Kai Ma
School of Electrical Engineering, Yanshan University, Qinhuangdao, China
Transport based Graph Kernels
Nov 02, 2020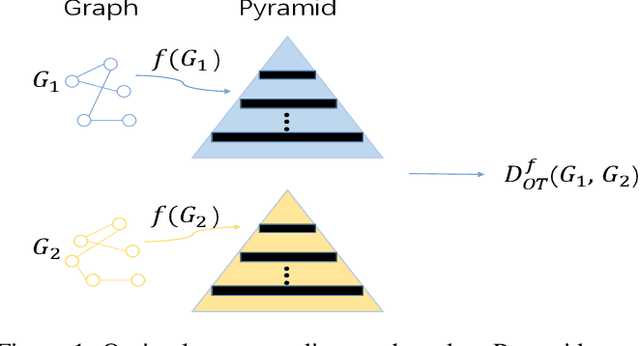
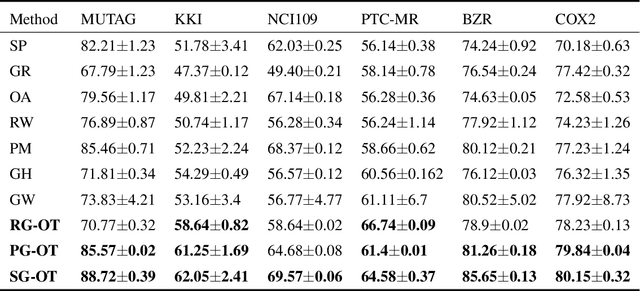
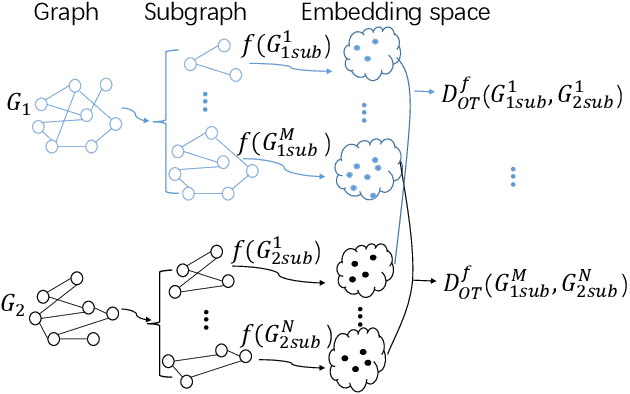
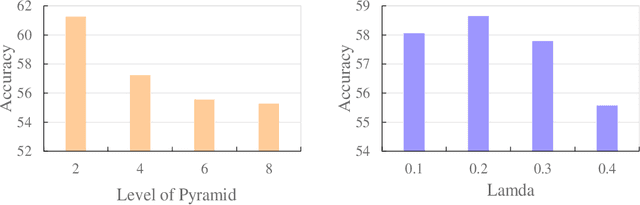
Abstract:Graph kernel is a powerful tool measuring the similarity between graphs. Most of the existing graph kernels focused on node labels or attributes and ignored graph hierarchical structure information. In order to effectively utilize graph hierarchical structure information, we propose pyramid graph kernel based on optimal transport (OT). Each graph is embedded into hierarchical structures of the pyramid. Then, the OT distance is utilized to measure the similarity between graphs in hierarchical structures. We also utilize the OT distance to measure the similarity between subgraphs and propose subgraph kernel based on OT. The positive semidefinite (p.s.d) of graph kernels based on optimal transport distance is not necessarily possible. We further propose regularized graph kernel based on OT where we add the kernel regularization to the original optimal transport distance to obtain p.s.d kernel matrix. We evaluate the proposed graph kernels on several benchmark classification tasks and compare their performance with the existing state-of-the-art graph kernels. In most cases, our proposed graph kernel algorithms outperform the competing methods.
Ordinal Pattern Kernel for Brain Connectivity Network Classification
Aug 18, 2020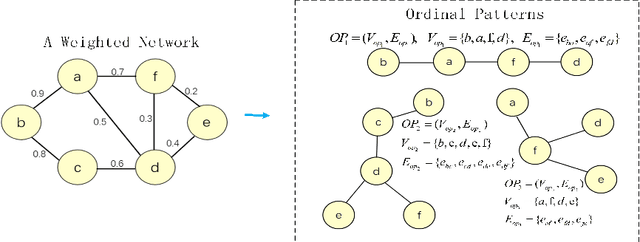
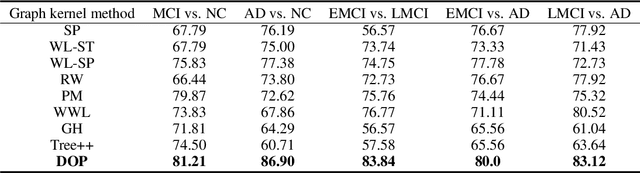
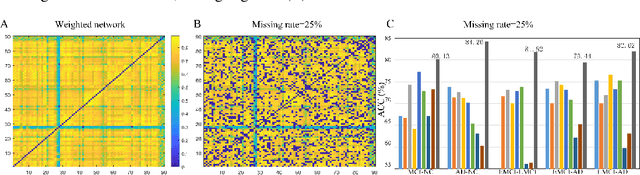
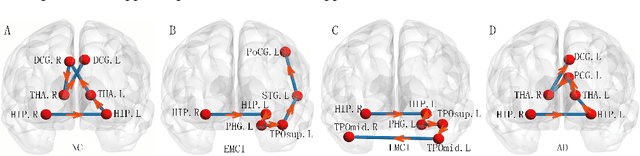
Abstract:Brain connectivity networks, which characterize the functional or structural interaction of brain regions, has been widely used for brain disease classification. Kernel-based method, such as graph kernel (i.e., kernel defined on graphs), has been proposed for measuring the similarity of brain networks, and yields the promising classification performance. However, most of graph kernels are built on unweighted graph (i.e., network) with edge present or not, and neglecting the valuable weight information of edges in brain connectivity network, with edge weights conveying the strengths of temporal correlation or fiber connection between brain regions. Accordingly, in this paper, we present an ordinal pattern kernel for brain connectivity network classification. Different with existing graph kernels that measures the topological similarity of unweighted graphs, the proposed ordinal pattern kernels calculate the similarity of weighted networks by comparing ordinal patterns from weighted networks. To evaluate the effectiveness of the proposed ordinal kernel, we further develop a depth-first-based ordinal pattern kernel, and perform extensive experiments in a real dataset of brain disease from ADNI database. The results demonstrate that our proposed ordinal pattern kernel can achieve better classification performance compared with state-of-the-art graph kernels.
MI^2GAN: Generative Adversarial Network for Medical Image Domain Adaptation using Mutual Information Constraint
Jul 30, 2020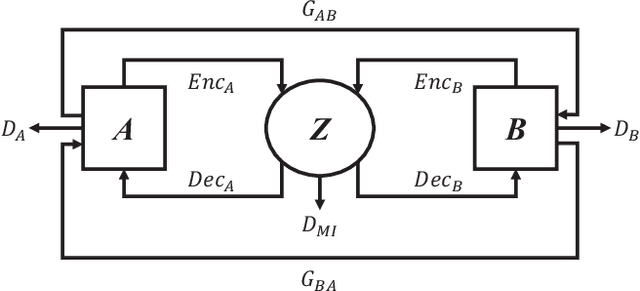
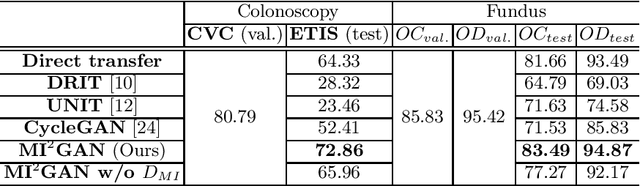
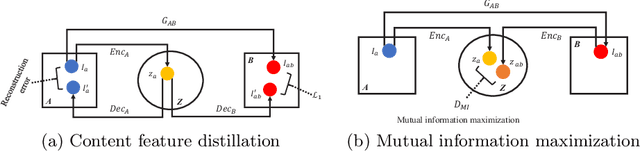
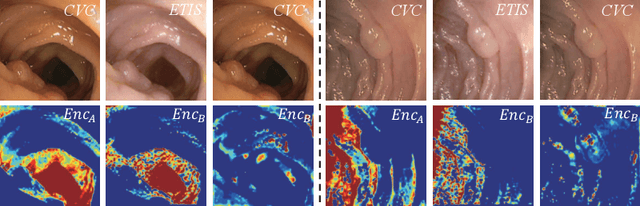
Abstract:Domain shift between medical images from multicentres is still an open question for the community, which degrades the generalization performance of deep learning models. Generative adversarial network (GAN), which synthesize plausible images, is one of the potential solutions to address the problem. However, the existing GAN-based approaches are prone to fail at preserving image-objects in image-to-image (I2I) translation, which reduces their practicality on domain adaptation tasks. In this paper, we propose a novel GAN (namely MI$^2$GAN) to maintain image-contents during cross-domain I2I translation. Particularly, we disentangle the content features from domain information for both the source and translated images, and then maximize the mutual information between the disentangled content features to preserve the image-objects. The proposed MI$^2$GAN is evaluated on two tasks---polyp segmentation using colonoscopic images and the segmentation of optic disc and cup in fundus images. The experimental results demonstrate that the proposed MI$^2$GAN can not only generate elegant translated images, but also significantly improve the generalization performance of widely used deep learning networks (e.g., U-Net).
TR-GAN: Topology Ranking GAN with Triplet Loss for Retinal Artery/Vein Classification
Jul 29, 2020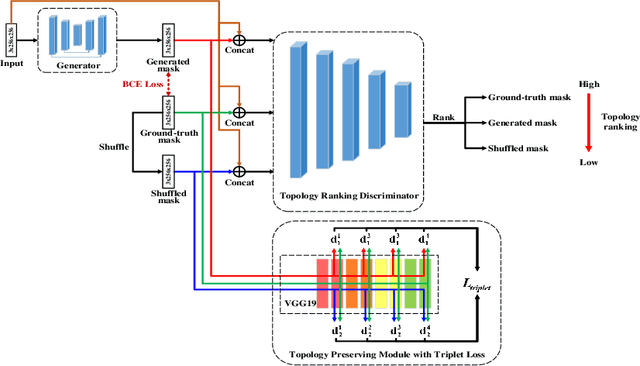
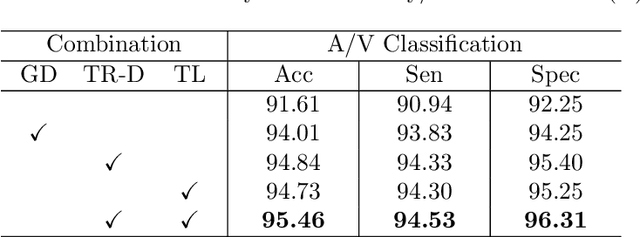
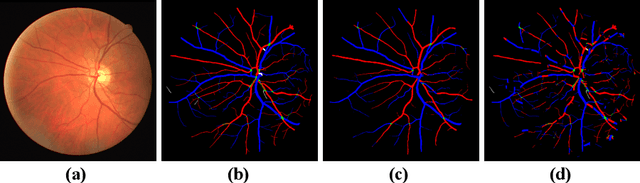
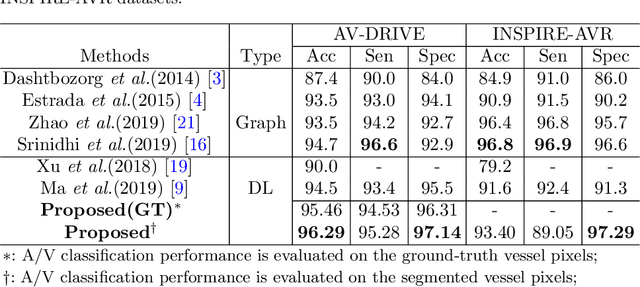
Abstract:Retinal artery/vein (A/V) classification lays the foundation for the quantitative analysis of retinal vessels, which is associated with potential risks of various cardiovascular and cerebral diseases. The topological connection relationship, which has been proved effective in improving the A/V classification performance for the conventional graph based method, has not been exploited by the deep learning based method. In this paper, we propose a Topology Ranking Generative Adversarial Network (TR-GAN) to improve the topology connectivity of the segmented arteries and veins, and further to boost the A/V classification performance. A topology ranking discriminator based on ordinal regression is proposed to rank the topological connectivity level of the ground-truth, the generated A/V mask and the intentionally shuffled mask. The ranking loss is further back-propagated to the generator to generate better connected A/V masks. In addition, a topology preserving module with triplet loss is also proposed to extract the high-level topological features and further to narrow the feature distance between the predicted A/V mask and the ground-truth. The proposed framework effectively increases the topological connectivity of the predicted A/V masks and achieves state-of-the-art A/V classification performance on the publicly available AV-DRIVE dataset.
Difficulty-aware Glaucoma Classification with Multi-Rater Consensus Modeling
Jul 29, 2020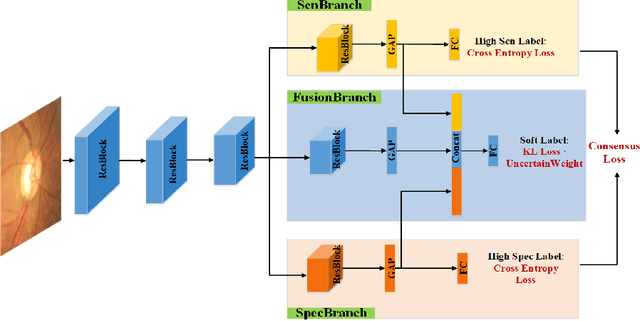
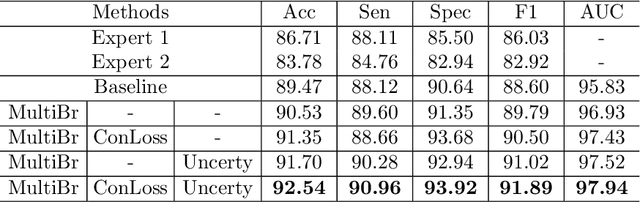
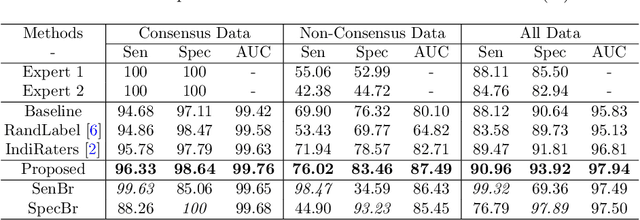
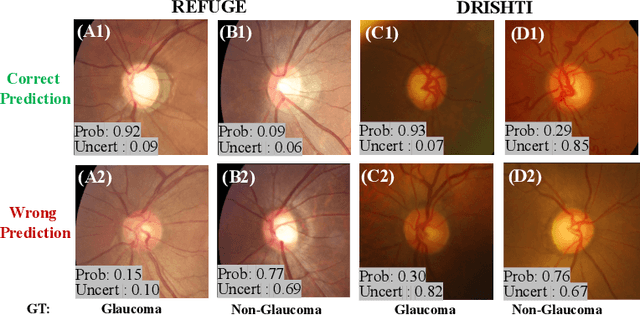
Abstract:Medical images are generally labeled by multiple experts before the final ground-truth labels are determined. Consensus or disagreement among experts regarding individual images reflects the gradeability and difficulty levels of the image. However, when being used for model training, only the final ground-truth label is utilized, while the critical information contained in the raw multi-rater gradings regarding the image being an easy/hard case is discarded. In this paper, we aim to take advantage of the raw multi-rater gradings to improve the deep learning model performance for the glaucoma classification task. Specifically, a multi-branch model structure is proposed to predict the most sensitive, most specifical and a balanced fused result for the input images. In order to encourage the sensitivity branch and specificity branch to generate consistent results for consensus labels and opposite results for disagreement labels, a consensus loss is proposed to constrain the output of the two branches. Meanwhile, the consistency/inconsistency between the prediction results of the two branches implies the image being an easy/hard case, which is further utilized to encourage the balanced fusion branch to concentrate more on the hard cases. Compared with models trained only with the final ground-truth labels, the proposed method using multi-rater consensus information has achieved superior performance, and it is also able to estimate the difficulty levels of individual input images when making the prediction.
Learning Crisp Edge Detector Using Logical Refinement Network
Jul 24, 2020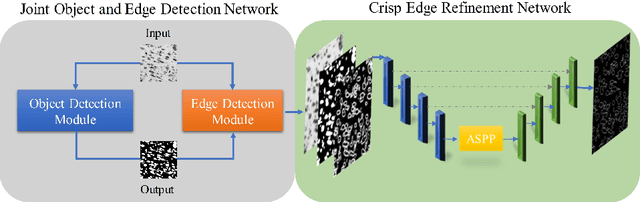
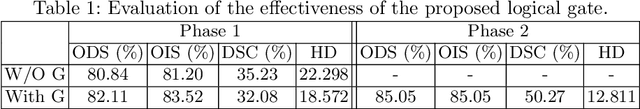
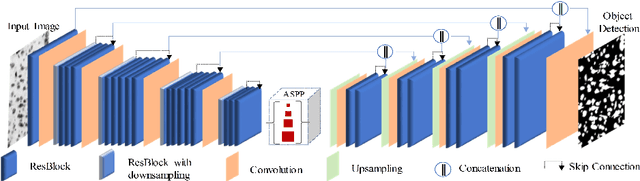
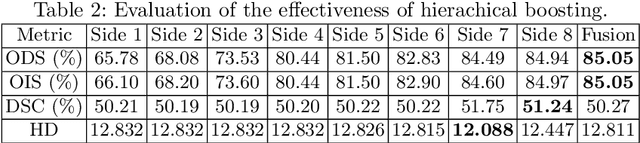
Abstract:Edge detection is a fundamental problem in different computer vision tasks. Recently, edge detection algorithms achieve satisfying improvement built upon deep learning. Although most of them report favorable evaluation scores, they often fail to accurately localize edges and give thick and blurry boundaries. In addition, most of them focus on 2D images and the challenging 3D edge detection is still under-explored. In this work, we propose a novel logical refinement network for crisp edge detection, which is motivated by the logical relationship between segmentation and edge maps and can be applied to both 2D and 3D images. The network consists of a joint object and edge detection network and a crisp edge refinement network, which predicts more accurate, clearer and thinner high quality binary edge maps without any post-processing. Extensive experiments are conducted on the 2D nuclei images from Kaggle 2018 Data Science Bowl and a private 3D microscopy images of a monkey brain, which show outstanding performance compared with state-of-the-art methods.
Leveraging Undiagnosed Data for Glaucoma Classification with Teacher-Student Learning
Jul 22, 2020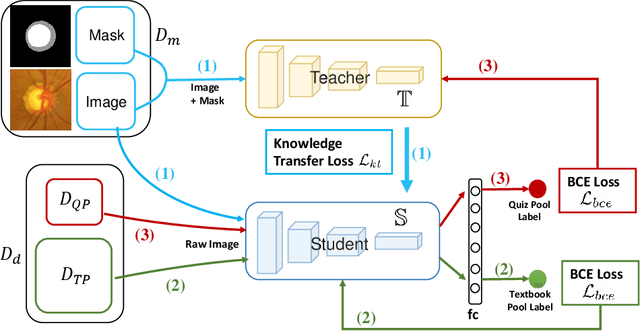
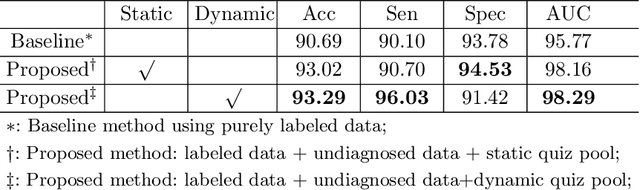
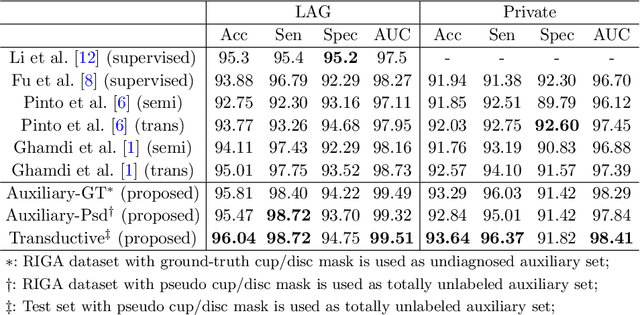
Abstract:Recently, deep learning has been adopted to the glaucoma classification task with performance comparable to that of human experts. However, a well trained deep learning model demands a large quantity of properly labeled data, which is relatively expensive since the accurate labeling of glaucoma requires years of specialist training. In order to alleviate this problem, we propose a glaucoma classification framework which takes advantage of not only the properly labeled images, but also undiagnosed images without glaucoma labels. To be more specific, the proposed framework is adapted from the teacher-student-learning paradigm. The teacher model encodes the wrapped information of undiagnosed images to a latent feature space, meanwhile the student model learns from the teacher through knowledge transfer to improve the glaucoma classification. For the model training procedure, we propose a novel training strategy that simulates the real-world teaching practice named as 'Learning To Teach with Knowledge Transfer (L2T-KT)', and establish a 'Quiz Pool' as the teacher's optimization target. Experiments show that the proposed framework is able to utilize the undiagnosed data effectively to improve the glaucoma prediction performance.
Instance-aware Self-supervised Learning for Nuclei Segmentation
Jul 22, 2020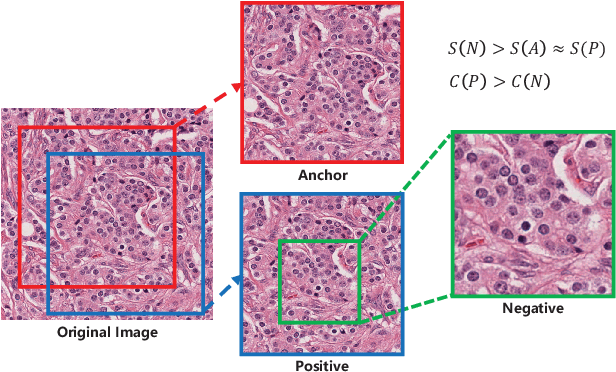
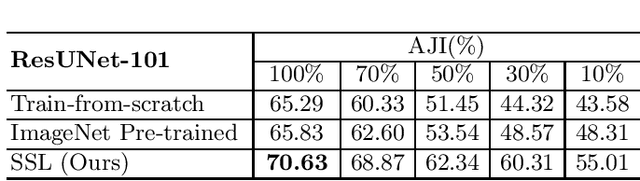
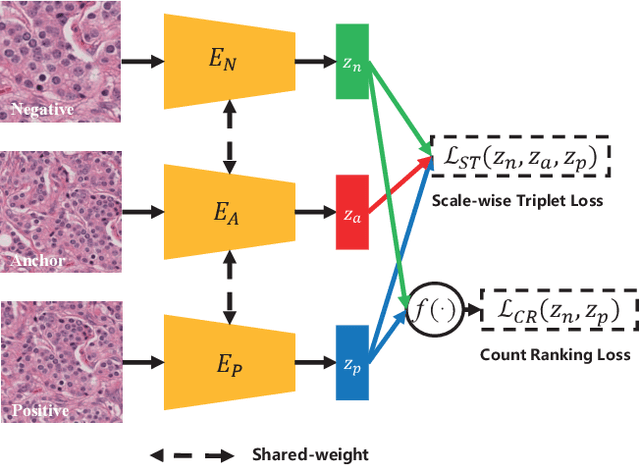
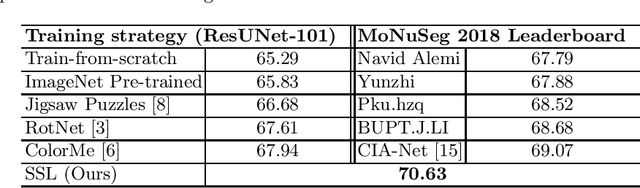
Abstract:Due to the wide existence and large morphological variances of nuclei, accurate nuclei instance segmentation is still one of the most challenging tasks in computational pathology. The annotating of nuclei instances, requiring experienced pathologists to manually draw the contours, is extremely laborious and expensive, which often results in the deficiency of annotated data. The deep learning based segmentation approaches, which highly rely on the quantity of training data, are difficult to fully demonstrate their capacity in this area. In this paper, we propose a novel self-supervised learning framework to deeply exploit the capacity of widely-used convolutional neural networks (CNNs) on the nuclei instance segmentation task. The proposed approach involves two sub-tasks (i.e., scale-wise triplet learning and count ranking), which enable neural networks to implicitly leverage the prior-knowledge of nuclei size and quantity, and accordingly mine the instance-aware feature representations from the raw data. Experimental results on the publicly available MoNuSeg dataset show that the proposed self-supervised learning approach can remarkably boost the segmentation accuracy of nuclei instance---a new state-of-the-art average Aggregated Jaccard Index (AJI) of 70.63%, is achieved by our self-supervised ResUNet-101. To our best knowledge, this is the first work focusing on the self-supervised learning for instance segmentation.
Comparing to Learn: Surpassing ImageNet Pretraining on Radiographs By Comparing Image Representations
Jul 22, 2020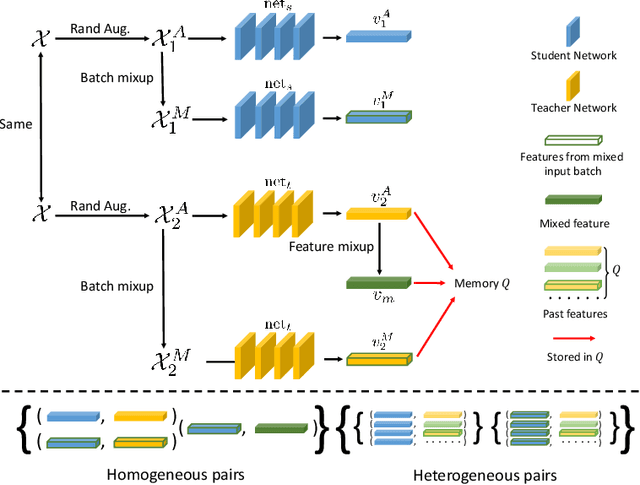
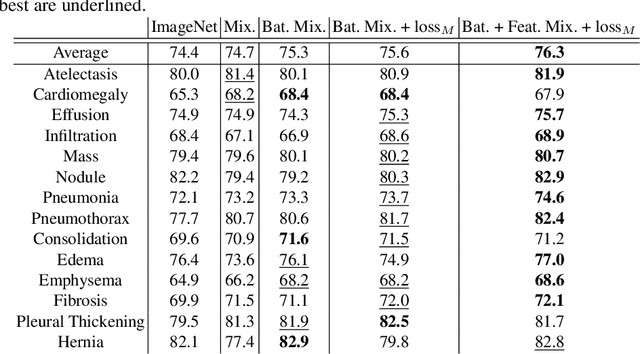

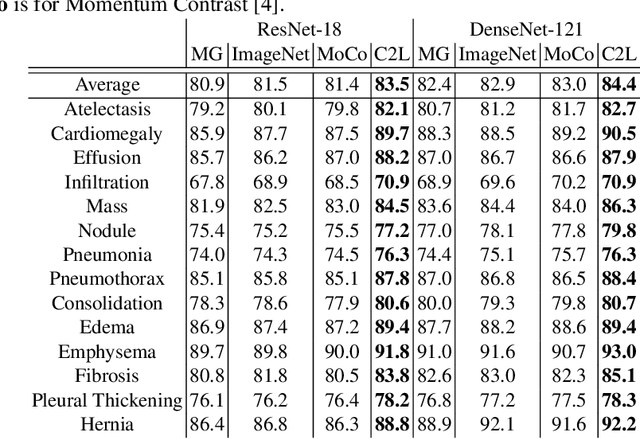
Abstract:In deep learning era, pretrained models play an important role in medical image analysis, in which ImageNet pretraining has been widely adopted as the best way. However, it is undeniable that there exists an obvious domain gap between natural images and medical images. To bridge this gap, we propose a new pretraining method which learns from 700k radiographs given no manual annotations. We call our method as Comparing to Learn (C2L) because it learns robust features by comparing different image representations. To verify the effectiveness of C2L, we conduct comprehensive ablation studies and evaluate it on different tasks and datasets. The experimental results on radiographs show that C2L can outperform ImageNet pretraining and previous state-of-the-art approaches significantly. Code and models are available.
GREEN: a Graph REsidual rE-ranking Network for Grading Diabetic Retinopathy
Jul 21, 2020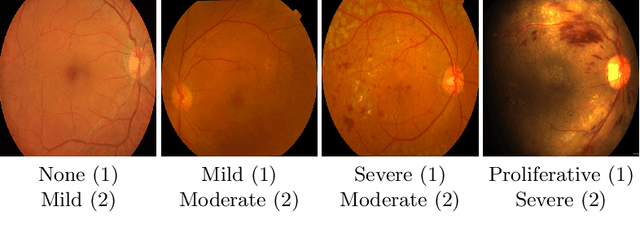
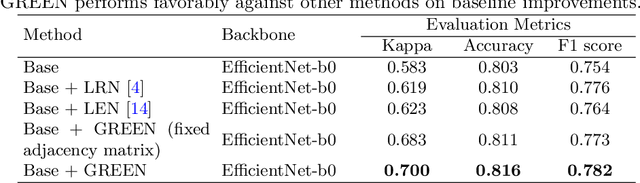
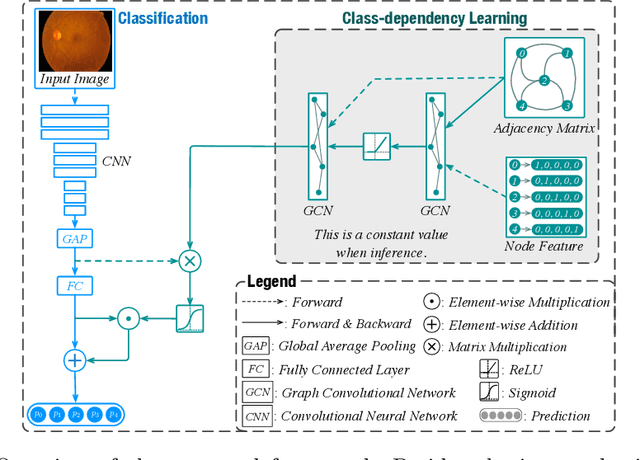
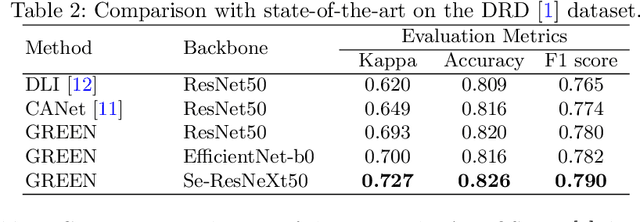
Abstract:The automatic grading of diabetic retinopathy (DR) facilitates medical diagnosis for both patients and physicians. Existing researches formulate DR grading as an image classification problem. As the stages/categories of DR correlate with each other, the relationship between different classes cannot be explicitly described via a one-hot label because it is empirically estimated by different physicians with different outcomes. This class correlation limits existing networks to achieve effective classification. In this paper, we propose a Graph REsidual rE-ranking Network (GREEN) to introduce a class dependency prior into the original image classification network. The class dependency prior is represented by a graph convolutional network with an adjacency matrix. This prior augments image classification pipeline by re-ranking classification results in a residual aggregation manner. Experiments on the standard benchmarks have shown that GREEN performs favorably against state-of-the-art approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge