Hugo J. Kuijf
for the ALFA study
Explainable artificial intelligence (XAI) in deep learning-based medical image analysis
Jul 22, 2021
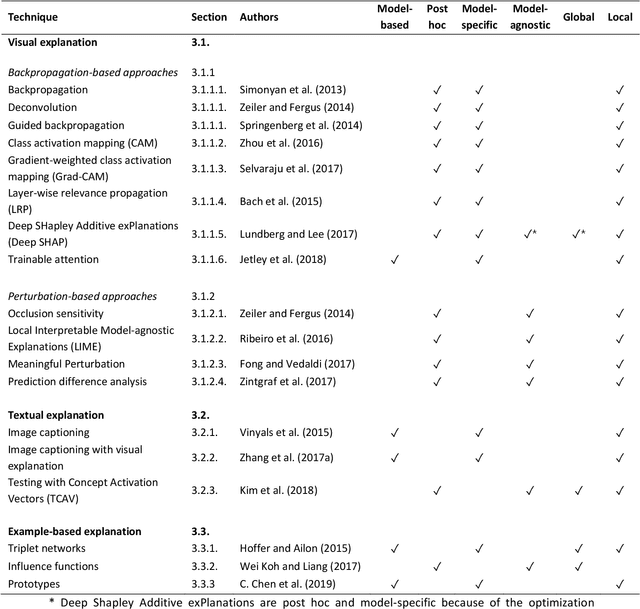
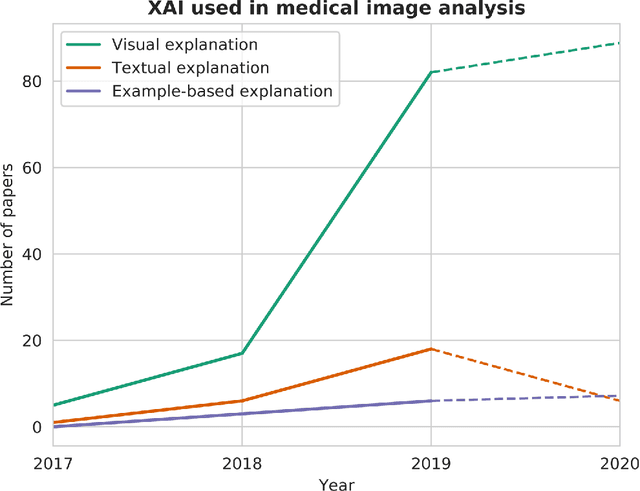
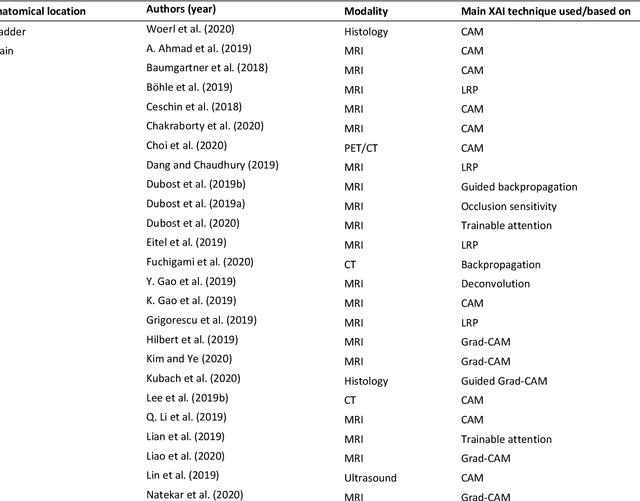
Abstract:With an increase in deep learning-based methods, the call for explainability of such methods grows, especially in high-stakes decision making areas such as medical image analysis. This survey presents an overview of eXplainable Artificial Intelligence (XAI) used in deep learning-based medical image analysis. A framework of XAI criteria is introduced to classify deep learning-based medical image analysis methods. Papers on XAI techniques in medical image analysis are then surveyed and categorized according to the framework and according to anatomical location. The paper concludes with an outlook of future opportunities for XAI in medical image analysis.
Using uncertainty estimation to reduce false positives in liver lesion detection
Jan 26, 2021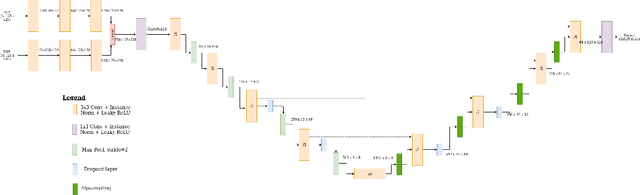
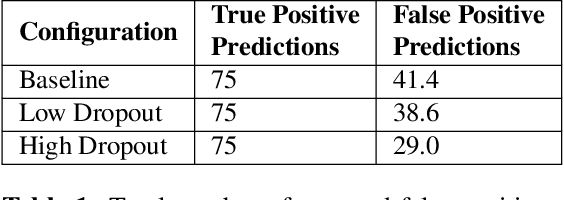
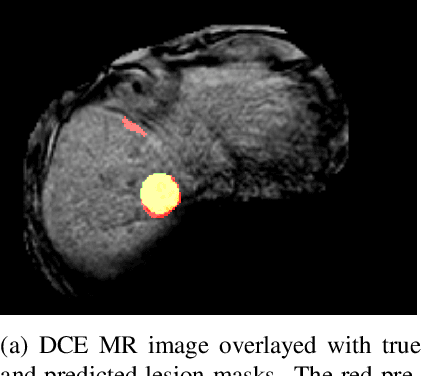
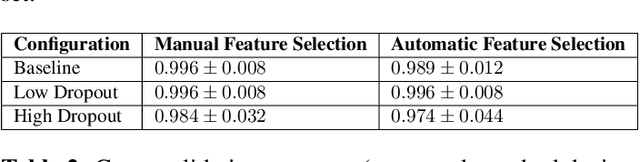
Abstract:Despite the successes of deep learning techniques at detecting objects in medical images, false positive detections occur which may hinder an accurate diagnosis. We propose a technique to reduce false positive detections made by a neural network using an SVM classifier trained with features derived from the uncertainty map of the neural network prediction. We demonstrate the effectiveness of this method for the detection of liver lesions on a dataset of abdominal MR images. We find that the use of a dropout rate of 0.5 produces the least number of false positives in the neural network predictions and the trained classifier filters out approximately 90% of these false positives detections in the test-set.
Variational Autoencoders with a Structural Similarity Loss in Time of Flight MRAs
Jan 20, 2021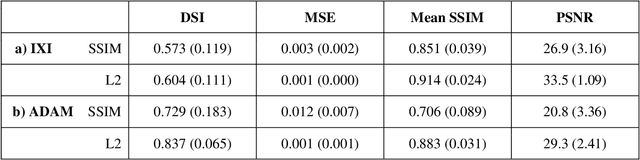
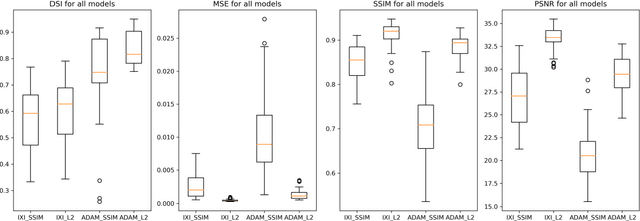
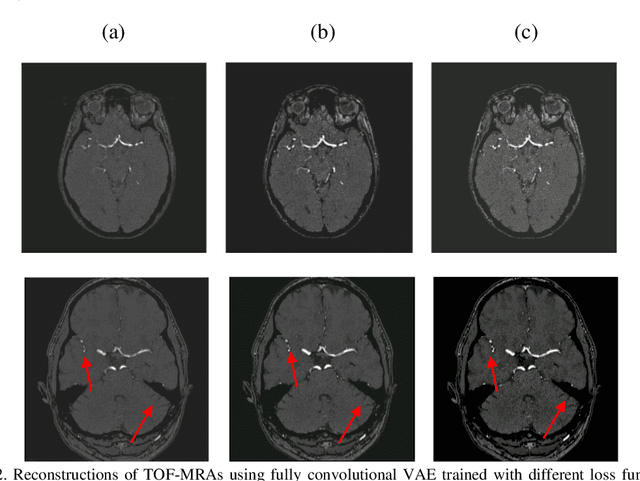
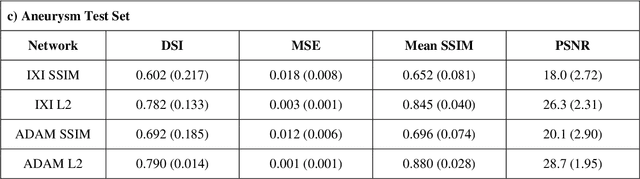
Abstract:Time-of-Flight Magnetic Resonance Angiographs (TOF-MRAs) enable visualization and analysis of cerebral arteries. This analysis may indicate normal variation of the configuration of the cerebrovascular system or vessel abnormalities, such as aneurysms. A model would be useful to represent normal cerebrovascular structure and variabilities in a healthy population and to differentiate from abnormalities. Current anomaly detection using autoencoding convolutional neural networks usually use a voxelwise mean-error for optimization. We propose optimizing a variational-autoencoder (VAE) with structural similarity loss (SSIM) for TOF-MRA reconstruction. A patch-trained 2D fully-convolutional VAE was optimized for TOF-MRA reconstruction by comparing vessel segmentations of original and reconstructed MRAs. The method was trained and tested on two datasets: the IXI dataset, and a subset from the ADAM challenge. Both trained networks were tested on a dataset including subjects with aneurysms. We compared VAE optimization with L2-loss and SSIM-loss. Performance was evaluated between original and reconstructed MRAs using mean square error, mean-SSIM, peak-signal-to-noise-ratio and dice similarity index (DSI) of segmented vessels. The L2-optimized VAE outperforms SSIM, with improved reconstruction metrics and DSIs for both datasets. Optimization using SSIM performed best for visual image quality, but with discrepancy in quantitative reconstruction and vascular segmentation. The larger, more diverse IXI dataset had overall better performance. Reconstruction metrics, including SSIM, were lower for MRAs including aneurysms. A SSIM-optimized VAE improved the visual perceptive image quality of TOF-MRA reconstructions. A L2-optimized VAE performed best for TOF-MRA reconstruction, where the vascular segmentation is important. SSIM is a potential metric for anomaly detection of MRAs.
Liver segmentation and metastases detection in MR images using convolutional neural networks
Oct 15, 2019
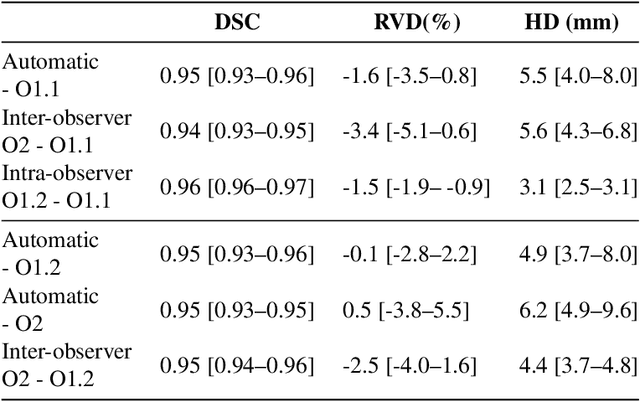
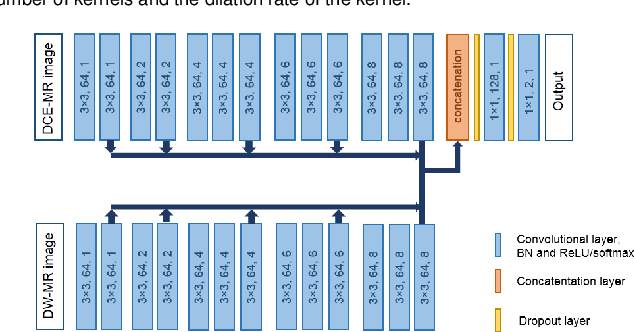
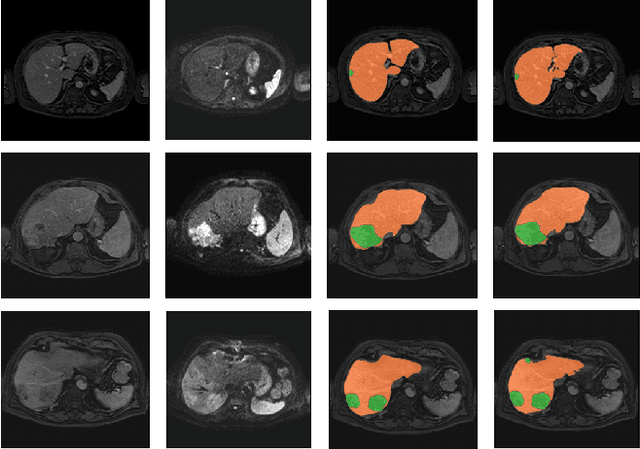
Abstract:Primary tumors have a high likelihood of developing metastases in the liver and early detection of these metastases is crucial for patient outcome. We propose a method based on convolutional neural networks (CNN) to detect liver metastases. First, the liver was automatically segmented using the six phases of abdominal dynamic contrast enhanced (DCE) MR images. Next, DCE-MR and diffusion weighted (DW) MR images are used for metastases detection within the liver mask. The liver segmentations have a median Dice similarity coefficient of 0.95 compared with manual annotations. The metastases detection method has a sensitivity of 99.8% with a median of 2 false positives per image. The combination of the two MR sequences in a dual pathway network is proven valuable for the detection of liver metastases. In conclusion, a high quality liver segmentation can be obtained in which we can successfully detect liver metastases.
Optimal input configuration of dynamic contrast enhanced MRI in convolutional neural networks for liver segmentation
Aug 22, 2019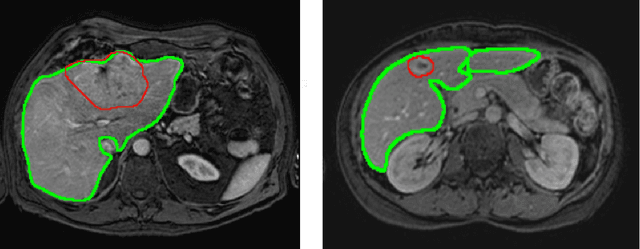
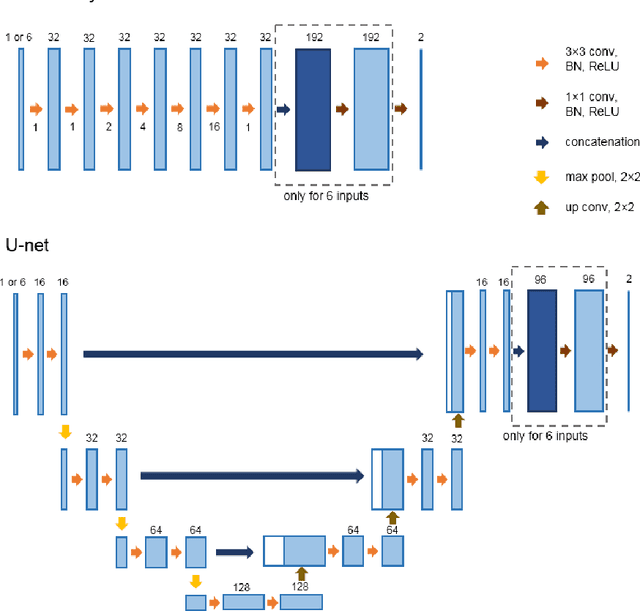
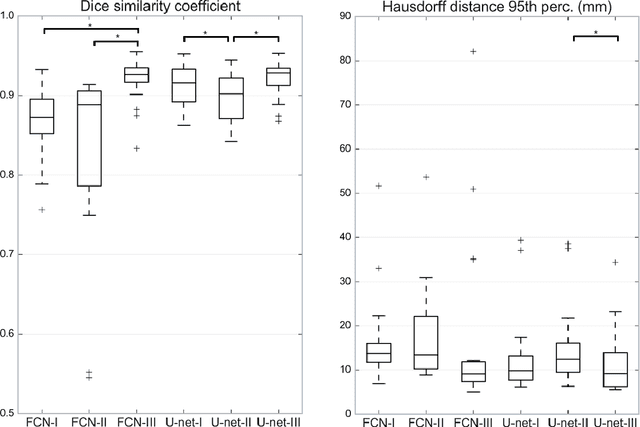
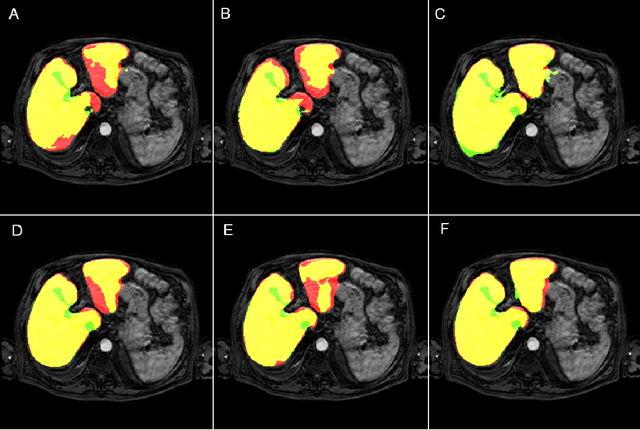
Abstract:Most MRI liver segmentation methods use a structural 3D scan as input, such as a T1 or T2 weighted scan. Segmentation performance may be improved by utilizing both structural and functional information, as contained in dynamic contrast enhanced (DCE) MR series. Dynamic information can be incorporated in a segmentation method based on convolutional neural networks in a number of ways. In this study, the optimal input configuration of DCE MR images for convolutional neural networks (CNNs) is studied. The performance of three different input configurations for CNNs is studied for a liver segmentation task. The three configurations are I) one phase image of the DCE-MR series as input image; II) the separate phases of the DCE-MR as input images; and III) the separate phases of the DCE-MR as channels of one input image. The three input configurations are fed into a dilated fully convolutional network and into a small U-net. The CNNs were trained using 19 annotated DCE-MR series and tested on another 19 annotated DCE-MR series. The performance of the three input configurations for both networks is evaluated against manual annotations. The results show that both neural networks perform better when the separate phases of the DCE-MR series are used as channels of an input image in comparison to one phase as input image or the separate phases as input images. No significant difference between the performances of the two network architectures was found for the separate phases as channels of an input image.
Standardized Assessment of Automatic Segmentation of White Matter Hyperintensities and Results of the WMH Segmentation Challenge
Apr 01, 2019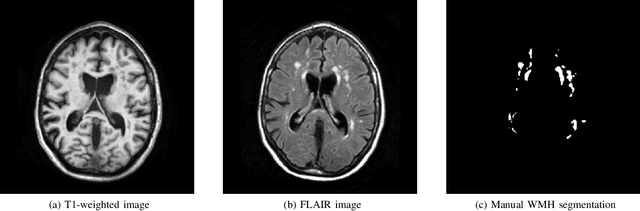
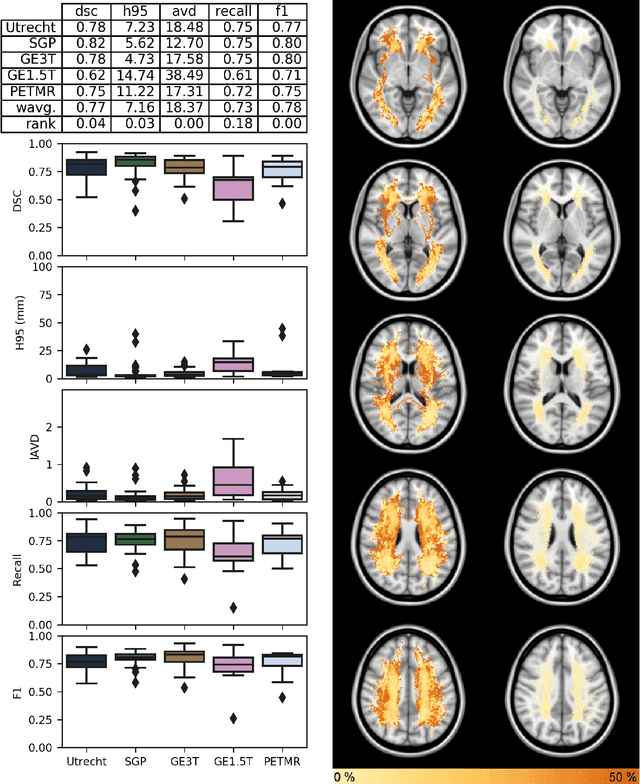
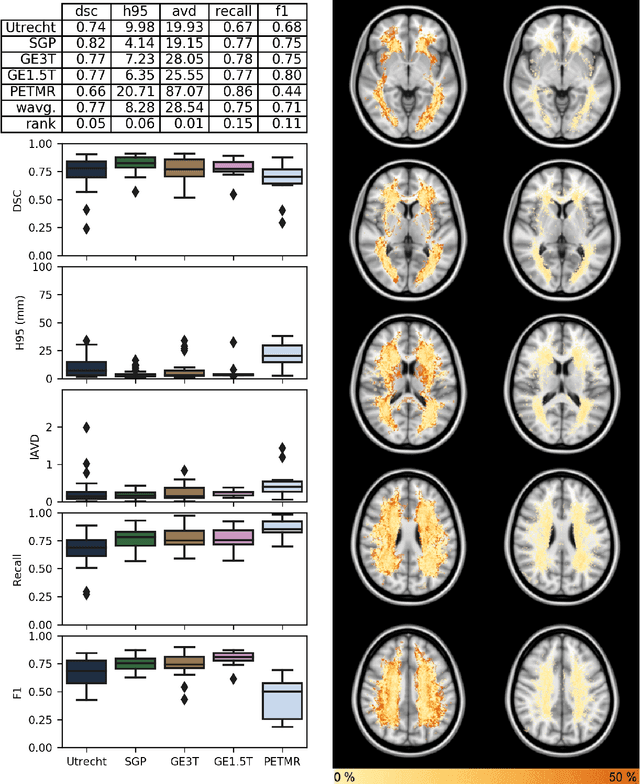
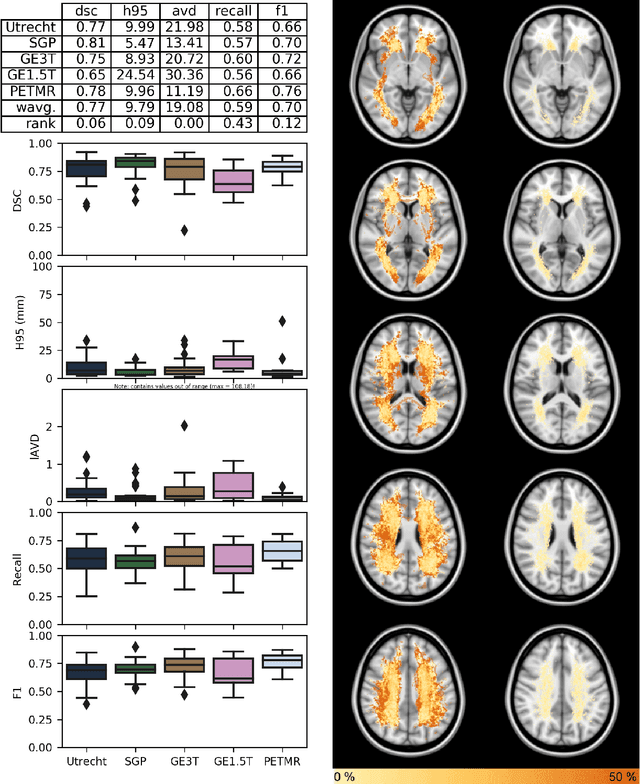
Abstract:Quantification of cerebral white matter hyperintensities (WMH) of presumed vascular origin is of key importance in many neurological research studies. Currently, measurements are often still obtained from manual segmentations on brain MR images, which is a laborious procedure. Automatic WMH segmentation methods exist, but a standardized comparison of the performance of such methods is lacking. We organized a scientific challenge, in which developers could evaluate their method on a standardized multi-center/-scanner image dataset, giving an objective comparison: the WMH Segmentation Challenge (https://wmh.isi.uu.nl/). Sixty T1+FLAIR images from three MR scanners were released with manual WMH segmentations for training. A test set of 110 images from five MR scanners was used for evaluation. Segmentation methods had to be containerized and submitted to the challenge organizers. Five evaluation metrics were used to rank the methods: (1) Dice similarity coefficient, (2) modified Hausdorff distance (95th percentile), (3) absolute log-transformed volume difference, (4) sensitivity for detecting individual lesions, and (5) F1-score for individual lesions. Additionally, methods were ranked on their inter-scanner robustness. Twenty participants submitted their method for evaluation. This paper provides a detailed analysis of the results. In brief, there is a cluster of four methods that rank significantly better than the other methods, with one clear winner. The inter-scanner robustness ranking shows that not all methods generalize to unseen scanners. The challenge remains open for future submissions and provides a public platform for method evaluation.
Response monitoring of breast cancer on DCE-MRI using convolutional neural network-generated seed points and constrained volume growing
Nov 22, 2018Abstract:Response of breast cancer to neoadjuvant chemotherapy (NAC) can be monitored using the change in visible tumor on magnetic resonance imaging (MRI). In our current workflow, seed points are manually placed in areas of enhancement likely to contain cancer. A constrained volume growing method uses these manually placed seed points as input and generates a tumor segmentation. This method is rigorously validated using complete pathological embedding. In this study, we propose to exploit deep learning for fast and automatic seed point detection, replacing manual seed point placement in our existing and well-validated workflow. The seed point generator was developed in early breast cancer patients with pathology-proven segmentations (N=100), operated shortly after MRI. It consisted of an ensemble of three independently trained fully convolutional dilated neural networks that classified breast voxels as tumor or non-tumor. Subsequently, local maxima were used as seed points for volume growing in patients receiving NAC (N=10). The percentage of tumor volume change was evaluated against semi-automatic segmentations. The primary cancer was localized in 95% of the tumors at the cost of 0.9 false positive per patient. False positives included focally enhancing regions of unknown origin and parts of the intramammary blood vessels. Volume growing from the seed points showed a median tumor volume decrease of 70% (interquartile range: 50%-77%), comparable to the semi-automatic segmentations (median: 70%, interquartile range 23%-76%). To conclude, a fast and automatic seed point generator was developed, fully automating a well-validated semi-automatic workflow for response monitoring of breast cancer to neoadjuvant chemotherapy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge