Georg Langs
Department of Biomedical Imaging and Image-guided Therapy, Computational Imaging Research Lab, Medical University Vienna, Austria
MApLe: Multi-instance Alignment of Diagnostic Reports and Large Medical Images
Apr 15, 2026Abstract:In diagnostic reports, experts encode complex imaging data into clinically actionable information. They describe subtle pathological findings that are meaningful in their anatomical context. Reports follow relatively consistent structures, expressing diagnostic information with few words that are often associated with tiny but consequential image observations. Standard vision language models struggle to identify the associations between these informative text components and small locations in the images. Here, we propose "MApLe", a multi-task, multi-instance vision language alignment approach that overcomes these limitations. It disentangles the concepts of anatomical region and diagnostic finding, and links local image information to sentences in a patch-wise approach. Our method consists of a text embedding trained to capture anatomical and diagnostic concepts in sentences, a patch-wise image encoder conditioned on anatomical structures, and a multi-instance alignment of these representations. We demonstrate that MApLe can successfully align different image regions and multiple diagnostic findings in free-text reports. We show that our model improves the alignment performance compared to state-of-the-art baseline models when evaluated on several downstream tasks. The code is available at https://github.com/cirmuw/MApLe.
HyperFitS -- Hypernetwork Fitting Spectra for metabolic quantification of ${}^1$H MR spectroscopic imaging
Apr 03, 2026Abstract:Purpose: Proton magnetic resonance spectroscopic imaging ($^1$H MRSI) enables the mapping of whole-brain metabolites concentrations in-vivo. However, a long-standing problem for its clinical applicability is the metabolic quantification, which can require extensive time for spectral fitting. Recently, deep learning methods have been able to provide whole-brain metabolic quantification in only a few seconds. However, neural network implementations often lack configurability and require retraining to change predefined parameter settings. Methods: We introduce HyperFitS, a hypernetwork for spectral fitting for metabolite quantification in whole-brain $^1$H MRSI that flexibly adapts to a broad range of baseline corrections and water suppression factors. Metabolite maps of human subjects acquired at 3T and 7T with isotropic resolutions of 10 mm, 3.4 mm and 2 mm by water-suppressed and water-unsuppressed MRSI were quantified with HyperFitS and compared to conventional LCModel fitting. Results: Metabolic maps show a substantial agreement between the new and gold-standard methods, with significantly faster fitting times by HyperFitS. Quantitative results further highlight the impact of baseline parametrization on metabolic quantification, which can alter results by up to 30%. Conclusion: HyperFitS shows strong agreement with state-of-the-art conventional methods, while reducing processing times from hours to a few seconds. Compared to prior deep learning based spectral fitting methods, HyperFitS enables a wide range of configurability and can adapt to data quality acquired with multiple protocols and field strengths without retraining.
Chronological Contrastive Learning: Few-Shot Progression Assessment in Irreversible Diseases
Mar 23, 2026Abstract:Quantitative disease severity scoring in medical imaging is costly, time-consuming, and subject to inter-reader variability. At the same time, clinical archives contain far more longitudinal imaging data than expert-annotated severity scores. Existing self-supervised methods typically ignore this chronological structure. We introduce ChronoCon, a contrastive learning approach that replaces label-based ranking losses with rankings derived solely from the visitation order of a patient's longitudinal scans. Under the clinically plausible assumption of monotonic progression in irreversible diseases, the method learns disease-relevant representations without using any expert labels. This generalizes the idea of Rank-N-Contrast from label distances to temporal ordering. Evaluated on rheumatoid arthritis radiographs for severity assessment, the learned representations substantially improve label efficiency. In low-label settings, ChronoCon significantly outperforms a fully supervised baseline initialized from ImageNet weights. In a few-shot learning experiment, fine-tuning ChronoCon on expert scores from only five patients yields an intraclass correlation coefficient of 86% for severity score prediction. These results demonstrate the potential of chronological contrastive learning to exploit routinely available imaging metadata to reduce annotation requirements in the irreversible disease domain. Code is available at https://github.com/cirmuw/ChronoCon.
MRI-derived quantification of hepatic vessel-to-volume ratios in chronic liver disease using a deep learning approach
Oct 09, 2025Abstract:Background: We aimed to quantify hepatic vessel volumes across chronic liver disease stages and healthy controls using deep learning-based magnetic resonance imaging (MRI) analysis, and assess correlations with biomarkers for liver (dys)function and fibrosis/portal hypertension. Methods: We assessed retrospectively healthy controls, non-advanced and advanced chronic liver disease (ACLD) patients using a 3D U-Net model for hepatic vessel segmentation on portal venous phase gadoxetic acid-enhanced 3-T MRI. Total (TVVR), hepatic (HVVR), and intrahepatic portal vein-to-volume ratios (PVVR) were compared between groups and correlated with: albumin-bilirubin (ALBI) and model for end-stage liver disease-sodium (MELD-Na) score, and fibrosis/portal hypertension (Fibrosis-4 [FIB-4] score, liver stiffness measurement [LSM], hepatic venous pressure gradient [HVPG], platelet count [PLT], and spleen volume). Results: We included 197 subjects, aged 54.9 $\pm$ 13.8 years (mean $\pm$ standard deviation), 111 males (56.3\%): 35 healthy controls, 44 non-ACLD, and 118 ACLD patients. TVVR and HVVR were highest in controls (3.9; 2.1), intermediate in non-ACLD (2.8; 1.7), and lowest in ACLD patients (2.3; 1.0) ($p \leq 0.001$). PVVR was reduced in both non-ACLD and ACLD patients (both 1.2) compared to controls (1.7) ($p \leq 0.001$), but showed no difference between CLD groups ($p = 0.999$). HVVR significantly correlated indirectly with FIB-4, ALBI, MELD-Na, LSM, and spleen volume ($\rho$ ranging from -0.27 to -0.40), and directly with PLT ($\rho = 0.36$). TVVR and PVVR showed similar but weaker correlations. Conclusions: Deep learning-based hepatic vessel volumetry demonstrated differences between healthy liver and chronic liver disease stages and shows correlations with established markers of disease severity.
Temporal Representation Learning of Phenotype Trajectories for pCR Prediction in Breast Cancer
Sep 18, 2025Abstract:Effective therapy decisions require models that predict the individual response to treatment. This is challenging since the progression of disease and response to treatment vary substantially across patients. Here, we propose to learn a representation of the early dynamics of treatment response from imaging data to predict pathological complete response (pCR) in breast cancer patients undergoing neoadjuvant chemotherapy (NACT). The longitudinal change in magnetic resonance imaging (MRI) data of the breast forms trajectories in the latent space, serving as basis for prediction of successful response. The multi-task model represents appearance, fosters temporal continuity and accounts for the comparably high heterogeneity in the non-responder cohort.In experiments on the publicly available ISPY-2 dataset, a linear classifier in the latent trajectory space achieves a balanced accuracy of 0.761 using only pre-treatment data (T0), 0.811 using early response (T0 + T1), and 0.861 using four imaging time points (T0 -> T3). The code will be made available upon paper acceptance.
AREPAS: Anomaly Detection in Fine-Grained Anatomy with Reconstruction-Based Semantic Patch-Scoring
Sep 16, 2025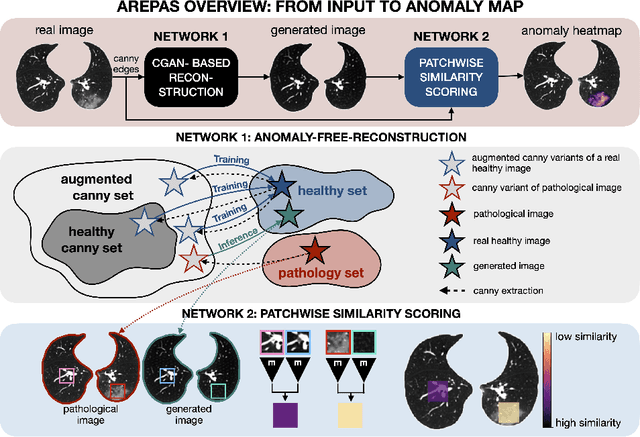
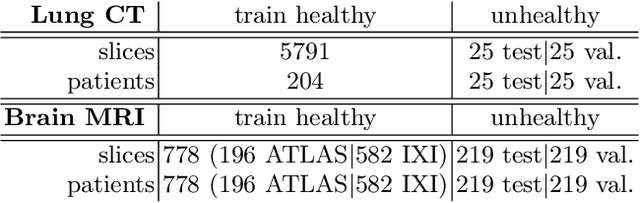
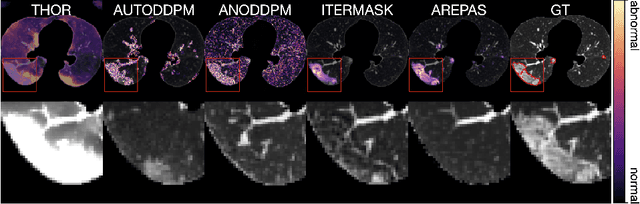
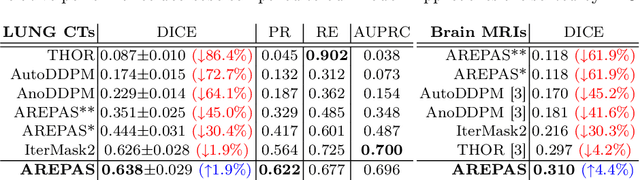
Abstract:Early detection of newly emerging diseases, lesion severity assessment, differentiation of medical conditions and automated screening are examples for the wide applicability and importance of anomaly detection (AD) and unsupervised segmentation in medicine. Normal fine-grained tissue variability such as present in pulmonary anatomy is a major challenge for existing generative AD methods. Here, we propose a novel generative AD approach addressing this issue. It consists of an image-to-image translation for anomaly-free reconstruction and a subsequent patch similarity scoring between observed and generated image-pairs for precise anomaly localization. We validate the new method on chest computed tomography (CT) scans for the detection and segmentation of infectious disease lesions. To assess generalizability, we evaluate the method on an ischemic stroke lesion segmentation task in T1-weighted brain MRI. Results show improved pixel-level anomaly segmentation in both chest CTs and brain MRIs, with relative DICE score improvements of +1.9% and +4.4%, respectively, compared to other state-of-the-art reconstruction-based methods.
Improving Vessel Segmentation with Multi-Task Learning and Auxiliary Data Available Only During Model Training
Sep 04, 2025



Abstract:Liver vessel segmentation in magnetic resonance imaging data is important for the computational analysis of vascular remodelling, associated with a wide spectrum of diffuse liver diseases. Existing approaches rely on contrast enhanced imaging data, but the necessary dedicated imaging sequences are not uniformly acquired. Images without contrast enhancement are acquired more frequently, but vessel segmentation is challenging, and requires large-scale annotated data. We propose a multi-task learning framework to segment vessels in liver MRI without contrast. It exploits auxiliary contrast enhanced MRI data available only during training to reduce the need for annotated training examples. Our approach draws on paired native and contrast enhanced data with and without vessel annotations for model training. Results show that auxiliary data improves the accuracy of vessel segmentation, even if they are not available during inference. The advantage is most pronounced if only few annotations are available for training, since the feature representation benefits from the shared task structure. A validation of this approach to augment a model for brain tumor segmentation confirms its benefits across different domains. An auxiliary informative imaging modality can augment expert annotations even if it is only available during training.
Conditional Fetal Brain Atlas Learning for Automatic Tissue Segmentation
Aug 06, 2025Abstract:Magnetic Resonance Imaging (MRI) of the fetal brain has become a key tool for studying brain development in vivo. Yet, its assessment remains challenging due to variability in brain maturation, imaging protocols, and uncertain estimates of Gestational Age (GA). To overcome these, brain atlases provide a standardized reference framework that facilitates objective evaluation and comparison across subjects by aligning the atlas and subjects in a common coordinate system. In this work, we introduce a novel deep-learning framework for generating continuous, age-specific fetal brain atlases for real-time fetal brain tissue segmentation. The framework combines a direct registration model with a conditional discriminator. Trained on a curated dataset of 219 neurotypical fetal MRIs spanning from 21 to 37 weeks of gestation. The method achieves high registration accuracy, captures dynamic anatomical changes with sharp structural detail, and robust segmentation performance with an average Dice Similarity Coefficient (DSC) of 86.3% across six brain tissues. Furthermore, volumetric analysis of the generated atlases reveals detailed neurotypical growth trajectories, providing valuable insights into the maturation of the fetal brain. This approach enables individualized developmental assessment with minimal pre-processing and real-time performance, supporting both research and clinical applications. The model code is available at https://github.com/cirmuw/fetal-brain-atlas
Identifying Signatures of Image Phenotypes to Track Treatment Response in Liver Disease
Jul 16, 2025



Abstract:Quantifiable image patterns associated with disease progression and treatment response are critical tools for guiding individual treatment, and for developing novel therapies. Here, we show that unsupervised machine learning can identify a pattern vocabulary of liver tissue in magnetic resonance images that quantifies treatment response in diffuse liver disease. Deep clustering networks simultaneously encode and cluster patches of medical images into a low-dimensional latent space to establish a tissue vocabulary. The resulting tissue types capture differential tissue change and its location in the liver associated with treatment response. We demonstrate the utility of the vocabulary on a randomized controlled trial cohort of non-alcoholic steatohepatitis patients. First, we use the vocabulary to compare longitudinal liver change in a placebo and a treatment cohort. Results show that the method identifies specific liver tissue change pathways associated with treatment, and enables a better separation between treatment groups than established non-imaging measures. Moreover, we show that the vocabulary can predict biopsy derived features from non-invasive imaging data. We validate the method on a separate replication cohort to demonstrate the applicability of the proposed method.
Towards contrast- and pathology-agnostic clinical fetal brain MRI segmentation using SynthSeg
Apr 14, 2025



Abstract:Magnetic resonance imaging (MRI) has played a crucial role in fetal neurodevelopmental research. Structural annotations of MR images are an important step for quantitative analysis of the developing human brain, with Deep learning providing an automated alternative for this otherwise tedious manual process. However, segmentation performances of Convolutional Neural Networks often suffer from domain shift, where the network fails when applied to subjects that deviate from the distribution with which it is trained on. In this work, we aim to train networks capable of automatically segmenting fetal brain MRIs with a wide range of domain shifts pertaining to differences in subject physiology and acquisition environments, in particular shape-based differences commonly observed in pathological cases. We introduce a novel data-driven train-time sampling strategy that seeks to fully exploit the diversity of a given training dataset to enhance the domain generalizability of the trained networks. We adapted our sampler, together with other existing data augmentation techniques, to the SynthSeg framework, a generator that utilizes domain randomization to generate diverse training data, and ran thorough experimentations and ablation studies on a wide range of training/testing data to test the validity of the approaches. Our networks achieved notable improvements in the segmentation quality on testing subjects with intense anatomical abnormalities (p < 1e-4), though at the cost of a slighter decrease in performance in cases with fewer abnormalities. Our work also lays the foundation for future works on creating and adapting data-driven sampling strategies for other training pipelines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge