Ge Wang
Xi'an Jiaotong University
What Does the Gradient Tell When Attacking the Graph Structure
Aug 26, 2022
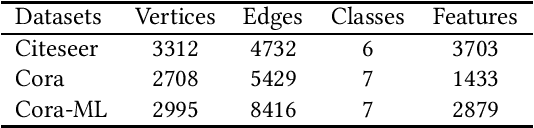
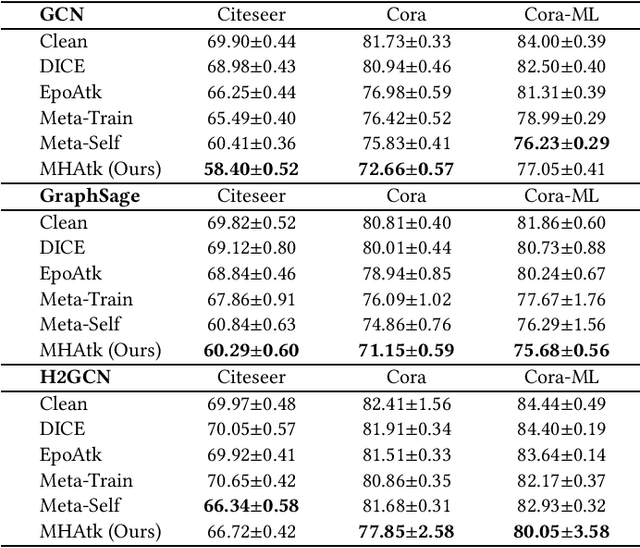
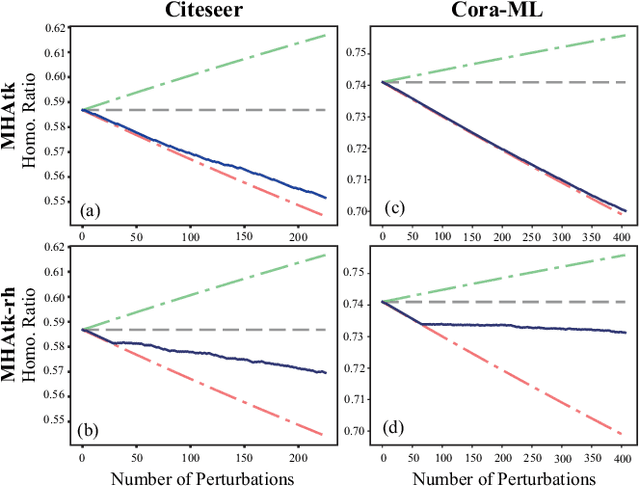
Abstract:Recent studies have proven that graph neural networks are vulnerable to adversarial attacks. Attackers can rely solely on the training labels to disrupt the performance of the agnostic victim model by edge perturbations. Researchers observe that the saliency-based attackers tend to add edges rather than delete them, which is previously explained by the fact that adding edges pollutes the nodes' features by aggregation while removing edges only leads to some loss of information. In this paper, we further prove that the attackers perturb graphs by adding inter-class edges, which also manifests as a reduction in the homophily of the perturbed graph. From this point of view, saliency-based attackers still have room for improvement in capability and imperceptibility. The message passing of the GNN-based surrogate model leads to the oversmoothing of nodes connected by inter-class edges, preventing attackers from obtaining the distinctiveness of node features. To solve this issue, we introduce a multi-hop aggregated message passing to preserve attribute differences between nodes. In addition, we propose a regularization term to restrict the homophily variance to enhance the attack imperceptibility. Experiments verify that our proposed surrogate model improves the attacker's versatility and the regularization term helps to limit the homophily of the perturbed graph.
FD-MAR: Fourier Dual-domain Network for CT Metal Artifact Reduction
Jul 24, 2022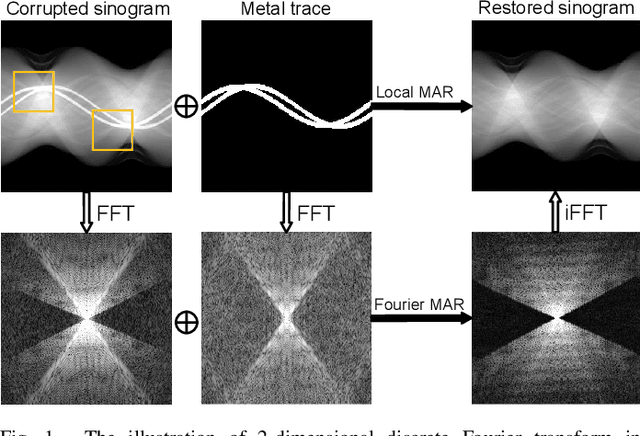
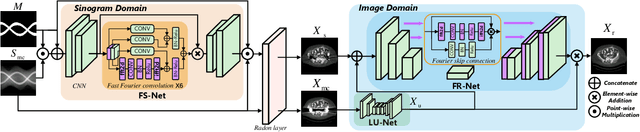
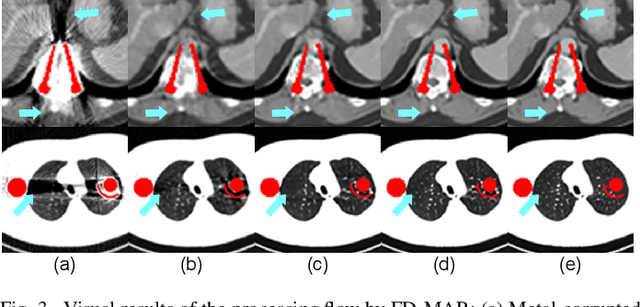
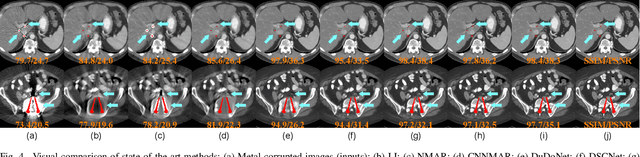
Abstract:The presence of high-density objects such as metal implants and dental fillings can introduce severely streak-like artifacts in computed tomography (CT) images, greatly limiting subsequent diagnosis. Although various deep neural networks-based methods have been proposed for metal artifact reduction (MAR), they usually suffer from poor performance due to limited exploitation of global context in the sinogram domain, secondary artifacts introduced in the image domain, and the requirement of precise metal masks. To address these issues, this paper explores fast Fourier convolution for MAR in both sinogram and image domains, and proposes a Fourier dual-domain network for MAR, termed FD-MAR. Specifically, we first propose a Fourier sinogram restoration network, which can leverage sinogram-wide receptive context to fill in the metal-corrupted region from uncorrupted region and, hence, is robust to the metal trace. Second, we propose a Fourier refinement network in the image domain, which can refine the reconstructed images in a local-to-global manner by exploring image-wide context information. As a result, the proposed FD-MAR can explore the sinogram- and image-wide receptive fields for MAR. By optimizing FD-MAR with a composite loss function, extensive experimental results demonstrate the superiority of the proposed FD-MAR over the state-of-the-art MAR methods in terms of quantitative metrics and visual comparison. Notably, FD-MAR does not require precise metal masks, which is of great importance in clinical routine.
Decorrelative Network Architecture for Robust Electrocardiogram Classification
Jul 19, 2022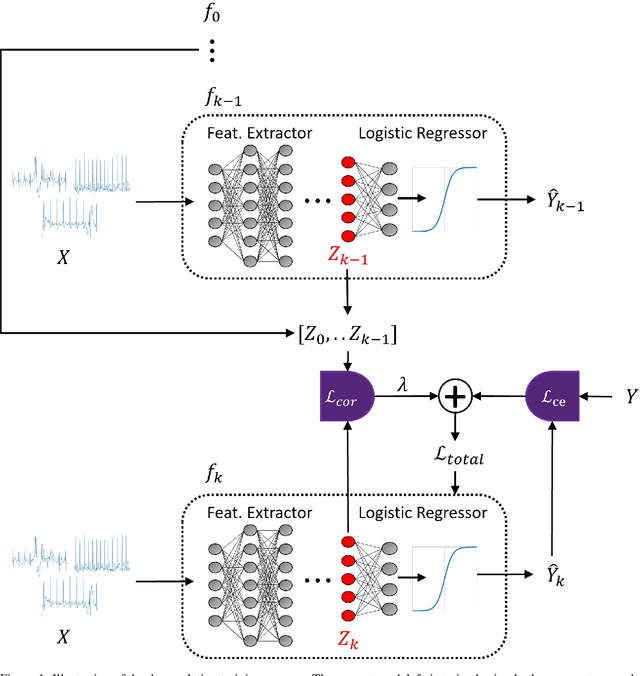
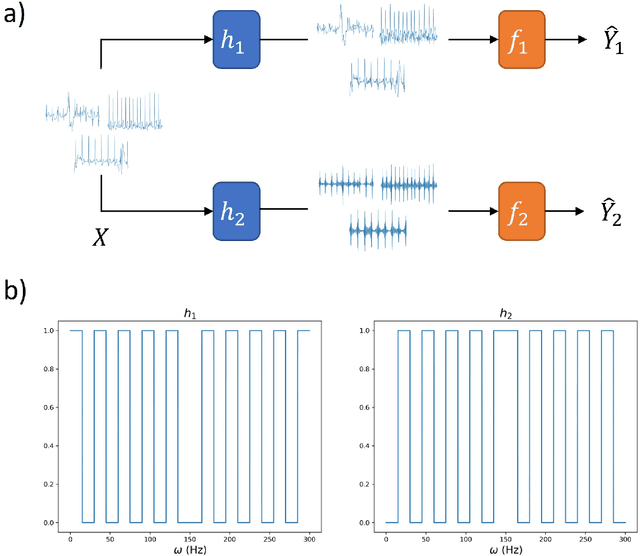
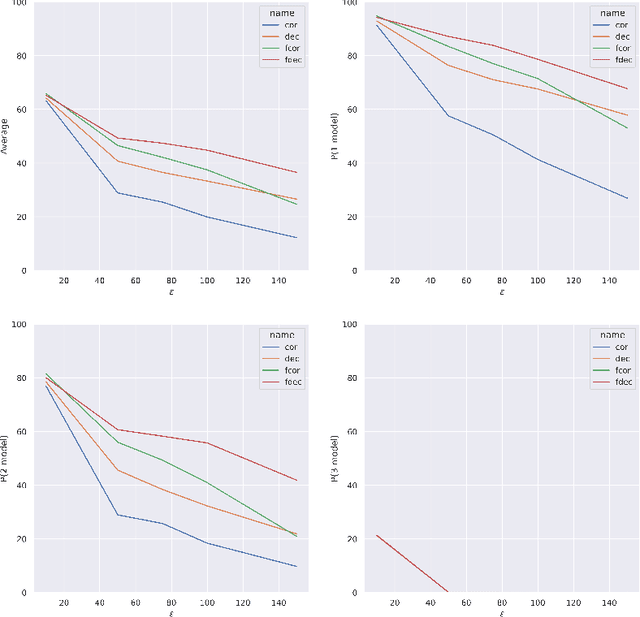
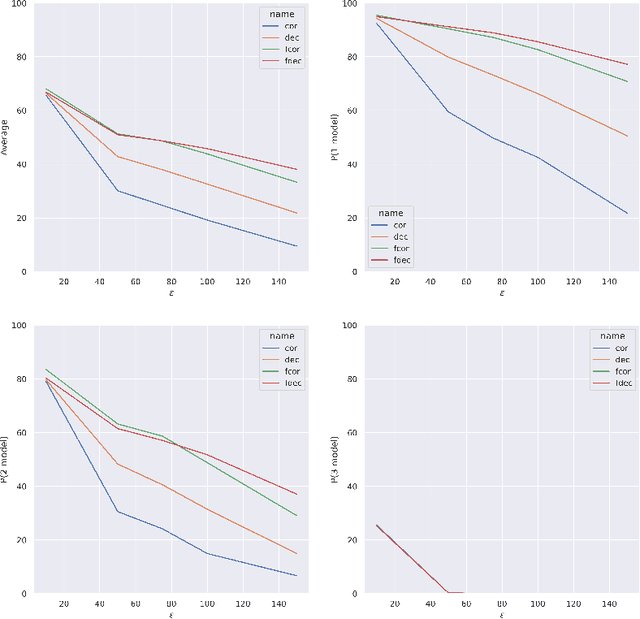
Abstract:Artificial intelligence has made great progresses in medical data analysis, but the lack of robustness and interpretability has kept these methods from being widely deployed. In particular, data-driven models are vulnerable to adversarial attacks, which are small, targeted perturbations that dramatically degrade model performance. As a recent example, while deep learning has shown impressive performance in electrocardiogram (ECG) classification, Han et al. crafted realistic perturbations that fooled the network 74% of the time [2020]. Current adversarial defense paradigms are computationally intensive and impractical for many high dimensional problems. Previous research indicates that a network vulnerability is related to the features learned during training. We propose a novel approach based on ensemble decorrelation and Fourier partitioning for training parallel network arms into a decorrelated architecture to learn complementary features, significantly reducing the chance of a perturbation fooling all arms of the deep learning model. We test our approach in ECG classification, demonstrating a much-improved 77.2% chance of at least one correct network arm on the strongest adversarial attack tested, in contrast to a 21.7% chance from a comparable ensemble. Our approach does not require expensive optimization with adversarial samples, and thus can be scaled to large problems. These methods can easily be applied to other tasks for improved network robustness.
HOME: High-Order Mixed-Moment-based Embedding for Representation Learning
Jul 15, 2022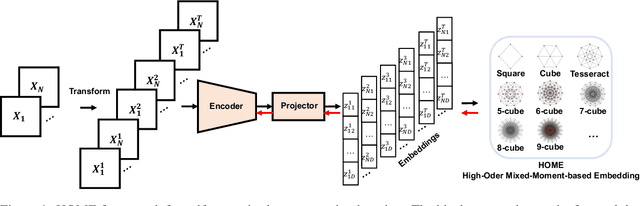
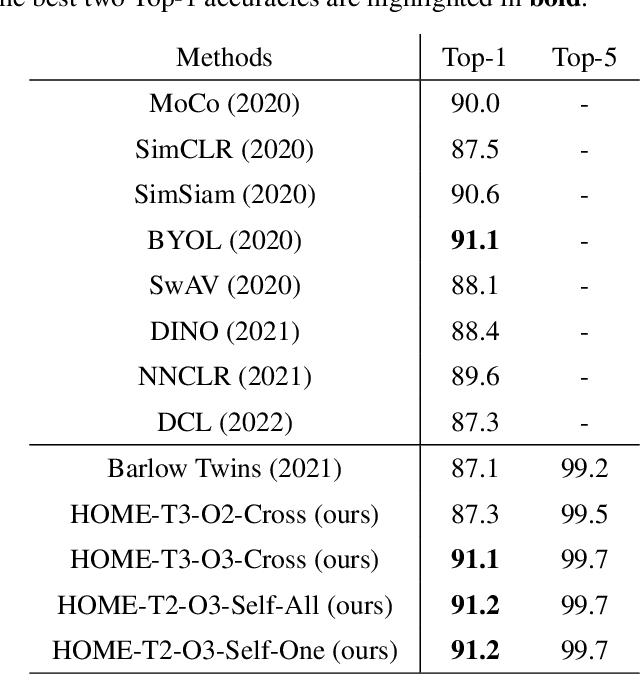
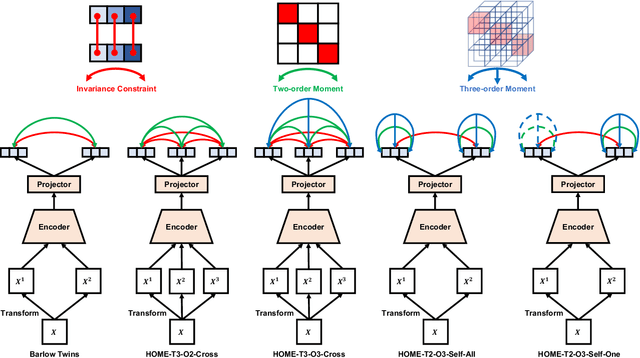
Abstract:Minimum redundancy among different elements of an embedding in a latent space is a fundamental requirement or major preference in representation learning to capture intrinsic informational structures. Current self-supervised learning methods minimize a pair-wise covariance matrix to reduce the feature redundancy and produce promising results. However, such representation features of multiple variables may contain the redundancy among more than two feature variables that cannot be minimized via the pairwise regularization. Here we propose the High-Order Mixed-Moment-based Embedding (HOME) strategy to reduce the redundancy between any sets of feature variables, which is to our best knowledge the first attempt to utilize high-order statistics/information in this context. Multivariate mutual information is minimum if and only if multiple variables are mutually independent, which suggests the necessary conditions of factorized mixed moments among multiple variables. Based on these statistical and information theoretic principles, our general HOME framework is presented for self-supervised representation learning. Our initial experiments show that a simple version in the form of a three-order HOME scheme already significantly outperforms the current two-order baseline method (i.e., Barlow Twins) in terms of the linear evaluation on representation features.
DLME: Deep Local-flatness Manifold Embedding
Jul 07, 2022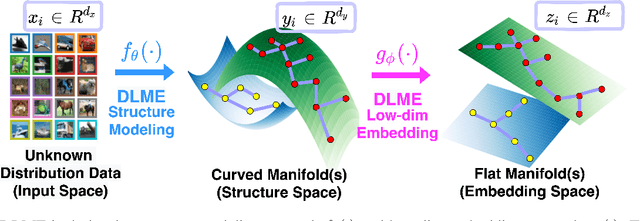
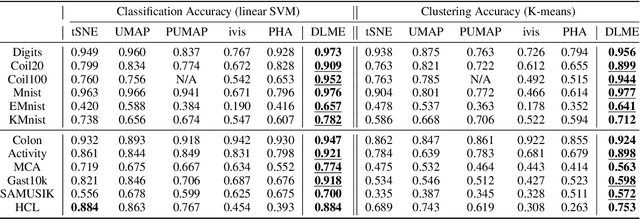
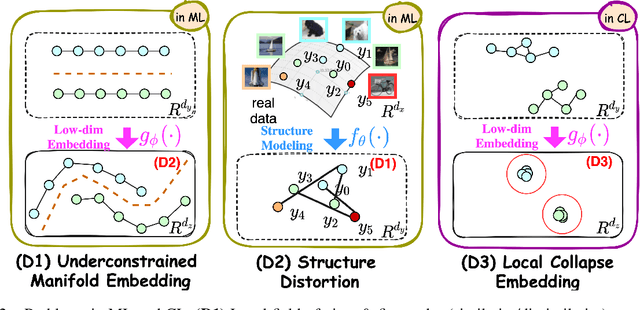
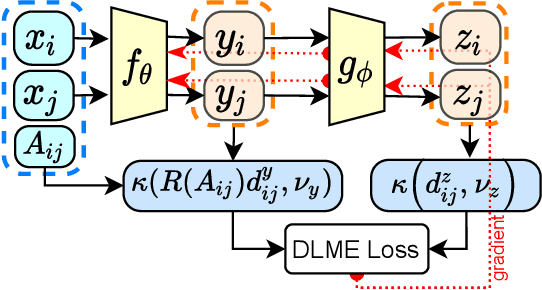
Abstract:Manifold learning~(ML) aims to find low-dimensional embedding from high-dimensional data. Previous works focus on handcraft or easy datasets with simple and ideal scenarios; however, we find they perform poorly on real-world datasets with under-sampling data. Generally, ML methods primarily model data structure and subsequently process a low-dimensional embedding, where the poor local connectivity of under-sampling data in the former step and inappropriate optimization objectives in the later step will lead to \emph{structural distortion} and \emph{underconstrained embedding}. To solve this problem, we propose Deep Local-flatness Manifold Embedding (DLME), a novel ML framework to obtain reliable manifold embedding by reducing distortion. Our proposed DLME constructs semantic manifolds by data augmentation and overcomes \emph{structural distortion} problems with the help of its smooth framework. To overcome \emph{underconstrained embedding}, we design a specific loss for DLME and mathematically demonstrate that it leads to a more suitable embedding based on our proposed Local Flatness Assumption. In the experiments, by showing the effectiveness of DLME on downstream classification, clustering, and visualization tasks with three types of datasets (toy, biological, and image), our experimental results show that DLME outperforms SOTA ML \& contrastive learning (CL) methods.
Segmentation-free PVC for Cardiac SPECT using a Densely-connected Multi-dimensional Dynamic Network
Jun 24, 2022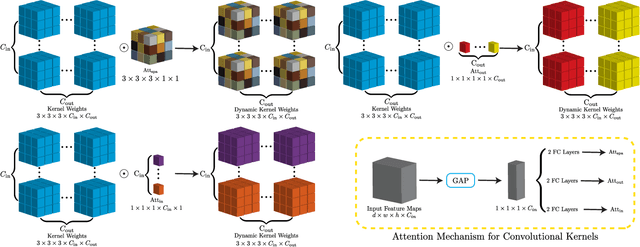
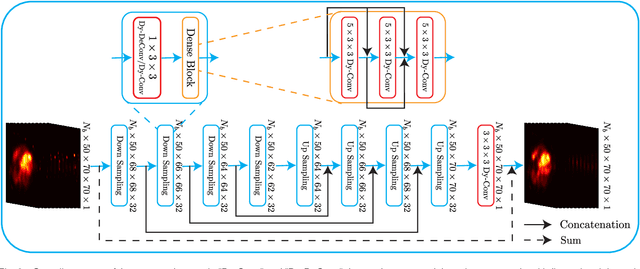
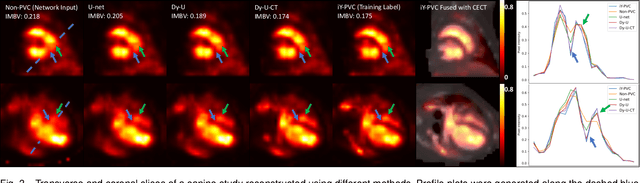

Abstract:In nuclear imaging, limited resolution causes partial volume effects (PVEs) that affect image sharpness and quantitative accuracy. Partial volume correction (PVC) methods incorporating high-resolution anatomical information from CT or MRI have been demonstrated to be effective. However, such anatomical-guided methods typically require tedious image registration and segmentation steps. Accurately segmented organ templates are also hard to obtain, particularly in cardiac SPECT imaging, due to the lack of hybrid SPECT/CT scanners with high-end CT and associated motion artifacts. Slight mis-registration/mis-segmentation would result in severe degradation in image quality after PVC. In this work, we develop a deep-learning-based method for fast cardiac SPECT PVC without anatomical information and associated organ segmentation. The proposed network involves a densely-connected multi-dimensional dynamic mechanism, allowing the convolutional kernels to be adapted based on the input images, even after the network is fully trained. Intramyocardial blood volume (IMBV) is introduced as an additional clinical-relevant loss function for network optimization. The proposed network demonstrated promising performance on 28 canine studies acquired on a GE Discovery NM/CT 570c dedicated cardiac SPECT scanner with a 64-slice CT using Technetium-99m-labeled red blood cells. This work showed that the proposed network with densely-connected dynamic mechanism produced superior results compared with the same network without such mechanism. Results also showed that the proposed network without anatomical information could produce images with statistically comparable IMBV measurements to the images generated by anatomical-guided PVC methods, which could be helpful in clinical translation.
Region-enhanced Deep Graph Convolutional Networks for Rumor Detection
Jun 22, 2022Abstract:Social media has been rapidly developing in the public sphere due to its ease of spreading new information, which leads to the circulation of rumors. However, detecting rumors from such a massive amount of information is becoming an increasingly arduous challenge. Previous work generally obtained valuable features from propagation information. It should be noted that most methods only target the propagation structure while ignoring the rumor transmission pattern. This limited focus severely restricts the collection of spread data. To solve this problem, the authors of the present study are motivated to explore the regionalized propagation patterns of rumors. Specifically, a novel region-enhanced deep graph convolutional network (RDGCN) that enhances the propagation features of rumors by learning regionalized propagation patterns and trains to learn the propagation patterns by unsupervised learning is proposed. In addition, a source-enhanced residual graph convolution layer (SRGCL) is designed to improve the graph neural network (GNN) oversmoothness and increase the depth limit of the rumor detection methods-based GNN. Experiments on Twitter15 and Twitter16 show that the proposed model performs better than the baseline approach on rumor detection and early rumor detection.
Self-Supervised Representation Learning With MUlti-Segmental Informational Coding
Jun 13, 2022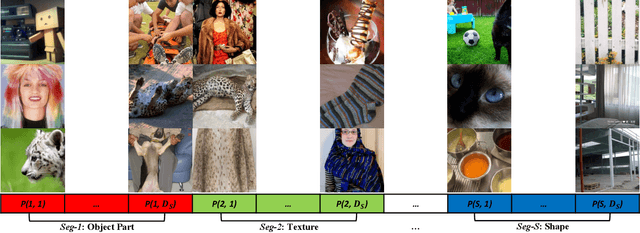
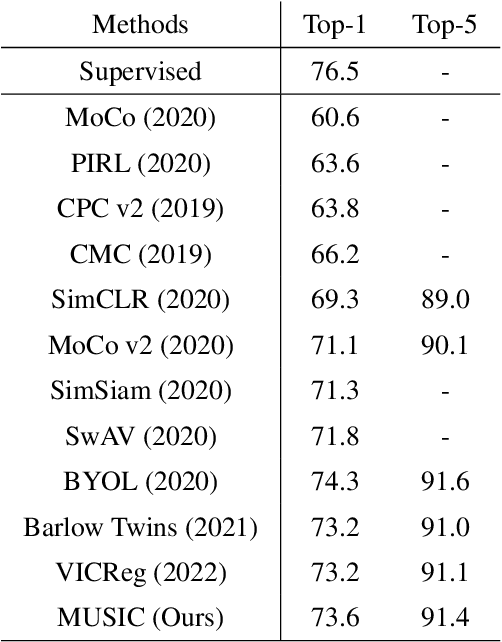
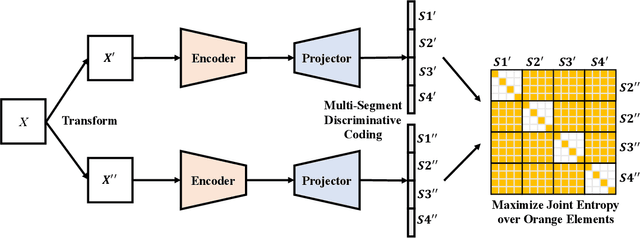
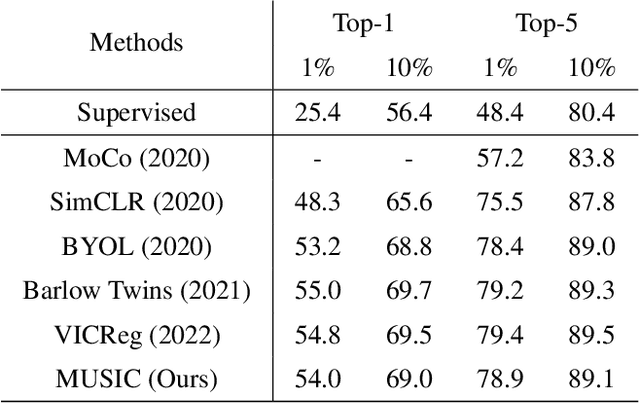
Abstract:Self-supervised representation learning maps high-dimensional data into a meaningful embedding space, where samples of similar semantic contents are close to each other. Most of the recent representation learning methods maximize cosine similarity or minimize the distance between the embedding features of different views from the same sample usually on the $l2$ normalized unit hypersphere. To prevent the trivial solutions that all samples have the same embedding feature, various techniques have been developed, such as contrastive learning, stop gradient, variance and covariance regularization, etc. In this study, we propose MUlti-Segmental Informational Coding (MUSIC) for self-supervised representation learning. MUSIC divides the embedding feature into multiple segments that discriminatively partition samples into different semantic clusters and different segments focus on different partition principles. Information theory measurements are directly used to optimize MUSIC and theoretically guarantee trivial solutions are avoided. MUSIC does not depend on commonly used techniques, such as memory bank or large batches, asymmetry networks, gradient stopping, momentum weight updating, etc, making the training framework flexible. Our experiments demonstrate that MUSIC achieves better results than most related Barlow Twins and VICReg methods on ImageNet classification with linear probing, and requires neither deep projectors nor large feature dimensions. Code will be made available.
Unsupervised Contrastive Learning based Transformer for Lung Nodule Detection
Apr 30, 2022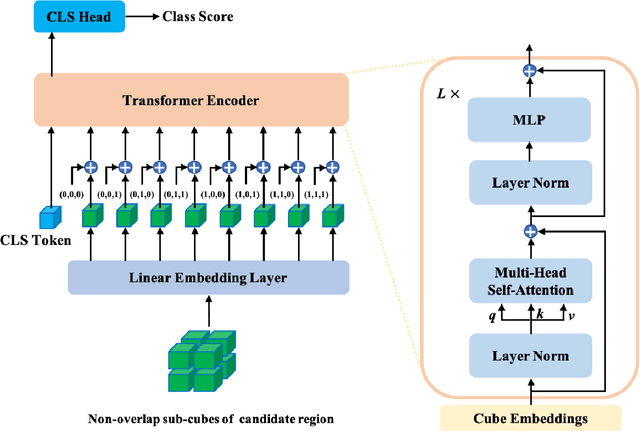
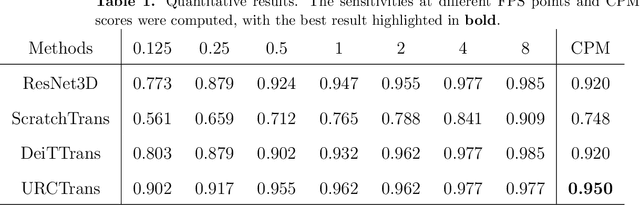
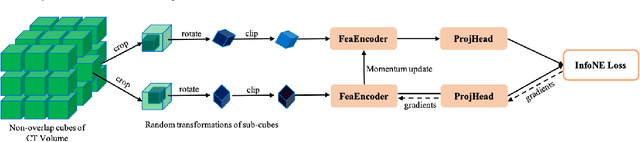
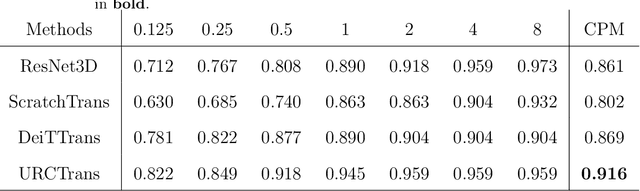
Abstract:Early detection of lung nodules with computed tomography (CT) is critical for the longer survival of lung cancer patients and better quality of life. Computer-aided detection/diagnosis (CAD) is proven valuable as a second or concurrent reader in this context. However, accurate detection of lung nodules remains a challenge for such CAD systems and even radiologists due to not only the variability in size, location, and appearance of lung nodules but also the complexity of lung structures. This leads to a high false-positive rate with CAD, compromising its clinical efficacy. Motivated by recent computer vision techniques, here we present a self-supervised region-based 3D transformer model to identify lung nodules among a set of candidate regions. Specifically, a 3D vision transformer (ViT) is developed that divides a CT image volume into a sequence of non-overlap cubes, extracts embedding features from each cube with an embedding layer, and analyzes all embedding features with a self-attention mechanism for the prediction. To effectively train the transformer model on a relatively small dataset, the region-based contrastive learning method is used to boost the performance by pre-training the 3D transformer with public CT images. Our experiments show that the proposed method can significantly improve the performance of lung nodule screening in comparison with the commonly used 3D convolutional neural networks.
Multimodal Dual Emotion with Fusion of Visual Sentiment for Rumor Detection
Apr 25, 2022
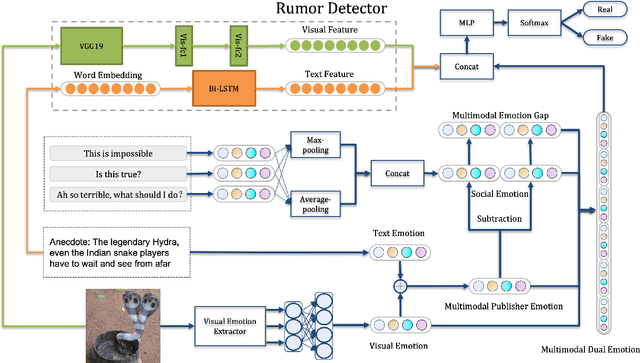
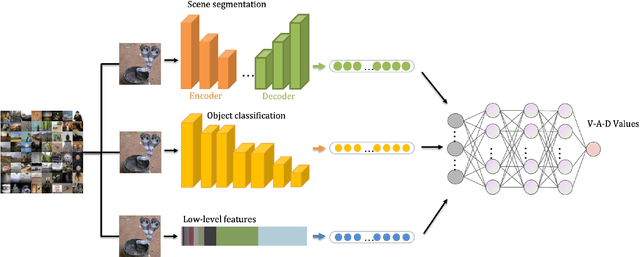
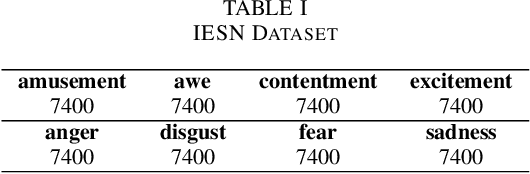
Abstract:In recent years, rumors have had a devastating impact on society, making rumor detection a significant challenge. However, the studies on rumor detection ignore the intense emotions of images in the rumor content. This paper verifies that the image emotion improves the rumor detection efficiency. A Multimodal Dual Emotion feature in rumor detection, which consists of visual and textual emotions, is proposed. To the best of our knowledge, this is the first study which uses visual emotion in rumor detection. The experiments on real datasets verify that the proposed features outperform the state-of-the-art sentiment features, and can be extended in rumor detectors while improving their performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge