Katherine Heller
Participatory Systems for Personalized Prediction
Feb 08, 2023



Abstract:Machine learning models are often personalized based on information that is protected, sensitive, self-reported, or costly to acquire. These models use information about people, but do not facilitate nor inform their \emph{consent}. Individuals cannot opt out of reporting information that a model needs to personalize their predictions, nor tell if they would benefit from personalization in the first place. In this work, we introduce a new family of prediction models, called \emph{participatory systems}, that allow individuals to opt into personalization at prediction time. We present a model-agnostic algorithm to learn participatory systems for supervised learning tasks where models are personalized with categorical group attributes. We conduct a comprehensive empirical study of participatory systems in clinical prediction tasks, comparing them to common approaches for personalization and imputation. Our results demonstrate that participatory systems can facilitate and inform consent in a way that improves performance and privacy across all groups who report personal data.
Evaluation Gaps in Machine Learning Practice
May 11, 2022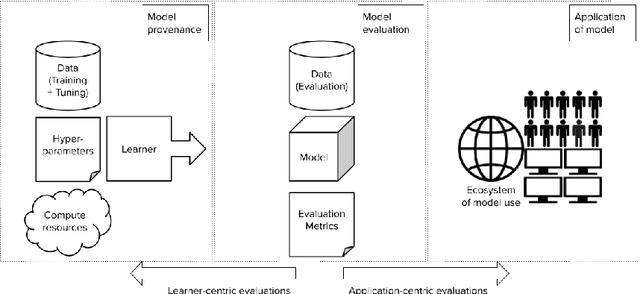

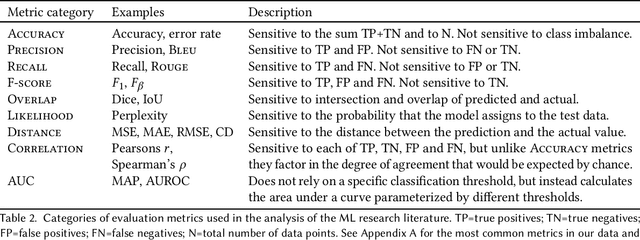
Abstract:Forming a reliable judgement of a machine learning (ML) model's appropriateness for an application ecosystem is critical for its responsible use, and requires considering a broad range of factors including harms, benefits, and responsibilities. In practice, however, evaluations of ML models frequently focus on only a narrow range of decontextualized predictive behaviours. We examine the evaluation gaps between the idealized breadth of evaluation concerns and the observed narrow focus of actual evaluations. Through an empirical study of papers from recent high-profile conferences in the Computer Vision and Natural Language Processing communities, we demonstrate a general focus on a handful of evaluation methods. By considering the metrics and test data distributions used in these methods, we draw attention to which properties of models are centered in the field, revealing the properties that are frequently neglected or sidelined during evaluation. By studying these properties, we demonstrate the machine learning discipline's implicit assumption of a range of commitments which have normative impacts; these include commitments to consequentialism, abstractability from context, the quantifiability of impacts, the limited role of model inputs in evaluation, and the equivalence of different failure modes. Shedding light on these assumptions enables us to question their appropriateness for ML system contexts, pointing the way towards more contextualized evaluation methodologies for robustly examining the trustworthiness of ML models
Disability prediction in multiple sclerosis using performance outcome measures and demographic data
Apr 08, 2022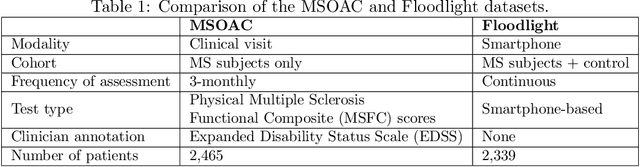
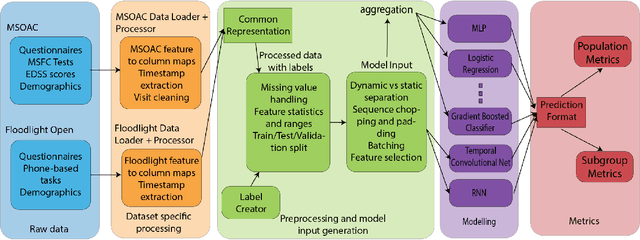
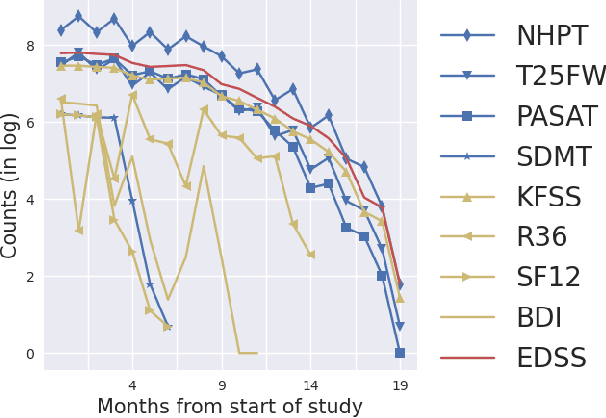
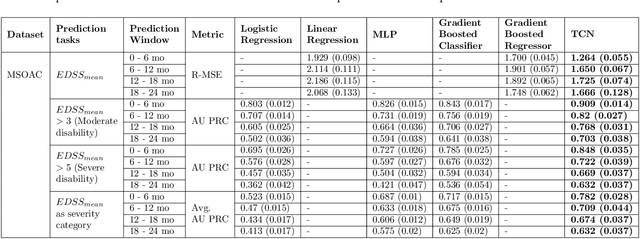
Abstract:Literature on machine learning for multiple sclerosis has primarily focused on the use of neuroimaging data such as magnetic resonance imaging and clinical laboratory tests for disease identification. However, studies have shown that these modalities are not consistent with disease activity such as symptoms or disease progression. Furthermore, the cost of collecting data from these modalities is high, leading to scarce evaluations. In this work, we used multi-dimensional, affordable, physical and smartphone-based performance outcome measures (POM) in conjunction with demographic data to predict multiple sclerosis disease progression. We performed a rigorous benchmarking exercise on two datasets and present results across 13 clinically actionable prediction endpoints and 6 machine learning models. To the best of our knowledge, our results are the first to show that it is possible to predict disease progression using POMs and demographic data in the context of both clinical trials and smartphone-base studies by using two datasets. Moreover, we investigate our models to understand the impact of different POMs and demographics on model performance through feature ablation studies. We also show that model performance is similar across different demographic subgroups (based on age and sex). To enable this work, we developed an end-to-end reusable pre-processing and machine learning framework which allows quicker experimentation over disparate MS datasets.
Healthsheet: Development of a Transparency Artifact for Health Datasets
Feb 26, 2022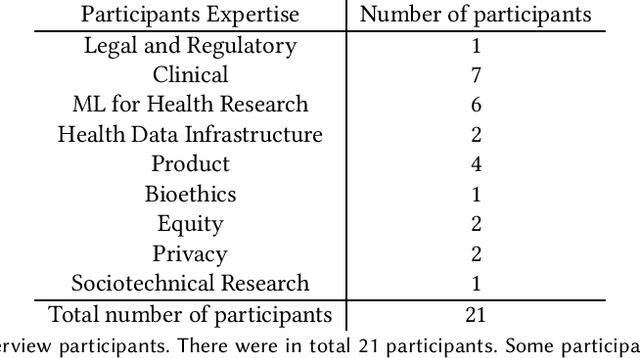
Abstract:Machine learning (ML) approaches have demonstrated promising results in a wide range of healthcare applications. Data plays a crucial role in developing ML-based healthcare systems that directly affect people's lives. Many of the ethical issues surrounding the use of ML in healthcare stem from structural inequalities underlying the way we collect, use, and handle data. Developing guidelines to improve documentation practices regarding the creation, use, and maintenance of ML healthcare datasets is therefore of critical importance. In this work, we introduce Healthsheet, a contextualized adaptation of the original datasheet questionnaire ~\cite{gebru2018datasheets} for health-specific applications. Through a series of semi-structured interviews, we adapt the datasheets for healthcare data documentation. As part of the Healthsheet development process and to understand the obstacles researchers face in creating datasheets, we worked with three publicly-available healthcare datasets as our case studies, each with different types of structured data: Electronic health Records (EHR), clinical trial study data, and smartphone-based performance outcome measures. Our findings from the interviewee study and case studies show 1) that datasheets should be contextualized for healthcare, 2) that despite incentives to adopt accountability practices such as datasheets, there is a lack of consistency in the broader use of these practices 3) how the ML for health community views datasheets and particularly \textit{Healthsheets} as diagnostic tool to surface the limitations and strength of datasets and 4) the relative importance of different fields in the datasheet to healthcare concerns.
Maintaining fairness across distribution shift: do we have viable solutions for real-world applications?
Feb 02, 2022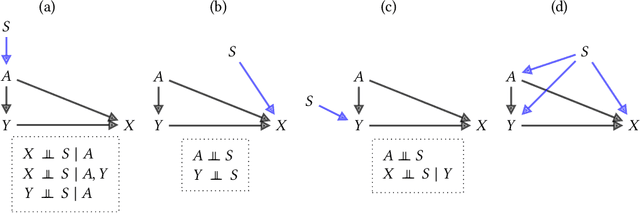
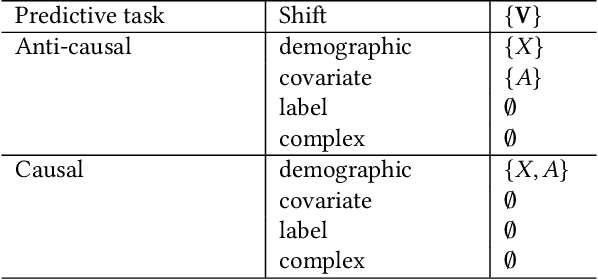
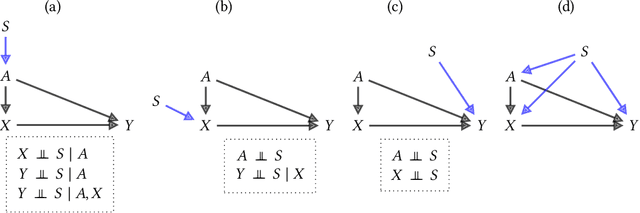
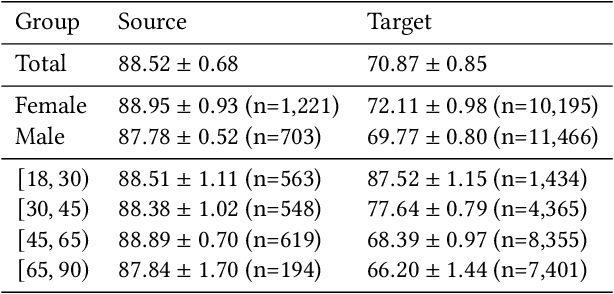
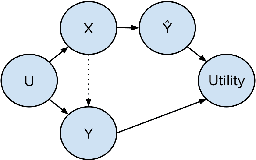
Abstract:Fairness and robustness are often considered as orthogonal dimensions when evaluating machine learning models. However, recent work has revealed interactions between fairness and robustness, showing that fairness properties are not necessarily maintained under distribution shift. In healthcare settings, this can result in e.g. a model that performs fairly according to a selected metric in "hospital A" showing unfairness when deployed in "hospital B". While a nascent field has emerged to develop provable fair and robust models, it typically relies on strong assumptions about the shift, limiting its impact for real-world applications. In this work, we explore the settings in which recently proposed mitigation strategies are applicable by referring to a causal framing. Using examples of predictive models in dermatology and electronic health records, we show that real-world applications are complex and often invalidate the assumptions of such methods. Our work hence highlights technical, practical, and engineering gaps that prevent the development of robustly fair machine learning models for real-world applications. Finally, we discuss potential remedies at each step of the machine learning pipeline.
Deep Cox Mixtures for Survival Regression
Jan 16, 2021

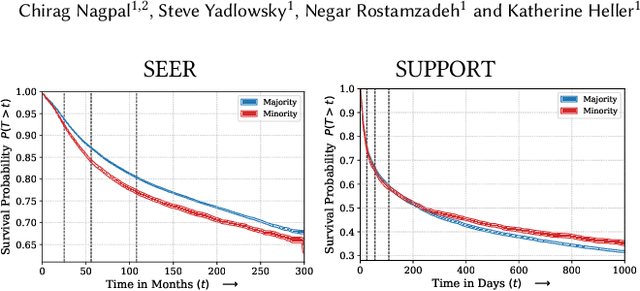

Abstract:Survival analysis is a challenging variation of regression modeling because of the presence of censoring, where the outcome measurement is only partially known, due to, for example, loss to follow up. Such problems come up frequently in medical applications, making survival analysis a key endeavor in biostatistics and machine learning for healthcare, with Cox regression models being amongst the most commonly employed models. We describe a new approach for survival analysis regression models, based on learning mixtures of Cox regressions to model individual survival distributions. We propose an approximation to the Expectation Maximization algorithm for this model that does hard assignments to mixture groups to make optimization efficient. In each group assignment, we fit the hazard ratios within each group using deep neural networks, and the baseline hazard for each mixture component non-parametrically. We perform experiments on multiple real world datasets, and look at the mortality rates of patients across ethnicity and gender. We emphasize the importance of calibration in healthcare settings and demonstrate that our approach outperforms classical and modern survival analysis baselines, both in terms of discriminative performance and calibration, with large gains in performance on the minority demographics.
Underspecification Presents Challenges for Credibility in Modern Machine Learning
Nov 06, 2020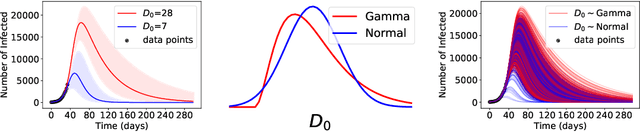


Abstract:ML models often exhibit unexpectedly poor behavior when they are deployed in real-world domains. We identify underspecification as a key reason for these failures. An ML pipeline is underspecified when it can return many predictors with equivalently strong held-out performance in the training domain. Underspecification is common in modern ML pipelines, such as those based on deep learning. Predictors returned by underspecified pipelines are often treated as equivalent based on their training domain performance, but we show here that such predictors can behave very differently in deployment domains. This ambiguity can lead to instability and poor model behavior in practice, and is a distinct failure mode from previously identified issues arising from structural mismatch between training and deployment domains. We show that this problem appears in a wide variety of practical ML pipelines, using examples from computer vision, medical imaging, natural language processing, clinical risk prediction based on electronic health records, and medical genomics. Our results show the need to explicitly account for underspecification in modeling pipelines that are intended for real-world deployment in any domain.
Efficient and Scalable Bayesian Neural Nets with Rank-1 Factors
May 14, 2020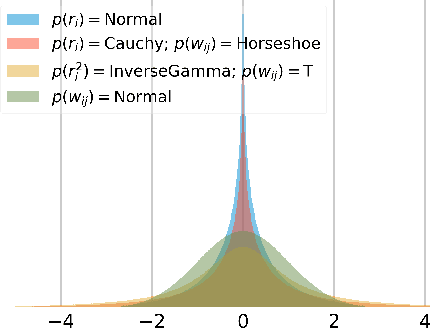
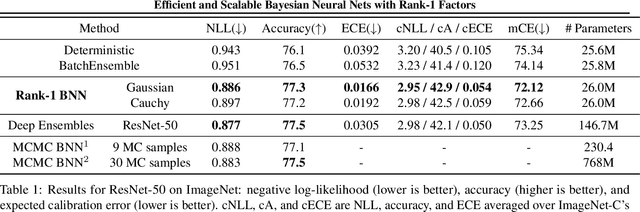
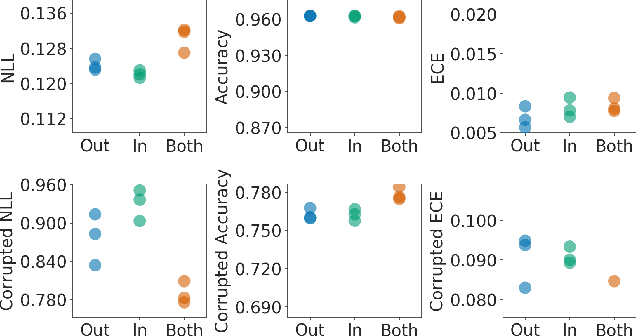
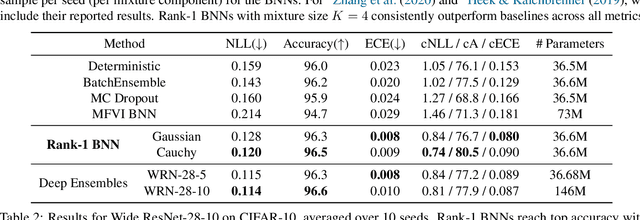
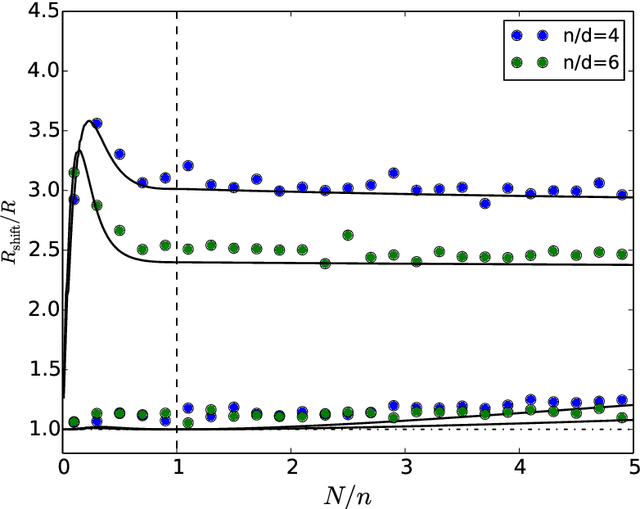
Abstract:Bayesian neural networks (BNNs) demonstrate promising success in improving the robustness and uncertainty quantification of modern deep learning. However, they generally struggle with underfitting at scale and parameter efficiency. On the other hand, deep ensembles have emerged as alternatives for uncertainty quantification that, while outperforming BNNs on certain problems, also suffer from efficiency issues. It remains unclear how to combine the strengths of these two approaches and remediate their common issues. To tackle this challenge, we propose a rank-1 parameterization of BNNs, where each weight matrix involves only a distribution on a rank-1 subspace. We also revisit the use of mixture approximate posteriors to capture multiple modes, where unlike typical mixtures, this approach admits a significantly smaller memory increase (e.g., only a 0.4% increase for a ResNet-50 mixture of size 10). We perform a systematic empirical study on the choices of prior, variational posterior, and methods to improve training. For ResNet-50 on ImageNet, Wide ResNet 28-10 on CIFAR-10/100, and an RNN on MIMIC-III, rank-1 BNNs achieve state-of-the-art performance across log-likelihood, accuracy, and calibration on the test sets and out-of-distribution variants.
Federated and Differentially Private Learning for Electronic Health Records
Nov 13, 2019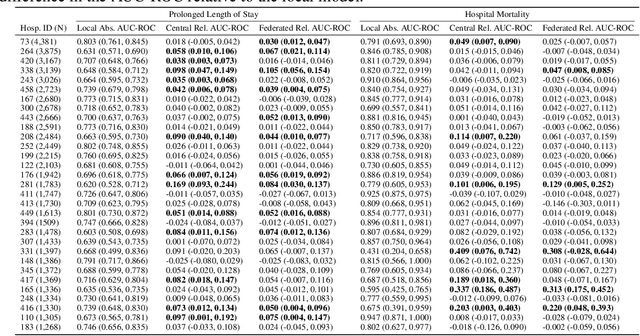
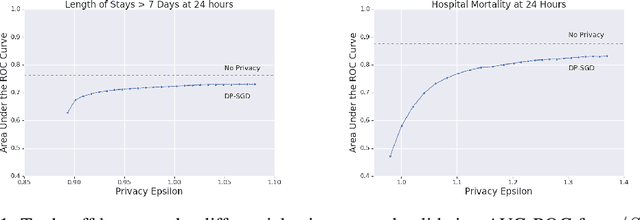
Abstract:The use of collaborative and decentralized machine learning techniques such as federated learning have the potential to enable the development and deployment of clinical risk predictions models in low-resource settings without requiring sensitive data be shared or stored in a central repository. This process necessitates communication of model weights or updates between collaborating entities, but it is unclear to what extent patient privacy is compromised as a result. To gain insight into this question, we study the efficacy of centralized versus federated learning in both private and non-private settings. The clinical prediction tasks we consider are the prediction of prolonged length of stay and in-hospital mortality across thirty one hospitals in the eICU Collaborative Research Database. We find that while it is straightforward to apply differentially private stochastic gradient descent to achieve strong privacy bounds when training in a centralized setting, it is considerably more difficult to do so in the federated setting.
Analyzing the Role of Model Uncertainty for Electronic Health Records
Jun 10, 2019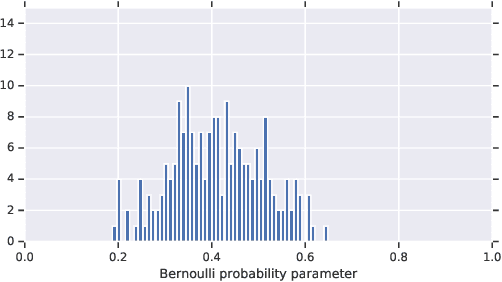
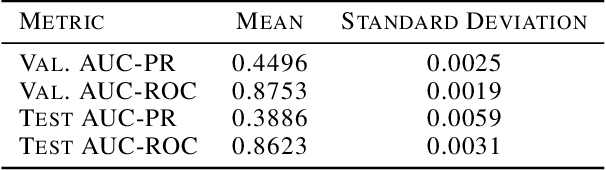
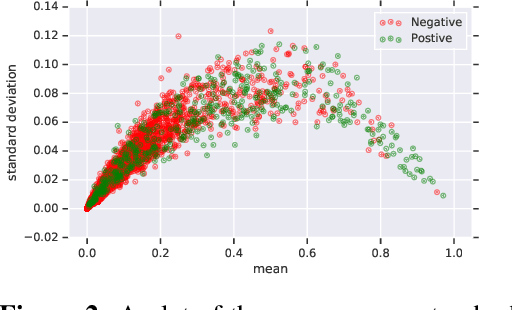
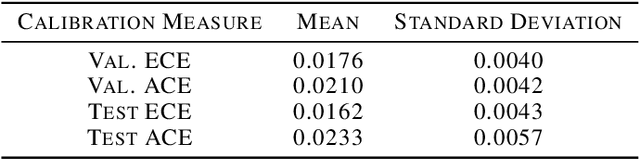
Abstract:In medicine, both ethical and monetary costs of incorrect predictions can be significant, and the complexity of the problems often necessitates increasingly complex models. Recent work has shown that changing just the random seed is enough for otherwise well-tuned deep neural networks to vary in their individual predicted probabilities. In light of this, we investigate the role of model uncertainty methods in the medical domain. Using RNN ensembles and various Bayesian RNNs, we show that population-level metrics, such as AUC-PR, AUC-ROC, log-likelihood, and calibration error, do not capture model uncertainty. Meanwhile, the presence of significant variability in patient-specific predictions and optimal decisions motivates the need for capturing model uncertainty. Understanding the uncertainty for individual patients is an area with clear clinical impact, such as determining when a model decision is likely to be brittle. We further show that RNNs with only Bayesian embeddings can be a more efficient way to capture model uncertainty compared to ensembles, and we analyze how model uncertainty is impacted across individual input features and patient subgroups.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge