Polyxeni Gkontra
Med-DualLoRA: Local Adaptation of Foundation Models for 3D Cardiac MRI
Mar 11, 2026Abstract:Foundation models (FMs) show great promise for robust downstream performance across medical imaging tasks and modalities, including cardiac magnetic resonance (CMR), following task-specific adaptation. However, adaptation using single-site data may lead to suboptimal performance and increased model bias, while centralized fine-tuning on clinical data is often infeasible due to privacy constraints. Federated fine-tuning offers a privacy-preserving alternative; yet conventional approaches struggle under heterogeneous, non-IID multi-center data and incur substantial communication overhead when adapting large models. In this work, we study federated FM fine-tuning for 3D CMR disease detection and propose Med-DualLoRA, a client-aware parameter-efficient fine-tuning (PEFT) federated framework that disentangles globally shared and local low-rank adaptations (LoRA) through additive decomposition. Global and local LoRA modules are trained locally, but only the global component is shared and aggregated across sites, keeping local adapters private. This design improves personalization while significantly reducing communication cost, and experiments show that adapting only two transformer blocks preserves performance while further improving efficiency. We evaluate our method on a multi-center state-of-the-art cine 3D CMR FM fine-tuned for disease detection using ACDC and combined M\&Ms datasets, treating each vendor as a federated client. Med-DualLoRA achieves statistically significant improved performance (balanced accuracy 0.768, specificity 0.612) compared to other federated PEFT baselines, while maintaining communication efficiency. Our approach provides a scalable solution for local federated adaptation of medical FMs under realistic clinical constraints.
Debiasing Cardiac Imaging with Controlled Latent Diffusion Models
Mar 28, 2024



Abstract:The progress in deep learning solutions for disease diagnosis and prognosis based on cardiac magnetic resonance imaging is hindered by highly imbalanced and biased training data. To address this issue, we propose a method to alleviate imbalances inherent in datasets through the generation of synthetic data based on sensitive attributes such as sex, age, body mass index, and health condition. We adopt ControlNet based on a denoising diffusion probabilistic model to condition on text assembled from patient metadata and cardiac geometry derived from segmentation masks using a large-cohort study, specifically, the UK Biobank. We assess our method by evaluating the realism of the generated images using established quantitative metrics. Furthermore, we conduct a downstream classification task aimed at debiasing a classifier by rectifying imbalances within underrepresented groups through synthetically generated samples. Our experiments demonstrate the effectiveness of the proposed approach in mitigating dataset imbalances, such as the scarcity of younger patients or individuals with normal BMI level suffering from heart failure. This work represents a major step towards the adoption of synthetic data for the development of fair and generalizable models for medical classification tasks. Notably, we conduct all our experiments using a single, consumer-level GPU to highlight the feasibility of our approach within resource-constrained environments. Our code is available at https://github.com/faildeny/debiasing-cardiac-mri.
Federated Learning for Multi-Center Imaging Diagnostics: A Study in Cardiovascular Disease
Jul 07, 2021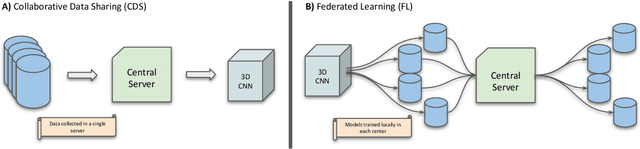
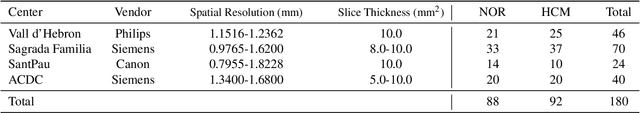
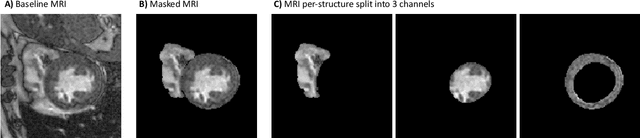
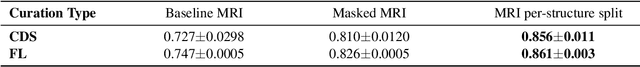
Abstract:Deep learning models can enable accurate and efficient disease diagnosis, but have thus far been hampered by the data scarcity present in the medical world. Automated diagnosis studies have been constrained by underpowered single-center datasets, and although some results have shown promise, their generalizability to other institutions remains questionable as the data heterogeneity between institutions is not taken into account. By allowing models to be trained in a distributed manner that preserves patients' privacy, federated learning promises to alleviate these issues, by enabling diligent multi-center studies. We present the first federated learning study on the modality of cardiovascular magnetic resonance (CMR) and use four centers derived from subsets of the M\&M and ACDC datasets, focusing on the diagnosis of hypertrophic cardiomyopathy (HCM). We adapt a 3D-CNN network pretrained on action recognition and explore two different ways of incorporating shape prior information to the model, and four different data augmentation set-ups, systematically analyzing their impact on the different collaborative learning choices. We show that despite the small size of data (180 subjects derived from four centers), the privacy preserving federated learning achieves promising results that are competitive with traditional centralized learning. We further find that federatively trained models exhibit increased robustness and are more sensitive to domain shift effects.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge