Junda Wang
ChatThero: An LLM-Supported Chatbot for Behavior Change and Therapeutic Support in Addiction Recovery
Aug 28, 2025
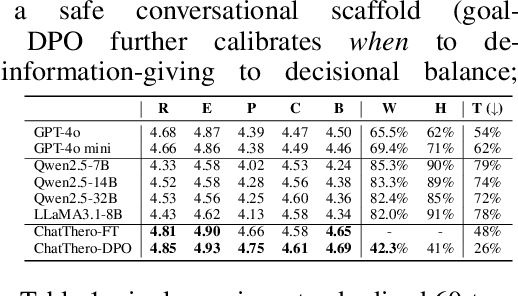
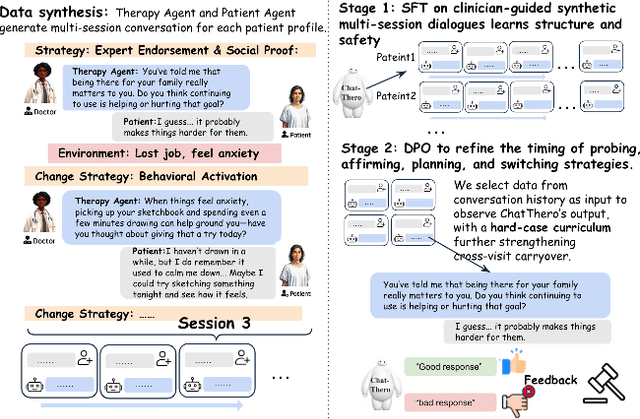

Abstract:Substance use disorders (SUDs) affect over 36 million people worldwide, yet few receive effective care due to stigma, motivational barriers, and limited personalized support. Although large language models (LLMs) show promise for mental-health assistance, most systems lack tight integration with clinically validated strategies, reducing effectiveness in addiction recovery. We present ChatThero, a multi-agent conversational framework that couples dynamic patient modeling with context-sensitive therapeutic dialogue and adaptive persuasive strategies grounded in cognitive behavioral therapy (CBT) and motivational interviewing (MI). We build a high-fidelity synthetic benchmark spanning Easy, Medium, and Hard resistance levels, and train ChatThero with a two-stage pipeline comprising supervised fine-tuning (SFT) followed by direct preference optimization (DPO). In evaluation, ChatThero yields a 41.5\% average gain in patient motivation, a 0.49\% increase in treatment confidence, and resolves hard cases with 26\% fewer turns than GPT-4o, and both automated and human clinical assessments rate it higher in empathy, responsiveness, and behavioral realism. The framework supports rigorous, privacy-preserving study of therapeutic conversation and provides a robust, replicable basis for research and clinical translation.
Incremental Online Learning of Randomized Neural Network with Forward Regularization
Dec 17, 2024Abstract:Online learning of deep neural networks suffers from challenges such as hysteretic non-incremental updating, increasing memory usage, past retrospective retraining, and catastrophic forgetting. To alleviate these drawbacks and achieve progressive immediate decision-making, we propose a novel Incremental Online Learning (IOL) process of Randomized Neural Networks (Randomized NN), a framework facilitating continuous improvements to Randomized NN performance in restrictive online scenarios. Within the framework, we further introduce IOL with ridge regularization (-R) and IOL with forward regularization (-F). -R generates stepwise incremental updates without retrospective retraining and avoids catastrophic forgetting. Moreover, we substituted -R with -F as it enhanced precognition learning ability using semi-supervision and realized better online regrets to offline global experts compared to -R during IOL. The algorithms of IOL for Randomized NN with -R/-F on non-stationary batch stream were derived respectively, featuring recursive weight updates and variable learning rates. Additionally, we conducted a detailed analysis and theoretically derived relative cumulative regret bounds of the Randomized NN learners with -R/-F in IOL under adversarial assumptions using a novel methodology and presented several corollaries, from which we observed the superiority on online learning acceleration and regret bounds of employing -F in IOL. Finally, our proposed methods were rigorously examined across regression and classification tasks on diverse datasets, which distinctly validated the efficacy of IOL frameworks of Randomized NN and the advantages of forward regularization.
RARE: Retrieval-Augmented Reasoning Enhancement for Large Language Models
Dec 05, 2024Abstract:This work introduces RARE (Retrieval-Augmented Reasoning Enhancement), a versatile extension to the mutual reasoning framework (rStar), aimed at enhancing reasoning accuracy and factual integrity across large language models (LLMs) for complex, knowledge-intensive tasks such as commonsense and medical reasoning. RARE incorporates two innovative actions within the Monte Carlo Tree Search (MCTS) framework: A6, which generates search queries based on the initial problem statement, performs information retrieval using those queries, and augments reasoning with the retrieved data to formulate the final answer; and A7, which leverages information retrieval specifically for generated sub-questions and re-answers these sub-questions with the relevant contextual information. Additionally, a Retrieval-Augmented Factuality Scorer is proposed to replace the original discriminator, prioritizing reasoning paths that meet high standards of factuality. Experimental results with LLaMA 3.1 show that RARE enables open-source LLMs to achieve competitive performance with top open-source models like GPT-4 and GPT-4o. This research establishes RARE as a scalable solution for improving LLMs in domains where logical coherence and factual integrity are critical.
CRTRE: Causal Rule Generation with Target Trial Emulation Framework
Nov 10, 2024



Abstract:Causal inference and model interpretability are gaining increasing attention, particularly in the biomedical domain. Despite recent advance, decorrelating features in nonlinear environments with human-interpretable representations remains underexplored. In this study, we introduce a novel method called causal rule generation with target trial emulation framework (CRTRE), which applies randomize trial design principles to estimate the causal effect of association rules. We then incorporate such association rules for the downstream applications such as prediction of disease onsets. Extensive experiments on six healthcare datasets, including synthetic data, real-world disease collections, and MIMIC-III/IV, demonstrate the model's superior performance. Specifically, our method achieved a $\beta$ error of 0.907, outperforming DWR (1.024) and SVM (1.141). On real-world datasets, our model achieved accuracies of 0.789, 0.920, and 0.300 for Esophageal Cancer, Heart Disease, and Cauda Equina Syndrome prediction task, respectively, consistently surpassing baseline models. On the ICD code prediction tasks, it achieved AUC Macro scores of 92.8 on MIMIC-III and 96.7 on MIMIC-IV, outperforming the state-of-the-art models KEPT and MSMN. Expert evaluations further validate the model's effectiveness, causality, and interpretability.
Data Extraction Attacks in Retrieval-Augmented Generation via Backdoors
Nov 03, 2024



Abstract:Despite significant advancements, large language models (LLMs) still struggle with providing accurate answers when lacking domain-specific or up-to-date knowledge. Retrieval-Augmented Generation (RAG) addresses this limitation by incorporating external knowledge bases, but it also introduces new attack surfaces. In this paper, we investigate data extraction attacks targeting the knowledge databases of RAG systems. We demonstrate that previous attacks on RAG largely depend on the instruction-following capabilities of LLMs, and that simple fine-tuning can reduce the success rate of such attacks to nearly zero. This makes these attacks impractical since fine-tuning is a common practice when deploying LLMs in specific domains. To further reveal the vulnerability, we propose to backdoor RAG, where a small portion of poisoned data is injected during the fine-tuning phase to create a backdoor within the LLM. When this compromised LLM is integrated into a RAG system, attackers can exploit specific triggers in prompts to manipulate the LLM to leak documents from the retrieval database. By carefully designing the poisoned data, we achieve both verbatim and paraphrased document extraction. We show that with only 3\% poisoned data, our method achieves an average success rate of 79.7\% in verbatim extraction on Llama2-7B, with a ROUGE-L score of 64.21, and a 68.6\% average success rate in paraphrased extraction, with an average ROUGE score of 52.6 across four datasets. These results underscore the privacy risks associated with the supply chain when deploying RAG systems.
LEAF: Learning and Evaluation Augmented by Fact-Checking to Improve Factualness in Large Language Models
Oct 31, 2024



Abstract:Large language models (LLMs) have shown remarkable capabilities in various natural language processing tasks, yet they often struggle with maintaining factual accuracy, particularly in knowledge-intensive domains like healthcare. This study introduces LEAF: Learning and Evaluation Augmented by Fact-Checking, a novel approach designed to enhance the factual reliability of LLMs, with a focus on medical question answering (QA). LEAF utilizes a dual strategy to enhance the factual accuracy of responses from models such as Llama 3 70B Instruct and Llama 3 8B Instruct. The first strategy, Fact-Check-Then-RAG, improves Retrieval-Augmented Generation (RAG) by incorporating fact-checking results to guide the retrieval process without updating model parameters. The second strategy, Learning from Fact-Checks via Self-Training, involves supervised fine-tuning (SFT) on fact-checked responses or applying Simple Preference Optimization (SimPO) with fact-checking as a ranking mechanism, both updating LLM parameters from supervision. These findings suggest that integrating fact-checked responses whether through RAG enhancement or self-training enhances the reliability and factual correctness of LLM outputs, offering a promising solution for applications where information accuracy is crucial.
SemiHVision: Enhancing Medical Multimodal Models with a Semi-Human Annotated Dataset and Fine-Tuned Instruction Generation
Oct 19, 2024



Abstract:Multimodal large language models (MLLMs) have made significant strides, yet they face challenges in the medical domain due to limited specialized knowledge. While recent medical MLLMs demonstrate strong performance in lab settings, they often struggle in real-world applications, highlighting a substantial gap between research and practice. In this paper, we seek to address this gap at various stages of the end-to-end learning pipeline, including data collection, model fine-tuning, and evaluation. At the data collection stage, we introduce SemiHVision, a dataset that combines human annotations with automated augmentation techniques to improve both medical knowledge representation and diagnostic reasoning. For model fine-tuning, we trained PMC-Cambrian-8B-AN over 2400 H100 GPU hours, resulting in performance that surpasses public medical models like HuatuoGPT-Vision-34B (79.0% vs. 66.7%) and private general models like Claude3-Opus (55.7%) on traditional benchmarks such as SLAKE and VQA-RAD. In the evaluation phase, we observed that traditional benchmarks cannot accurately reflect realistic clinical task capabilities. To overcome this limitation and provide more targeted guidance for model evaluation, we introduce the JAMA Clinical Challenge, a novel benchmark specifically designed to evaluate diagnostic reasoning. On this benchmark, PMC-Cambrian-AN achieves state-of-the-art performance with a GPT-4 score of 1.29, significantly outperforming HuatuoGPT-Vision-34B (1.13) and Claude3-Opus (1.17), demonstrating its superior diagnostic reasoning abilities.
MedQA-CS: Benchmarking Large Language Models Clinical Skills Using an AI-SCE Framework
Oct 02, 2024



Abstract:Artificial intelligence (AI) and large language models (LLMs) in healthcare require advanced clinical skills (CS), yet current benchmarks fail to evaluate these comprehensively. We introduce MedQA-CS, an AI-SCE framework inspired by medical education's Objective Structured Clinical Examinations (OSCEs), to address this gap. MedQA-CS evaluates LLMs through two instruction-following tasks, LLM-as-medical-student and LLM-as-CS-examiner, designed to reflect real clinical scenarios. Our contributions include developing MedQA-CS, a comprehensive evaluation framework with publicly available data and expert annotations, and providing the quantitative and qualitative assessment of LLMs as reliable judges in CS evaluation. Our experiments show that MedQA-CS is a more challenging benchmark for evaluating clinical skills than traditional multiple-choice QA benchmarks (e.g., MedQA). Combined with existing benchmarks, MedQA-CS enables a more comprehensive evaluation of LLMs' clinical capabilities for both open- and closed-source LLMs.
UMass-BioNLP at MEDIQA-M3G 2024: DermPrompt -- A Systematic Exploration of Prompt Engineering with GPT-4V for Dermatological Diagnosis
Apr 27, 2024



Abstract:This paper presents our team's participation in the MEDIQA-ClinicalNLP2024 shared task B. We present a novel approach to diagnosing clinical dermatology cases by integrating large multimodal models, specifically leveraging the capabilities of GPT-4V under a retriever and a re-ranker framework. Our investigation reveals that GPT-4V, when used as a retrieval agent, can accurately retrieve the correct skin condition 85% of the time using dermatological images and brief patient histories. Additionally, we empirically show that Naive Chain-of-Thought (CoT) works well for retrieval while Medical Guidelines Grounded CoT is required for accurate dermatological diagnosis. Further, we introduce a Multi-Agent Conversation (MAC) framework and show its superior performance and potential over the best CoT strategy. The experiments suggest that using naive CoT for retrieval and multi-agent conversation for critique-based diagnosis, GPT-4V can lead to an early and accurate diagnosis of dermatological conditions. The implications of this work extend to improving diagnostic workflows, supporting dermatological education, and enhancing patient care by providing a scalable, accessible, and accurate diagnostic tool.
JMLR: Joint Medical LLM and Retrieval Training for Enhancing Reasoning and Professional Question Answering Capability
Mar 02, 2024



Abstract:With the explosive growth of medical data and the rapid development of artificial intelligence technology, precision medicine has emerged as a key to enhancing the quality and efficiency of healthcare services. In this context, Large Language Models (LLMs) play an increasingly vital role in medical knowledge acquisition and question-answering systems. To further improve the performance of these systems in the medical domain, we introduce an innovative method that jointly trains an Information Retrieval (IR) system and an LLM during the fine-tuning phase. This approach, which we call Joint Medical LLM and Retrieval Training (JMLR), is designed to overcome the challenges faced by traditional models in handling medical question-answering tasks. By employing a synchronized training mechanism, JMLR reduces the demand for computational resources and enhances the model's ability to leverage medical knowledge for reasoning and answering questions. Our experimental results demonstrate that JMLR-13B (81.2% on Amboos, 61.3% on MedQA) outperforms models using conventional pre-training and fine-tuning Meditron-70B (76.4% on AMBOSS, 60.3% on MedQA). For models of the same 7B scale, JMLR-7B(68.7% on Amboos, 51.7% on MedQA) significantly outperforms other public models (Meditron-7B: 50.1%, 47.9%), proving its superiority in terms of cost (our training time: 37 hours, traditional method: 144 hours), efficiency, and effectiveness in medical question-answering tasks. Through this work, we provide a new and efficient knowledge enhancement tool for healthcare, demonstrating the great potential of integrating IR and LLM training in precision medical information retrieval and question-answering systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge