Jimeng Sun
KMM-CP: Practical Conformal Prediction under Covariate Shift via Selective Kernel Mean Matching
Mar 27, 2026Abstract:Uncertainty quantification is essential for deploying machine learning models in high-stakes domains such as scientific discovery and healthcare. Conformal Prediction (CP) provides finite-sample coverage guarantees under exchangeability, an assumption often violated in practice due to distribution shift. Under covariate shift, restoring validity requires importance weighting, yet accurate density-ratio estimation becomes unstable when training and test distributions exhibit limited support overlap. We propose KMM-CP, a conformal prediction framework based on Kernel Mean Matching (KMM) for covariate-shift correction. We show that KMM directly controls the bias-variance components governing conformal coverage error by minimizing RKHS moment discrepancy under explicit weight constraints, and establish asymptotic coverage guarantees under mild conditions. We then introduce a selective extension that identifies regions of reliable support overlap and restricts conformal correction to this subset, further improving stability in low-overlap regimes. Experiments on molecular property prediction benchmarks with realistic distribution shifts show that KMM-CP reduces coverage gap by over 50% compared to existing approaches. The code is available at https://github.com/siddharthal/KMM-CP.
A Practical Guide Towards Interpreting Time-Series Deep Clinical Predictive Models: A Reproducibility Study
Mar 25, 2026Abstract:Clinical decisions are high-stakes and require explicit justification, making model interpretability essential for auditing deep clinical models prior to deployment. As the ecosystem of model architectures and explainability methods expands, critical questions remain: Do architectural features like attention improve explainability? Do interpretability approaches generalize across clinical tasks? While prior benchmarking efforts exist, they often lack extensibility and reproducibility, and critically, fail to systematically examine how interpretability varies across the interplay of clinical tasks and model architectures. To address these gaps, we present a comprehensive benchmark evaluating interpretability methods across diverse clinical prediction tasks and model architectures. Our analysis reveals that: (1) attention when leveraged properly is a highly efficient approach for faithfully interpreting model predictions; (2) black-box interpreters like KernelSHAP and LIME are computationally infeasible for time-series clinical prediction tasks; and (3) several interpretability approaches are too unreliable to be trustworthy. From our findings, we discuss several guidelines on improving interpretability within clinical predictive pipelines. To support reproducibility and extensibility, we provide our implementations via PyHealth, a well-documented open-source framework: https://github.com/sunlabuiuc/PyHealth.
Bridging the Reproducibility Divide: Open Source Software's Role in Standardizing Healthcare AI
Mar 02, 2026Abstract:Our analysis of recent AI4H publications reveals that, despite a trend toward utilizing open datasets and sharing modeling code, 74% of AI4H papers still rely on private datasets or do not share their code. This is especially concerning in healthcare applications, where trust is essential. Furthermore, inconsistent and poorly documented data preprocessing pipelines result in variable model performance reports, even for identical tasks and datasets, making it challenging to evaluate the true effectiveness of AI models. Despite the challenges posed by the reproducibility crisis, addressing these issues through open practices offers substantial benefits. For instance, while the reproducibility mandate adds extra effort to research and publication, it significantly enhances the impact of the work. Our analysis shows that papers that used both public datasets and shared code received, on average, 110% more citations than those that do neither--more than doubling the citation count. Given the clear benefits of enhancing reproducibility, it is imperative for the AI4H community to take concrete steps to overcome existing barriers. The community should promote open science practices, establish standardized guidelines for data preprocessing, and develop robust benchmarks. Tackling these challenges through open-source development can improve reproducibility, which is essential for ensuring that AI models are safe, effective, and beneficial for patient care. This approach will help build more trustworthy AI systems that can be integrated into healthcare settings, ultimately contributing to better patient outcomes and advancing the field of medicine.
How Well Do Multimodal Models Reason on ECG Signals?
Feb 27, 2026Abstract:While multimodal large language models offer a promising solution to the "black box" nature of health AI by generating interpretable reasoning traces, verifying the validity of these traces remains a critical challenge. Existing evaluation methods are either unscalable, relying on manual clinician review, or superficial, utilizing proxy metrics (e.g. QA) that fail to capture the semantic correctness of clinical logic. In this work, we introduce a reproducible framework for evaluating reasoning in ECG signals. We propose decomposing reasoning into two distinct, components: (i) Perception, the accurate identification of patterns within the raw signal, and (ii) Deduction, the logical application of domain knowledge to those patterns. To evaluate Perception, we employ an agentic framework that generates code to empirically verify the temporal structures described in the reasoning trace. To evaluate Deduction, we measure the alignment of the model's logic against a structured database of established clinical criteria in a retrieval-based approach. This dual-verification method enables the scalable assessment of "true" reasoning capabilities.
ODEBrain: Continuous-Time EEG Graph for Modeling Dynamic Brain Networks
Feb 26, 2026Abstract:Modeling neural population dynamics is crucial for foundational neuroscientific research and various clinical applications. Conventional latent variable methods typically model continuous brain dynamics through discretizing time with recurrent architecture, which necessarily results in compounded cumulative prediction errors and failure of capturing instantaneous, nonlinear characteristics of EEGs. We propose ODEBRAIN, a Neural ODE latent dynamic forecasting framework to overcome these challenges by integrating spatio-temporal-frequency features into spectral graph nodes, followed by a Neural ODE modeling the continuous latent dynamics. Our design ensures that latent representations can capture stochastic variations of complex brain states at any given time point. Extensive experiments verify that ODEBRAIN can improve significantly over existing methods in forecasting EEG dynamics with enhanced robustness and generalization capabilities.
Making Conformal Predictors Robust in Healthcare Settings: a Case Study on EEG Classification
Feb 23, 2026Abstract:Quantifying uncertainty in clinical predictions is critical for high-stakes diagnosis tasks. Conformal prediction offers a principled approach by providing prediction sets with theoretical coverage guarantees. However, in practice, patient distribution shifts violate the i.i.d. assumptions underlying standard conformal methods, leading to poor coverage in healthcare settings. In this work, we evaluate several conformal prediction approaches on EEG seizure classification, a task with known distribution shift challenges and label uncertainty. We demonstrate that personalized calibration strategies can improve coverage by over 20 percentage points while maintaining comparable prediction set sizes. Our implementation is available via PyHealth, an open-source healthcare AI framework: https://github.com/sunlabuiuc/PyHealth.
Neural Signals Generate Clinical Notes in the Wild
Jan 29, 2026Abstract:Generating clinical reports that summarize abnormal patterns, diagnostic findings, and clinical interpretations from long-term EEG recordings remains labor-intensive. We curate a large-scale clinical EEG dataset with $9{,}922$ reports paired with approximately $11{,}000$ hours of EEG recordings from $9{,}048$ patients. We therefore develop CELM, the first clinical EEG-to-Language foundation model capable of summarizing long-duration, variable-length EEG recordings and performing end-to-end clinical report generation at multiple scales, including recording description, background activity, epileptiform abnormalities, events/seizures, and impressions. Experimental results show that, with patient history supervision, our method achieves $70\%$--$95\%$ average relative improvements in standard generation metrics (e.g., ROUGE-1 and METEOR) from $0.2$--$0.3$ to $0.4$--$0.6$. In the zero-shot setting without patient history, CELM attains generation scores in the range of $0.43$--$0.52$, compared to baselines of $0.17$--$0.26$. CELM integrates pretrained EEG foundation models with language models to enable scalable multimodal learning. We release our model and benchmark construction pipeline at [URL].
PyHealth 2.0: A Comprehensive Open-Source Toolkit for Accessible and Reproducible Clinical Deep Learning
Jan 23, 2026Abstract:Difficulty replicating baselines, high computational costs, and required domain expertise create persistent barriers to clinical AI research. To address these challenges, we introduce PyHealth 2.0, an enhanced clinical deep learning toolkit that enables predictive modeling in as few as 7 lines of code. PyHealth 2.0 offers three key contributions: (1) a comprehensive toolkit addressing reproducibility and compatibility challenges by unifying 15+ datasets, 20+ clinical tasks, 25+ models, 5+ interpretability methods, and uncertainty quantification including conformal prediction within a single framework that supports diverse clinical data modalities - signals, imaging, and electronic health records - with translation of 5+ medical coding standards; (2) accessibility-focused design accommodating multimodal data and diverse computational resources with up to 39x faster processing and 20x lower memory usage, enabling work from 16GB laptops to production systems; and (3) an active open-source community of 400+ members lowering domain expertise barriers through extensive documentation, reproducible research contributions, and collaborations with academic health systems and industry partners, including multi-language support via RHealth. PyHealth 2.0 establishes an open-source foundation and community advancing accessible, reproducible healthcare AI. Available at pip install pyhealth.
$\texttt{AMEND++}$: Benchmarking Eligibility Criteria Amendments in Clinical Trials
Jan 09, 2026Abstract:Clinical trial amendments frequently introduce delays, increased costs, and administrative burden, with eligibility criteria being the most commonly amended component. We introduce \textit{eligibility criteria amendment prediction}, a novel NLP task that aims to forecast whether the eligibility criteria of an initial trial protocol will undergo future amendments. To support this task, we release $\texttt{AMEND++}$, a benchmark suite comprising two datasets: $\texttt{AMEND}$, which captures eligibility-criteria version histories and amendment labels from public clinical trials, and $\verb|AMEND_LLM|$, a refined subset curated using an LLM-based denoising pipeline to isolate substantive changes. We further propose $\textit{Change-Aware Masked Language Modeling}$ (CAMLM), a revision-aware pretraining strategy that leverages historical edits to learn amendment-sensitive representations. Experiments across diverse baselines show that CAMLM consistently improves amendment prediction, enabling more robust and cost-effective clinical trial design.
Adaptation of Agentic AI
Dec 22, 2025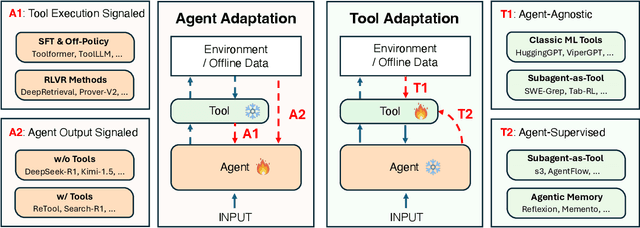
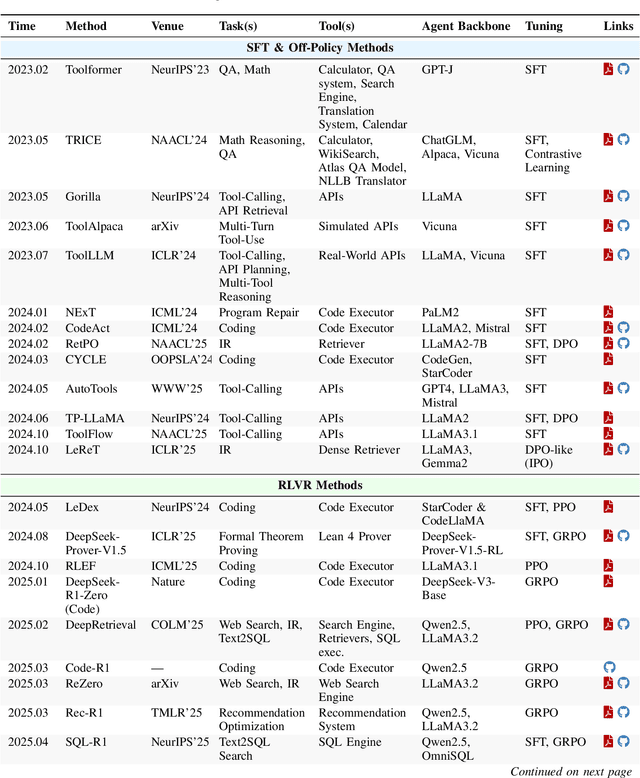
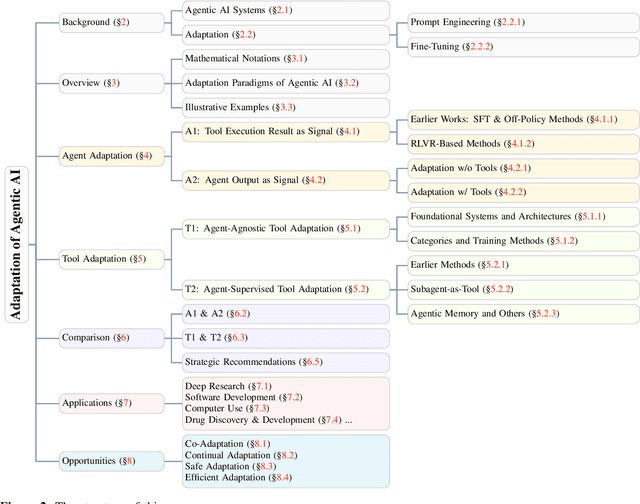
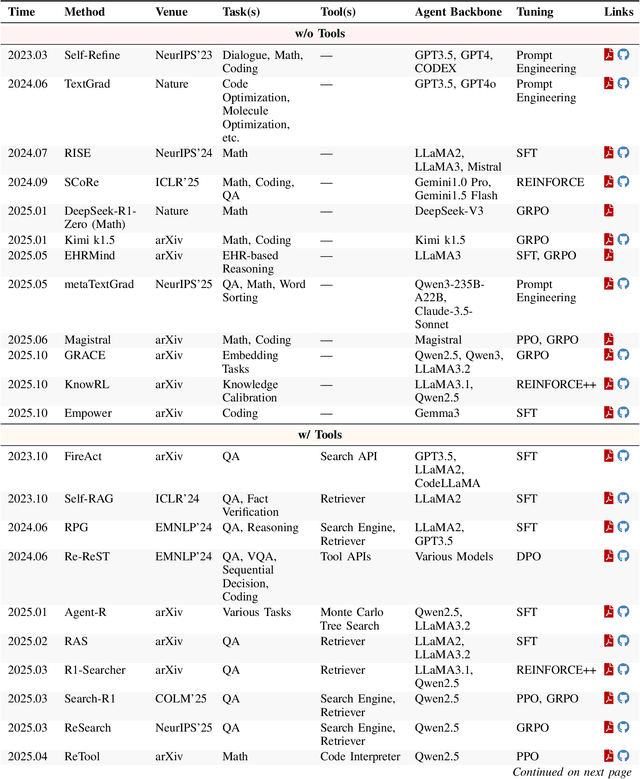
Abstract:Cutting-edge agentic AI systems are built on foundation models that can be adapted to plan, reason, and interact with external tools to perform increasingly complex and specialized tasks. As these systems grow in capability and scope, adaptation becomes a central mechanism for improving performance, reliability, and generalization. In this paper, we unify the rapidly expanding research landscape into a systematic framework that spans both agent adaptations and tool adaptations. We further decompose these into tool-execution-signaled and agent-output-signaled forms of agent adaptation, as well as agent-agnostic and agent-supervised forms of tool adaptation. We demonstrate that this framework helps clarify the design space of adaptation strategies in agentic AI, makes their trade-offs explicit, and provides practical guidance for selecting or switching among strategies during system design. We then review the representative approaches in each category, analyze their strengths and limitations, and highlight key open challenges and future opportunities. Overall, this paper aims to offer a conceptual foundation and practical roadmap for researchers and practitioners seeking to build more capable, efficient, and reliable agentic AI systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge