Michael Rutherford
Medical Image De-Identification Benchmark Challenge
Jul 31, 2025



Abstract:The de-identification (deID) of protected health information (PHI) and personally identifiable information (PII) is a fundamental requirement for sharing medical images, particularly through public repositories, to ensure compliance with patient privacy laws. In addition, preservation of non-PHI metadata to inform and enable downstream development of imaging artificial intelligence (AI) is an important consideration in biomedical research. The goal of MIDI-B was to provide a standardized platform for benchmarking of DICOM image deID tools based on a set of rules conformant to the HIPAA Safe Harbor regulation, the DICOM Attribute Confidentiality Profiles, and best practices in preservation of research-critical metadata, as defined by The Cancer Imaging Archive (TCIA). The challenge employed a large, diverse, multi-center, and multi-modality set of real de-identified radiology images with synthetic PHI/PII inserted. The MIDI-B Challenge consisted of three phases: training, validation, and test. Eighty individuals registered for the challenge. In the training phase, we encouraged participants to tune their algorithms using their in-house or public data. The validation and test phases utilized the DICOM images containing synthetic identifiers (of 216 and 322 subjects, respectively). Ten teams successfully completed the test phase of the challenge. To measure success of a rule-based approach to image deID, scores were computed as the percentage of correct actions from the total number of required actions. The scores ranged from 97.91% to 99.93%. Participants employed a variety of open-source and proprietary tools with customized configurations, large language models, and optical character recognition (OCR). In this paper we provide a comprehensive report on the MIDI-B Challenge's design, implementation, results, and lessons learned.
medigan: A Python Library of Pretrained Generative Models for Enriched Data Access in Medical Imaging
Sep 28, 2022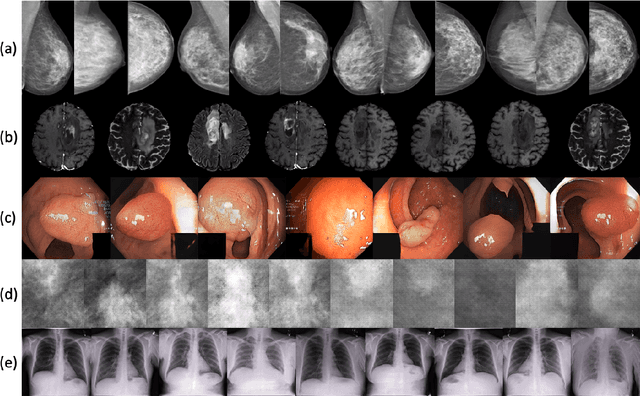
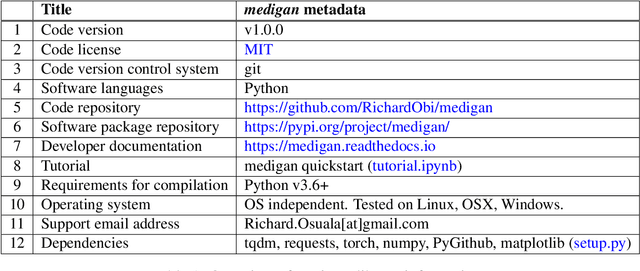
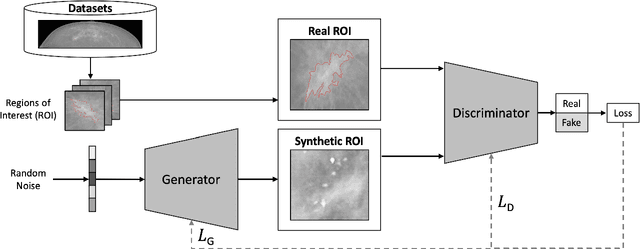
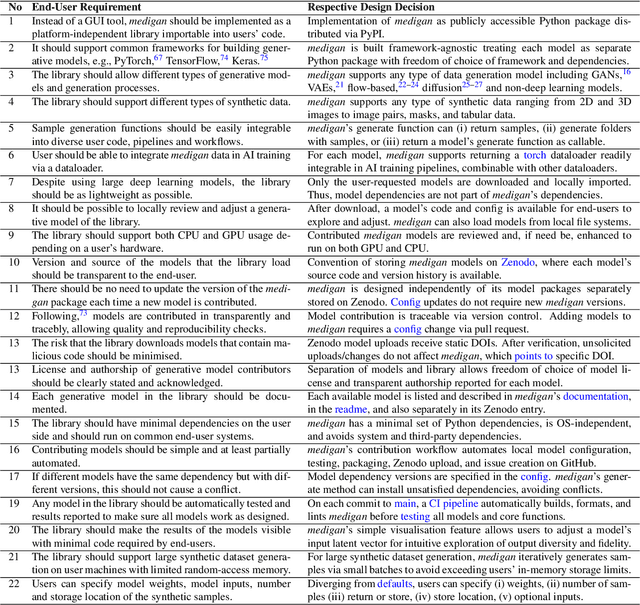
Abstract:Synthetic data generated by generative models can enhance the performance and capabilities of data-hungry deep learning models in medical imaging. However, there is (1) limited availability of (synthetic) datasets and (2) generative models are complex to train, which hinders their adoption in research and clinical applications. To reduce this entry barrier, we propose medigan, a one-stop shop for pretrained generative models implemented as an open-source framework-agnostic Python library. medigan allows researchers and developers to create, increase, and domain-adapt their training data in just a few lines of code. Guided by design decisions based on gathered end-user requirements, we implement medigan based on modular components for generative model (i) execution, (ii) visualisation, (iii) search & ranking, and (iv) contribution. The library's scalability and design is demonstrated by its growing number of integrated and readily-usable pretrained generative models consisting of 21 models utilising 9 different Generative Adversarial Network architectures trained on 11 datasets from 4 domains, namely, mammography, endoscopy, x-ray, and MRI. Furthermore, 3 applications of medigan are analysed in this work, which include (a) enabling community-wide sharing of restricted data, (b) investigating generative model evaluation metrics, and (c) improving clinical downstream tasks. In (b), extending on common medical image synthesis assessment and reporting standards, we show Fr\'echet Inception Distance variability based on image normalisation and radiology-specific feature extraction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge