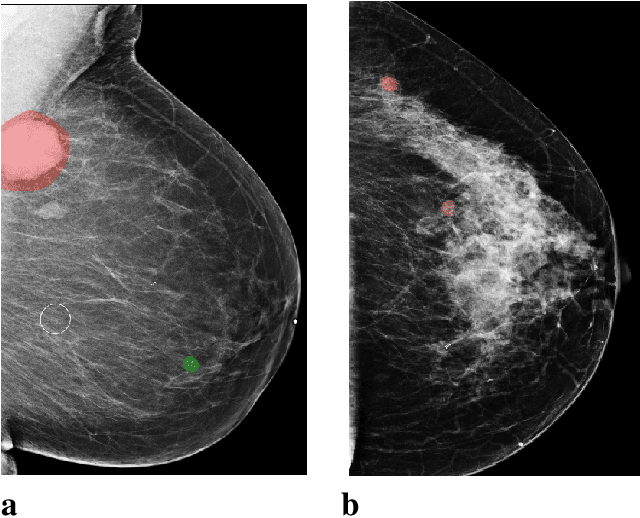
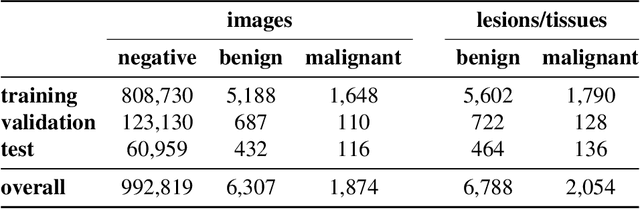
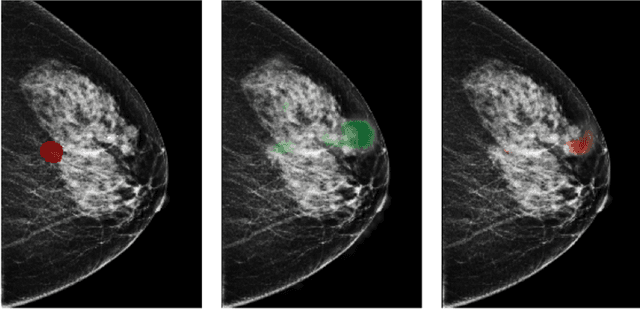
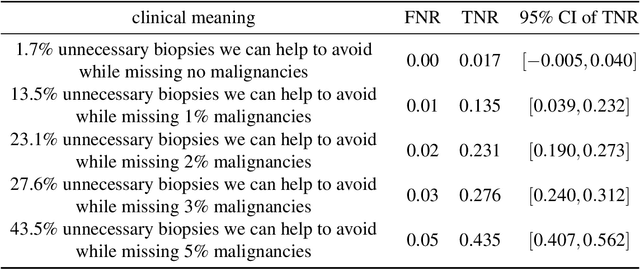
Abstract:Breast cancer is the most common cancer in women, and hundreds of thousands of unnecessary biopsies are done around the world at a tremendous cost. It is crucial to reduce the rate of biopsies that turn out to be benign tissue. In this study, we build deep neural networks (DNNs) to classify biopsied lesions as being either malignant or benign, with the goal of using these networks as second readers serving radiologists to further reduce the number of false positive findings. We enhance the performance of DNNs that are trained to learn from small image patches by integrating global context provided in the form of saliency maps learned from the entire image into their reasoning, similar to how radiologists consider global context when evaluating areas of interest. Our experiments are conducted on a dataset of 229,426 screening mammography exams from 141,473 patients. We achieve an AUC of 0.8 on a test set consisting of 464 benign and 136 malignant lesions.
An artificial intelligence system for predicting the deterioration of COVID-19 patients in the emergency department
Aug 04, 2020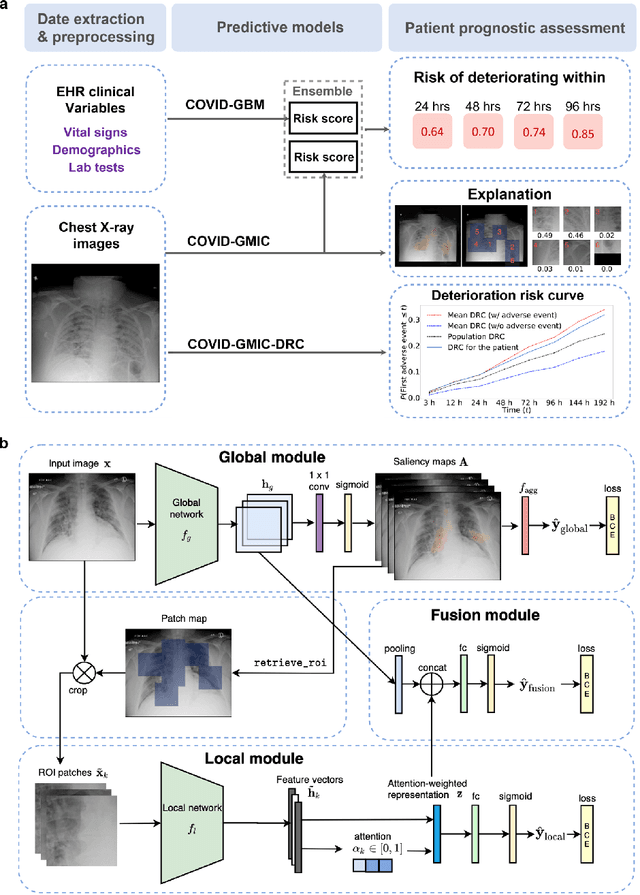
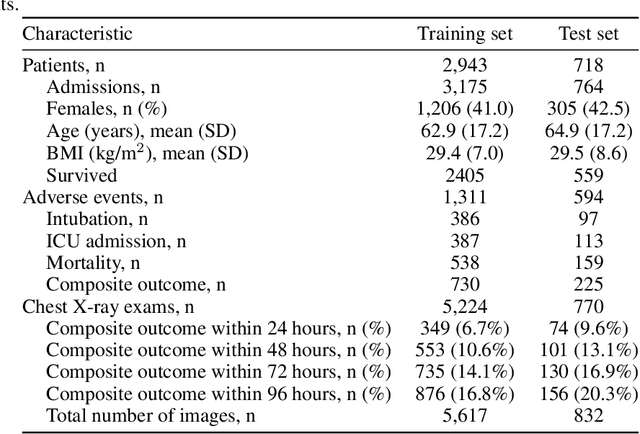
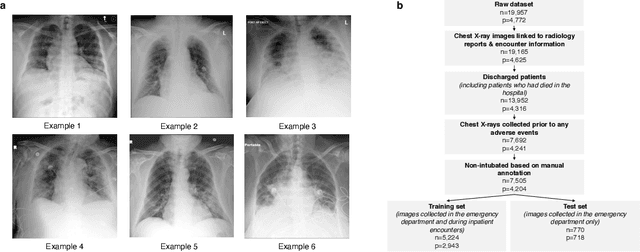
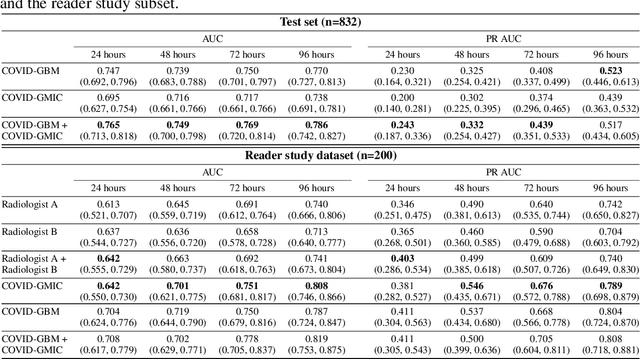
Abstract:During the COVID-19 pandemic, rapid and accurate triage of patients at the emergency department is critical to inform decision-making. We propose a data-driven approach for automatic prediction of deterioration risk using a deep neural network that learns from chest X-ray images, and a gradient boosting model that learns from routine clinical variables. Our AI prognosis system, trained using data from 3,661 patients, achieves an AUC of 0.786 (95% CI: 0.742-0.827) when predicting deterioration within 96 hours. The deep neural network extracts informative areas of chest X-ray images to assist clinicians in interpreting the predictions, and performs comparably to two radiologists in a reader study. In order to verify performance in a real clinical setting, we silently deployed a preliminary version of the deep neural network at NYU Langone Health during the first wave of the pandemic, which produced accurate predictions in real-time. In summary, our findings demonstrate the potential of the proposed system for assisting front-line physicians in the triage of COVID-19 patients.
Understanding the robustness of deep neural network classifiers for breast cancer screening
Mar 23, 2020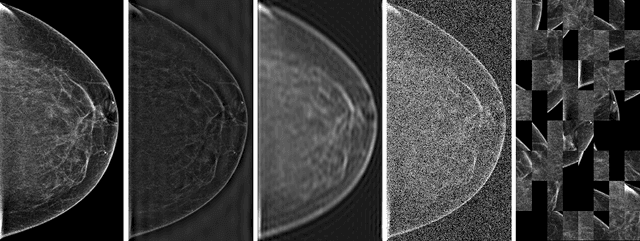
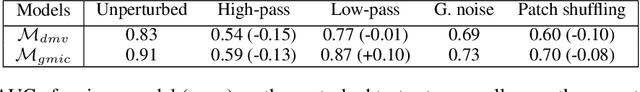
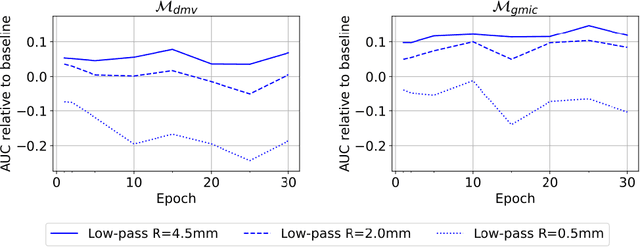
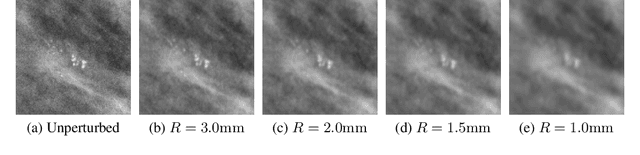
Abstract:Deep neural networks (DNNs) show promise in breast cancer screening, but their robustness to input perturbations must be better understood before they can be clinically implemented. There exists extensive literature on this subject in the context of natural images that can potentially be built upon. However, it cannot be assumed that conclusions about robustness will transfer from natural images to mammogram images, due to significant differences between the two image modalities. In order to determine whether conclusions will transfer, we measure the sensitivity of a radiologist-level screening mammogram image classifier to four commonly studied input perturbations that natural image classifiers are sensitive to. We find that mammogram image classifiers are also sensitive to these perturbations, which suggests that we can build on the existing literature. We also perform a detailed analysis on the effects of low-pass filtering, and find that it degrades the visibility of clinically meaningful features called microcalcifications. Since low-pass filtering removes semantically meaningful information that is predictive of breast cancer, we argue that it is undesirable for mammogram image classifiers to be invariant to it. This is in contrast to natural images, where we do not want DNNs to be sensitive to low-pass filtering due to its tendency to remove information that is human-incomprehensible.
An interpretable classifier for high-resolution breast cancer screening images utilizing weakly supervised localization
Feb 13, 2020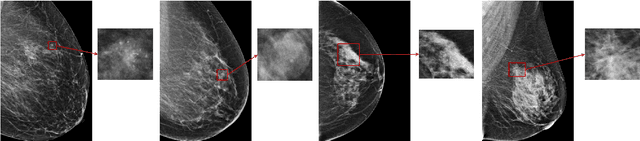
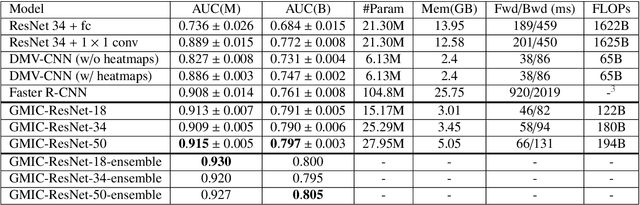
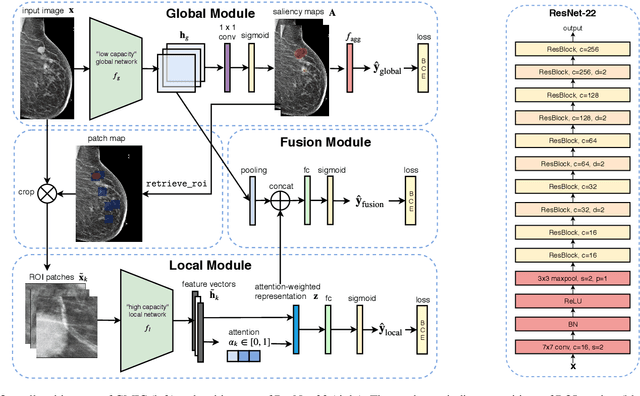
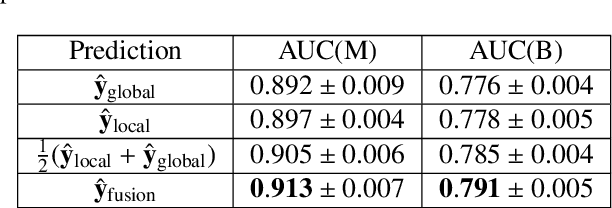
Abstract:Medical images differ from natural images in significantly higher resolutions and smaller regions of interest. Because of these differences, neural network architectures that work well for natural images might not be applicable to medical image analysis. In this work, we extend the globally-aware multiple instance classifier, a framework we proposed to address these unique properties of medical images. This model first uses a low-capacity, yet memory-efficient, network on the whole image to identify the most informative regions. It then applies another higher-capacity network to collect details from chosen regions. Finally, it employs a fusion module that aggregates global and local information to make a final prediction. While existing methods often require lesion segmentation during training, our model is trained with only image-level labels and can generate pixel-level saliency maps indicating possible malignant findings. We apply the model to screening mammography interpretation: predicting the presence or absence of benign and malignant lesions. On the NYU Breast Cancer Screening Dataset, consisting of more than one million images, our model achieves an AUC of 0.93 in classifying breasts with malignant findings, outperforming ResNet-34 and Faster R-CNN. Compared to ResNet-34, our model is 4.1x faster for inference while using 78.4% less GPU memory. Furthermore, we demonstrate, in a reader study, that our model surpasses radiologist-level AUC by a margin of 0.11. The proposed model is available online: https://github.com/nyukat/GMIC.
Improving localization-based approaches for breast cancer screening exam classification
Aug 01, 2019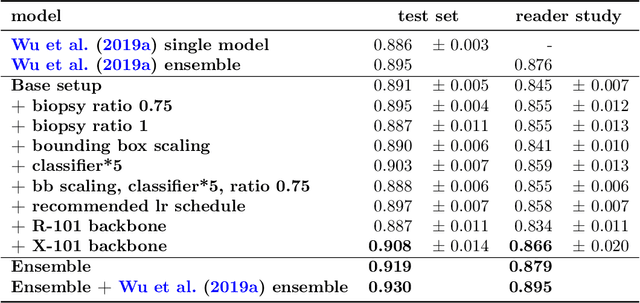
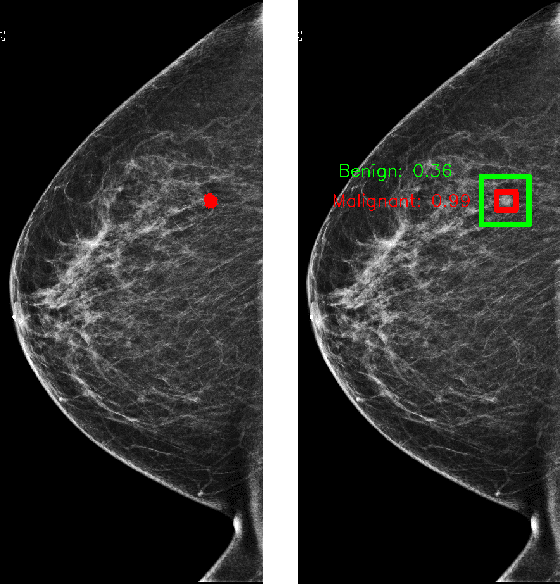
Abstract:We trained and evaluated a localization-based deep CNN for breast cancer screening exam classification on over 200,000 exams (over 1,000,000 images). Our model achieves an AUC of 0.919 in predicting malignancy in patients undergoing breast cancer screening, reducing the error rate of the baseline (Wu et al., 2019a) by 23%. In addition, the models generates bounding boxes for benign and malignant findings, providing interpretable predictions.
Screening Mammogram Classification with Prior Exams
Jul 30, 2019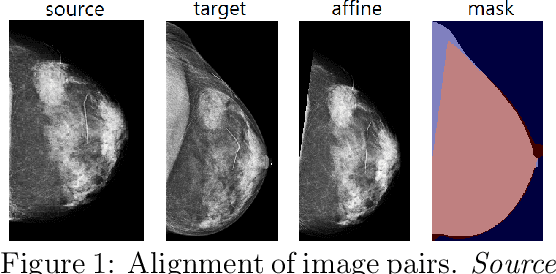
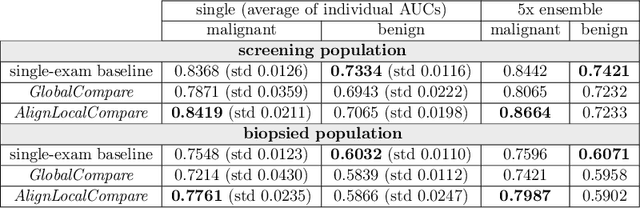
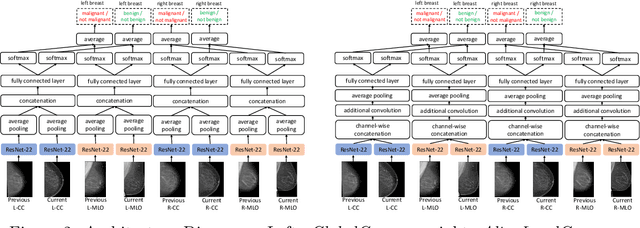
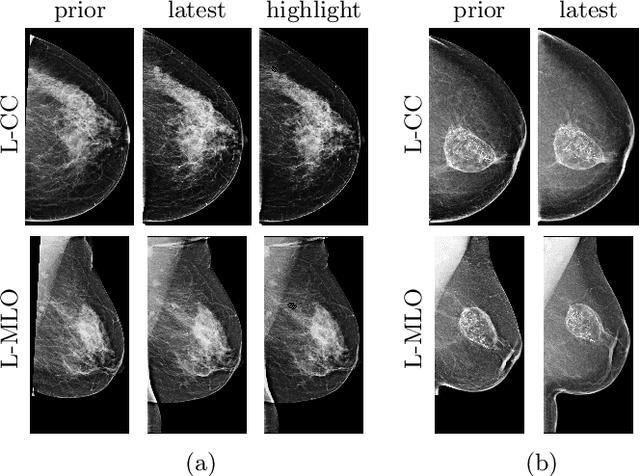
Abstract:Radiologists typically compare a patient's most recent breast cancer screening exam to their previous ones in making informed diagnoses. To reflect this practice, we propose new neural network models that compare pairs of screening mammograms from the same patient. We train and evaluate our proposed models on over 665,000 pairs of images (over 166,000 pairs of exams). Our best model achieves an AUC of 0.866 in predicting malignancy in patients who underwent breast cancer screening, reducing the error rate of the corresponding baseline.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge