Narges Razavian
Scaling Recurrence-aware Foundation Models for Clinical Records via Next-Visit Prediction
Mar 25, 2026Abstract:While large-scale pretraining has revolutionized language modeling, its potential remains underexplored in healthcare with structured electronic health records (EHRs). We present RAVEN, a novel generative pretraining strategy for sequential EHR data based on Recurrence-Aware next-Visit EveNt prediction. Leveraging a dataset of over one million unique individuals, our model learns to autoregressively generate tokenized clinical events for the next visit conditioned on patient history. We introduce regularization on predicting repeated events and highlight a key pitfall in EHR-based foundation model evaluations: repeated event tokens can inflate performance metrics when new onsets are not distinguished from subsequent occurrences. Furthermore, we empirically investigate the scaling behaviors in a data-constrained, compute-saturated regime, showing that simply increasing model size is suboptimal without commensurate increases in data volume. We evaluate our model via zero-shot prediction for forecasting the incidence of a diverse set of diseases, where it rivals fully fine-tuned representation-based Transformer models and outperforms widely used simulation-based next-token approaches. Finally, without additional parameter updates, we show that RAVEN can generalize to an external patient cohort under lossy clinical code mappings and feature coverage gaps.
Cerebra: A Multidisciplinary AI Board for Multimodal Dementia Characterization and Risk Assessment
Mar 24, 2026Abstract:Modern clinical practice increasingly depends on reasoning over heterogeneous, evolving, and incomplete patient data. Although recent advances in multimodal foundation models have improved performance on various clinical tasks, most existing models remain static, opaque, and poorly aligned with real-world clinical workflows. We present Cerebra, an interactive multi-agent AI team that coordinates specialized agents for EHR, clinical notes, and medical imaging analysis. These outputs are synthesized into a clinician-facing dashboard that combines visual analytics with a conversational interface, enabling clinicians to interrogate predictions and contextualize risk at the point of care. Cerebra supports privacy-preserving deployment by operating on structured representations and remains robust when modalities are incomplete. We evaluated Cerebra using a massive multi-institutional dataset spanning 3 million patients from four independent healthcare systems. Cerebra consistently outperformed both state-of-the-art single-modality models and large multimodal language model baselines. In dementia risk prediction, it achieved AUROCs up to 0.80, compared with 0.74 for the strongest single-modality model and 0.68 for language model baselines. For dementia diagnosis, it achieved an AUROC of 0.86, and for survival prediction, a C-index of 0.81. In a reader study with experienced physicians, Cerebra significantly improved expert performance, increasing accuracy by 17.5 percentage points in prospective dementia risk estimation. These results demonstrate Cerebra's potential for interpretable, robust decision support in clinical care.
A Multidisciplinary AI Board for Multimodal Dementia Characterization and Risk Assessment
Mar 23, 2026Abstract:Modern clinical practice increasingly depends on reasoning over heterogeneous, evolving, and incomplete patient data. Although recent advances in multimodal foundation models have improved performance on various clinical tasks, most existing models remain static, opaque, and poorly aligned with real-world clinical workflows. We present Cerebra, an interactive multi-agent AI team that coordinates specialized agents for EHR, clinical notes, and medical imaging analysis. These outputs are synthesized into a clinician-facing dashboard that combines visual analytics with a conversational interface, enabling clinicians to interrogate predictions and contextualize risk at the point of care. Cerebra supports privacy-preserving deployment by operating on structured representations and remains robust when modalities are incomplete. We evaluated Cerebra using a massive multi-institutional dataset spanning 3 million patients from four independent healthcare systems. Cerebra consistently outperformed both state-of-the-art single-modality models and large multimodal language model baselines. In dementia risk prediction, it achieved AUROCs up to 0.80, compared with 0.74 for the strongest single-modality model and 0.68 for language model baselines. For dementia diagnosis, it achieved an AUROC of 0.86, and for survival prediction, a C-index of 0.81. In a reader study with experienced physicians, Cerebra significantly improved expert performance, increasing accuracy by 17.5 percentage points in prospective dementia risk estimation. These results demonstrate Cerebra's potential for interpretable, robust decision support in clinical care.
3D Foundation AI Model for Generalizable Disease Detection in Head Computed Tomography
Feb 04, 2025



Abstract:Head computed tomography (CT) imaging is a widely-used imaging modality with multitudes of medical indications, particularly in assessing pathology of the brain, skull, and cerebrovascular system. It is commonly the first-line imaging in neurologic emergencies given its rapidity of image acquisition, safety, cost, and ubiquity. Deep learning models may facilitate detection of a wide range of diseases. However, the scarcity of high-quality labels and annotations, particularly among less common conditions, significantly hinders the development of powerful models. To address this challenge, we introduce FM-CT: a Foundation Model for Head CT for generalizable disease detection, trained using self-supervised learning. Our approach pre-trains a deep learning model on a large, diverse dataset of 361,663 non-contrast 3D head CT scans without the need for manual annotations, enabling the model to learn robust, generalizable features. To investigate the potential of self-supervised learning in head CT, we employed both discrimination with self-distillation and masked image modeling, and we construct our model in 3D rather than at the slice level (2D) to exploit the structure of head CT scans more comprehensively and efficiently. The model's downstream classification performance is evaluated using internal and three external datasets, encompassing both in-distribution (ID) and out-of-distribution (OOD) data. Our results demonstrate that the self-supervised foundation model significantly improves performance on downstream diagnostic tasks compared to models trained from scratch and previous 3D CT foundation models on scarce annotated datasets. This work highlights the effectiveness of self-supervised learning in medical imaging and sets a new benchmark for head CT image analysis in 3D, enabling broader use of artificial intelligence for head CT-based diagnosis.
Automatic Detection of Alzheimer's Disease with Multi-Modal Fusion of Clinical MRI Scans
Nov 30, 2023



Abstract:The aging population of the U.S. drives the prevalence of Alzheimer's disease. Brookmeyer et al. forecasts approximately 15 million Americans will have either clinical AD or mild cognitive impairment by 2060. In response to this urgent call, methods for early detection of Alzheimer's disease have been developed for prevention and pre-treatment. Notably, literature on the application of deep learning in the automatic detection of the disease has been proliferating. This study builds upon previous literature and maintains a focus on leveraging multi-modal information to enhance automatic detection. We aim to predict the stage of the disease - Cognitively Normal (CN), Mildly Cognitive Impairment (MCI), and Alzheimer's Disease (AD), based on two different types of brain MRI scans. We design an AlexNet-based deep learning model that learns the synergy of complementary information from both T1 and FLAIR MRI scans.
Making Self-supervised Learning Robust to Spurious Correlation via Learning-speed Aware Sampling
Nov 29, 2023



Abstract:Self-supervised learning (SSL) has emerged as a powerful technique for learning rich representations from unlabeled data. The data representations are able to capture many underlying attributes of data, and be useful in downstream prediction tasks. In real-world settings, spurious correlations between some attributes (e.g. race, gender and age) and labels for downstream tasks often exist, e.g. cancer is usually more prevalent among elderly patients. In this paper, we investigate SSL in the presence of spurious correlations and show that the SSL training loss can be minimized by capturing only a subset of the conspicuous features relevant to those sensitive attributes, despite the presence of other important predictive features for the downstream tasks. To address this issue, we investigate the learning dynamics of SSL and observe that the learning is slower for samples that conflict with such correlations (e.g. elder patients without cancer). Motivated by these findings, we propose a learning-speed aware SSL (LA-SSL) approach, in which we sample each training data with a probability that is inversely related to its learning speed. We evaluate LA-SSL on three datasets that exhibit spurious correlations between different attributes, demonstrating that it improves the robustness of pretrained representations on downstream classification tasks.
Multiple Instance Learning via Iterative Self-Paced Supervised Contrastive Learning
Oct 17, 2022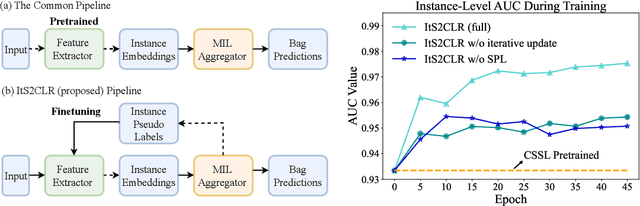
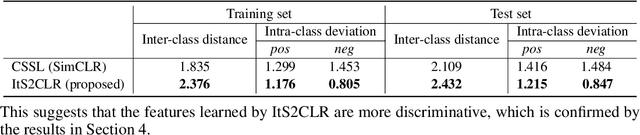
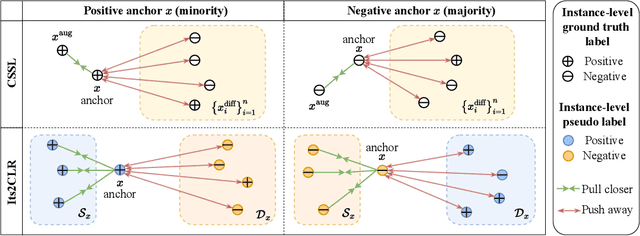

Abstract:Learning representations for individual instances when only bag-level labels are available is a fundamental challenge in multiple instance learning (MIL). Recent works have shown promising results using contrastive self-supervised learning (CSSL), which learns to push apart representations corresponding to two different randomly-selected instances. Unfortunately, in real-world applications such as medical image classification, there is often class imbalance, so randomly-selected instances mostly belong to the same majority class, which precludes CSSL from learning inter-class differences. To address this issue, we propose a novel framework, Iterative Self-paced Supervised Contrastive Learning for MIL Representations (ItS2CLR), which improves the learned representation by exploiting instance-level pseudo labels derived from the bag-level labels. The framework employs a novel self-paced sampling strategy to ensure the accuracy of pseudo labels. We evaluate ItS2CLR on three medical datasets, showing that it improves the quality of instance-level pseudo labels and representations, and outperforms existing MIL methods in terms of both bag and instance level accuracy.
Interpretable Prediction of Lung Squamous Cell Carcinoma Recurrence With Self-supervised Learning
Mar 23, 2022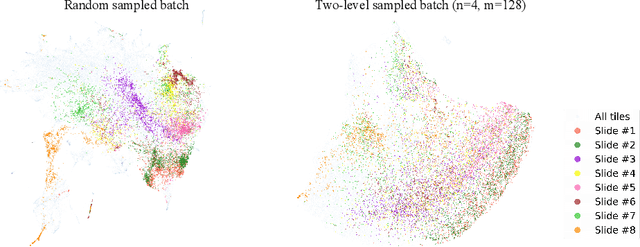
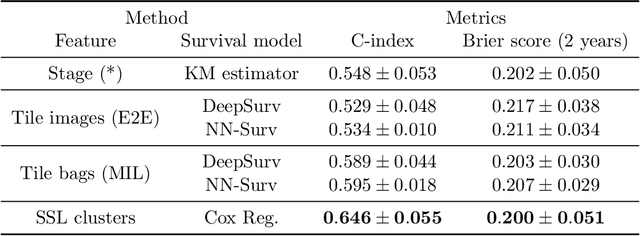
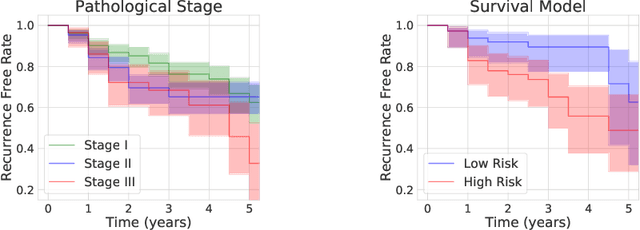

Abstract:Lung squamous cell carcinoma (LSCC) has a high recurrence and metastasis rate. Factors influencing recurrence and metastasis are currently unknown and there are no distinct histopathological or morphological features indicating the risks of recurrence and metastasis in LSCC. Our study focuses on the recurrence prediction of LSCC based on H&E-stained histopathological whole-slide images (WSI). Due to the small size of LSCC cohorts in terms of patients with available recurrence information, standard end-to-end learning with various convolutional neural networks for this task tends to overfit. Also, the predictions made by these models are hard to interpret. Histopathology WSIs are typically very large and are therefore processed as a set of smaller tiles. In this work, we propose a novel conditional self-supervised learning (SSL) method to learn representations of WSI at the tile level first, and leverage clustering algorithms to identify the tiles with similar histopathological representations. The resulting representations and clusters from self-supervision are used as features of a survival model for recurrence prediction at the patient level. Using two publicly available datasets from TCGA and CPTAC, we show that our LSCC recurrence prediction survival model outperforms both LSCC pathological stage-based approach and machine learning baselines such as multiple instance learning. The proposed method also enables us to explain the recurrence histopathological risk factors via the derived clusters. This can help pathologists derive new hypotheses regarding morphological features associated with LSCC recurrence.
Deep Probability Estimation
Nov 21, 2021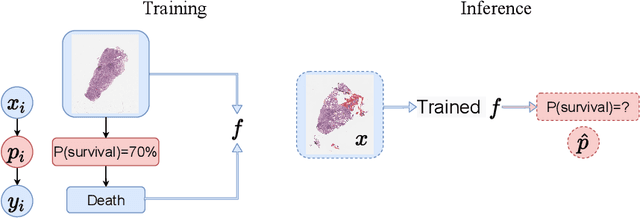
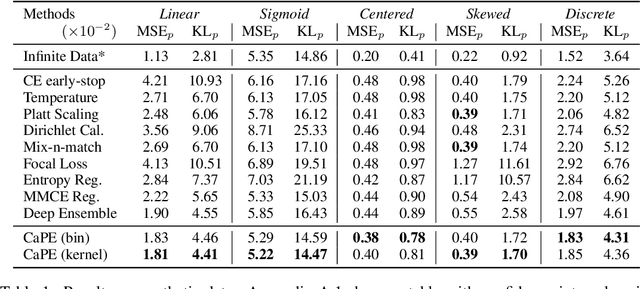
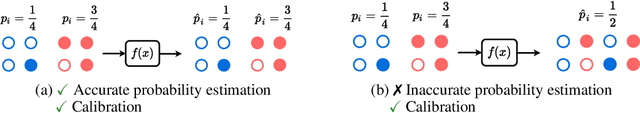
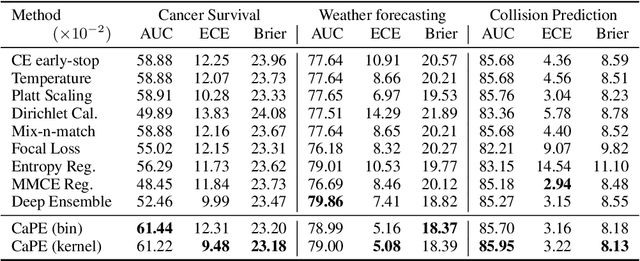
Abstract:Reliable probability estimation is of crucial importance in many real-world applications where there is inherent uncertainty, such as weather forecasting, medical prognosis, or collision avoidance in autonomous vehicles. Probability-estimation models are trained on observed outcomes (e.g. whether it has rained or not, or whether a patient has died or not), because the ground-truth probabilities of the events of interest are typically unknown. The problem is therefore analogous to binary classification, with the important difference that the objective is to estimate probabilities rather than predicting the specific outcome. The goal of this work is to investigate probability estimation from high-dimensional data using deep neural networks. There exist several methods to improve the probabilities generated by these models but they mostly focus on classification problems where the probabilities are related to model uncertainty. In the case of problems with inherent uncertainty, it is challenging to evaluate performance without access to ground-truth probabilities. To address this, we build a synthetic dataset to study and compare different computable metrics. We evaluate existing methods on the synthetic data as well as on three real-world probability estimation tasks, all of which involve inherent uncertainty: precipitation forecasting from radar images, predicting cancer patient survival from histopathology images, and predicting car crashes from dashcam videos. Finally, we also propose a new method for probability estimation using neural networks, which modifies the training process to promote output probabilities that are consistent with empirical probabilities computed from the data. The method outperforms existing approaches on most metrics on the simulated as well as real-world data.
Causal Effect Variational Autoencoder with Uniform Treatment
Nov 16, 2021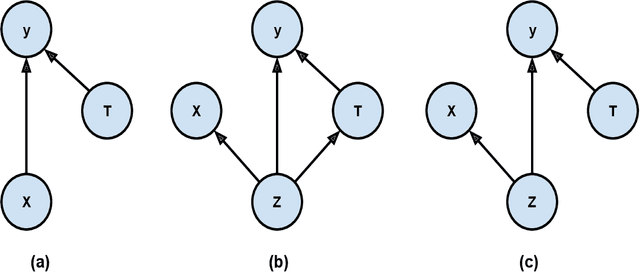
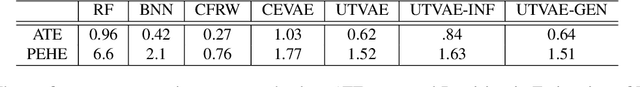
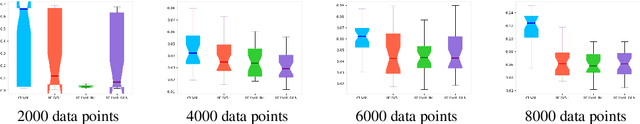
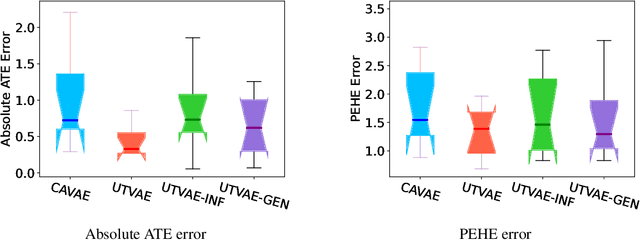
Abstract:Causal effect variational autoencoder (CEVAE) are trained to predict the outcome given observational treatment data, while uniform treatment variational autoencoders (UTVAE) are trained with uniform treatment distribution using importance sampling. In this paper, we show that using uniform treatment over observational treatment distribution leads to better causal inference by mitigating the distribution shift that occurs from training to test time. We also explore the combination of uniform and observational treatment distributions with inference and generative network training objectives to find a better training procedure for inferring treatment effect. Experimentally, we find that the proposed UTVAE yields better absolute average treatment effect error and precision in estimation of heterogeneous effect error than the CEVAE on synthetic and IHDP datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge