Karim Lekadir
Segmentation variability and radiomics stability for predicting Triple-Negative Breast Cancer subtype using Magnetic Resonance Imaging
Apr 02, 2025
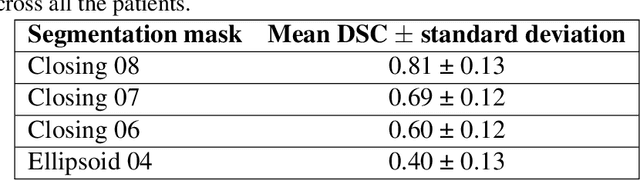
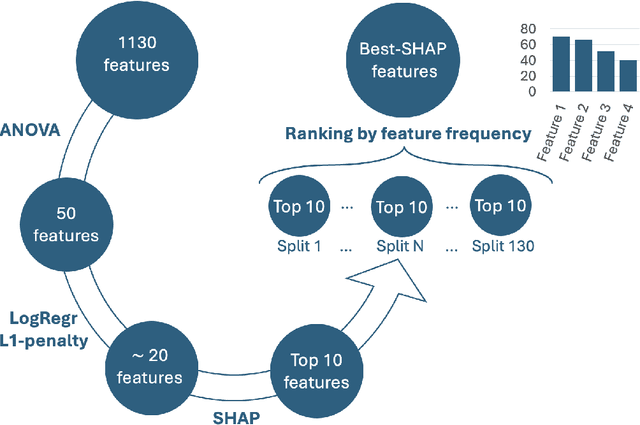
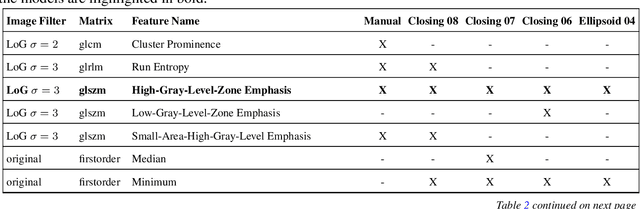
Abstract:Most papers caution against using predictive models for disease stratification based on unselected radiomic features, as these features are affected by contouring variability. Instead, they advocate for the use of the Intraclass Correlation Coefficient (ICC) as a measure of stability for feature selection. However, the direct effect of segmentation variability on the predictive models is rarely studied. This study investigates the impact of segmentation variability on feature stability and predictive performance in radiomics-based prediction of Triple-Negative Breast Cancer (TNBC) subtype using Magnetic Resonance Imaging. A total of 244 images from the Duke dataset were used, with segmentation variability introduced through modifications of manual segmentations. For each mask, explainable radiomic features were selected using the Shapley Additive exPlanations method and used to train logistic regression models. Feature stability across segmentations was assessed via ICC, Pearson's correlation, and reliability scores quantifying the relationship between feature stability and segmentation variability. Results indicate that segmentation accuracy does not significantly impact predictive performance. While incorporating peritumoral information may reduce feature reproducibility, it does not diminish feature predictive capability. Moreover, feature selection in predictive models is not inherently tied to feature stability with respect to segmentation, suggesting that an overreliance on ICC or reliability scores for feature selection might exclude valuable predictive features.
Federated nnU-Net for Privacy-Preserving Medical Image Segmentation
Mar 04, 2025



Abstract:The nnU-Net framework has played a crucial role in medical image segmentation and has become the gold standard in multitudes of applications targeting different diseases, organs, and modalities. However, so far it has been used primarily in a centralized approach where the data collected from hospitals are stored in one center and used to train the nnU-Net. This centralized approach has various limitations, such as leakage of sensitive patient information and violation of patient privacy. Federated learning is one of the approaches to train a segmentation model in a decentralized manner that helps preserve patient privacy. In this paper, we propose FednnU-Net, a federated learning extension of nnU-Net. We introduce two novel federated learning methods to the nnU-Net framework - Federated Fingerprint Extraction (FFE) and Asymmetric Federated Averaging (AsymFedAvg) - and experimentally show their consistent performance for breast, cardiac and fetal segmentation using 6 datasets representing samples from 18 institutions. Additionally, to further promote research and deployment of decentralized training in privacy constrained institutions, we make our plug-n-play framework public. The source-code is available at https://github.com/faildeny/FednnUNet .
Efficient MedSAMs: Segment Anything in Medical Images on Laptop
Dec 20, 2024


Abstract:Promptable segmentation foundation models have emerged as a transformative approach to addressing the diverse needs in medical images, but most existing models require expensive computing, posing a big barrier to their adoption in clinical practice. In this work, we organized the first international competition dedicated to promptable medical image segmentation, featuring a large-scale dataset spanning nine common imaging modalities from over 20 different institutions. The top teams developed lightweight segmentation foundation models and implemented an efficient inference pipeline that substantially reduced computational requirements while maintaining state-of-the-art segmentation accuracy. Moreover, the post-challenge phase advanced the algorithms through the design of performance booster and reproducibility tasks, resulting in improved algorithms and validated reproducibility of the winning solution. Furthermore, the best-performing algorithms have been incorporated into the open-source software with a user-friendly interface to facilitate clinical adoption. The data and code are publicly available to foster the further development of medical image segmentation foundation models and pave the way for impactful real-world applications.
Simulating Dynamic Tumor Contrast Enhancement in Breast MRI using Conditional Generative Adversarial Networks
Sep 27, 2024Abstract:This paper presents a method for virtual contrast enhancement in breast MRI, offering a promising non-invasive alternative to traditional contrast agent-based DCE-MRI acquisition. Using a conditional generative adversarial network, we predict DCE-MRI images, including jointly-generated sequences of multiple corresponding DCE-MRI timepoints, from non-contrast-enhanced MRIs, enabling tumor localization and characterization without the associated health risks. Furthermore, we qualitatively and quantitatively evaluate the synthetic DCE-MRI images, proposing a multi-metric Scaled Aggregate Measure (SAMe), assessing their utility in a tumor segmentation downstream task, and conclude with an analysis of the temporal patterns in multi-sequence DCE-MRI generation. Our approach demonstrates promising results in generating realistic and useful DCE-MRI sequences, highlighting the potential of virtual contrast enhancement for improving breast cancer diagnosis and treatment, particularly for patients where contrast agent administration is contraindicated.
PSFHS Challenge Report: Pubic Symphysis and Fetal Head Segmentation from Intrapartum Ultrasound Images
Sep 17, 2024



Abstract:Segmentation of the fetal and maternal structures, particularly intrapartum ultrasound imaging as advocated by the International Society of Ultrasound in Obstetrics and Gynecology (ISUOG) for monitoring labor progression, is a crucial first step for quantitative diagnosis and clinical decision-making. This requires specialized analysis by obstetrics professionals, in a task that i) is highly time- and cost-consuming and ii) often yields inconsistent results. The utility of automatic segmentation algorithms for biometry has been proven, though existing results remain suboptimal. To push forward advancements in this area, the Grand Challenge on Pubic Symphysis-Fetal Head Segmentation (PSFHS) was held alongside the 26th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI 2023). This challenge aimed to enhance the development of automatic segmentation algorithms at an international scale, providing the largest dataset to date with 5,101 intrapartum ultrasound images collected from two ultrasound machines across three hospitals from two institutions. The scientific community's enthusiastic participation led to the selection of the top 8 out of 179 entries from 193 registrants in the initial phase to proceed to the competition's second stage. These algorithms have elevated the state-of-the-art in automatic PSFHS from intrapartum ultrasound images. A thorough analysis of the results pinpointed ongoing challenges in the field and outlined recommendations for future work. The top solutions and the complete dataset remain publicly available, fostering further advancements in automatic segmentation and biometry for intrapartum ultrasound imaging.
Intrapartum Ultrasound Image Segmentation of Pubic Symphysis and Fetal Head Using Dual Student-Teacher Framework with CNN-ViT Collaborative Learning
Sep 11, 2024Abstract:The segmentation of the pubic symphysis and fetal head (PSFH) constitutes a pivotal step in monitoring labor progression and identifying potential delivery complications. Despite the advances in deep learning, the lack of annotated medical images hinders the training of segmentation. Traditional semi-supervised learning approaches primarily utilize a unified network model based on Convolutional Neural Networks (CNNs) and apply consistency regularization to mitigate the reliance on extensive annotated data. However, these methods often fall short in capturing the discriminative features of unlabeled data and in delineating the long-range dependencies inherent in the ambiguous boundaries of PSFH within ultrasound images. To address these limitations, we introduce a novel framework, the Dual-Student and Teacher Combining CNN and Transformer (DSTCT), which synergistically integrates the capabilities of CNNs and Transformers. Our framework comprises a Vision Transformer (ViT) as the teacher and two student mod ls one ViT and one CNN. This dual-student setup enables mutual supervision through the generation of both hard and soft pseudo-labels, with the consistency in their predictions being refined by minimizing the classifier determinacy discrepancy. The teacher model further reinforces learning within this architecture through the imposition of consistency regularization constraints. To augment the generalization abilities of our approach, we employ a blend of data and model perturbation techniques. Comprehensive evaluations on the benchmark dataset of the PSFH Segmentation Grand Challenge at MICCAI 2023 demonstrate our DSTCT framework outperformed ten contemporary semi-supervised segmentation methods. Code available at https://github.com/jjm1589/DSTCT.
Democratizing AI in Africa: FL for Low-Resource Edge Devices
Aug 30, 2024



Abstract:Africa faces significant challenges in healthcare delivery due to limited infrastructure and access to advanced medical technologies. This study explores the use of federated learning to overcome these barriers, focusing on perinatal health. We trained a fetal plane classifier using perinatal data from five African countries: Algeria, Ghana, Egypt, Malawi, and Uganda, along with data from Spanish hospitals. To incorporate the lack of computational resources in the analysis, we considered a heterogeneous set of devices, including a Raspberry Pi and several laptops, for model training. We demonstrate comparative performance between a centralized and a federated model, despite the compute limitations, and a significant improvement in model generalizability when compared to models trained only locally. These results show the potential for a future implementation at a large scale of a federated learning platform to bridge the accessibility gap and improve model generalizability with very little requirements.
Enhancing the Utility of Privacy-Preserving Cancer Classification using Synthetic Data
Jul 17, 2024Abstract:Deep learning holds immense promise for aiding radiologists in breast cancer detection. However, achieving optimal model performance is hampered by limitations in availability and sharing of data commonly associated to patient privacy concerns. Such concerns are further exacerbated, as traditional deep learning models can inadvertently leak sensitive training information. This work addresses these challenges exploring and quantifying the utility of privacy-preserving deep learning techniques, concretely, (i) differentially private stochastic gradient descent (DP-SGD) and (ii) fully synthetic training data generated by our proposed malignancy-conditioned generative adversarial network. We assess these methods via downstream malignancy classification of mammography masses using a transformer model. Our experimental results depict that synthetic data augmentation can improve privacy-utility tradeoffs in differentially private model training. Further, model pretraining on synthetic data achieves remarkable performance, which can be further increased with DP-SGD fine-tuning across all privacy guarantees. With this first in-depth exploration of privacy-preserving deep learning in breast imaging, we address current and emerging clinical privacy requirements and pave the way towards the adoption of private high-utility deep diagnostic models. Our reproducible codebase is publicly available at https://github.com/RichardObi/mammo_dp.
MAMA-MIA: A Large-Scale Multi-Center Breast Cancer DCE-MRI Benchmark Dataset with Expert Segmentations
Jun 19, 2024Abstract:Current research in breast cancer Magnetic Resonance Imaging (MRI), especially with Artificial Intelligence (AI), faces challenges due to the lack of expert segmentations. To address this, we introduce the MAMA-MIA dataset, comprising 1506 multi-center dynamic contrast-enhanced MRI cases with expert segmentations of primary tumors and non-mass enhancement areas. These cases were sourced from four publicly available collections in The Cancer Imaging Archive (TCIA). Initially, we trained a deep learning model to automatically segment the cases, generating preliminary segmentations that significantly reduced expert segmentation time. Sixteen experts, averaging 9 years of experience in breast cancer, then corrected these segmentations, resulting in the final expert segmentations. Additionally, two radiologists conducted a visual inspection of the automatic segmentations to support future quality control studies. Alongside the expert segmentations, we provide 49 harmonized demographic and clinical variables and the pretrained weights of the well-known nnUNet architecture trained using the DCE-MRI full-images and expert segmentations. This dataset aims to accelerate the development and benchmarking of deep learning models and foster innovation in breast cancer diagnostics and treatment planning.
Mitigating annotation shift in cancer classification using single image generative models
May 30, 2024Abstract:Artificial Intelligence (AI) has emerged as a valuable tool for assisting radiologists in breast cancer detection and diagnosis. However, the success of AI applications in this domain is restricted by the quantity and quality of available data, posing challenges due to limited and costly data annotation procedures that often lead to annotation shifts. This study simulates, analyses and mitigates annotation shifts in cancer classification in the breast mammography domain. First, a high-accuracy cancer risk prediction model is developed, which effectively distinguishes benign from malignant lesions. Next, model performance is used to quantify the impact of annotation shift. We uncover a substantial impact of annotation shift on multiclass classification performance particularly for malignant lesions. We thus propose a training data augmentation approach based on single-image generative models for the affected class, requiring as few as four in-domain annotations to considerably mitigate annotation shift, while also addressing dataset imbalance. Lastly, we further increase performance by proposing and validating an ensemble architecture based on multiple models trained under different data augmentation regimes. Our study offers key insights into annotation shift in deep learning breast cancer classification and explores the potential of single-image generative models to overcome domain shift challenges.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge