Luyao Shi
SYMDIREC: A Neuro-Symbolic Divide-Retrieve-Conquer Framework for Enhanced RTL Synthesis and Summarization
Mar 17, 2026Abstract:Register-Transfer Level (RTL) synthesis and summarization are central to hardware design automation but remain challenging for Large Language Models (LLMs) due to rigid HDL syntax, limited supervision, and weak alignment with natural language. Existing prompting and retrieval-augmented generation (RAG) methods have not incorporated symbolic planning, limiting their structural precision. We introduce SYMDIREC, a neuro-symbolic framework that decomposes RTL tasks into symbolic subgoals, retrieves relevant code via a fine-tuned retriever, and assembles verified outputs through LLM reasoning. Supporting both Verilog and VHDL without LLM fine-tuning, SYMDIREC achieves ~20% higher Pass@1 rates for synthesis and 15-20% ROUGE-L improvements for summarization over prompting and RAG baselines, demonstrating the benefits of symbolic guidance in RTL tasks.
Chain-of-Descriptions: Improving Code LLMs for VHDL Code Generation and Summarization
Jul 16, 2025Abstract:Large Language Models (LLMs) have become widely used across diverse NLP tasks and domains, demonstrating their adaptability and effectiveness. In the realm of Electronic Design Automation (EDA), LLMs show promise for tasks like Register-Transfer Level (RTL) code generation and summarization. However, despite the proliferation of LLMs for general code-related tasks, there's a dearth of research focused on evaluating and refining these models for hardware description languages (HDLs), notably VHDL. In this study, we evaluate the performance of existing code LLMs for VHDL code generation and summarization using various metrics and two datasets -- VHDL-Eval and VHDL-Xform. The latter, an in-house dataset, aims to gauge LLMs' understanding of functionally equivalent code. Our findings reveal consistent underperformance of these models across different metrics, underscoring a significant gap in their suitability for this domain. To address this challenge, we propose Chain-of-Descriptions (CoDes), a novel approach to enhance the performance of LLMs for VHDL code generation and summarization tasks. CoDes involves generating a series of intermediate descriptive steps based on: (i) the problem statement for code generation, and (ii) the VHDL code for summarization. These steps are then integrated with the original input prompt (problem statement or code) and provided as input to the LLMs to generate the final output. Our experiments demonstrate that the CoDes approach significantly surpasses the standard prompting strategy across various metrics on both datasets. This method not only improves the quality of VHDL code generation and summarization but also serves as a framework for future research aimed at enhancing code LLMs for VHDL.
Improving LLM-Powered EDA Assistants with RAFT
Jun 06, 2025Abstract:Electronic design engineers often struggle to efficiently access relevant information for tasks like design verification and technology development. While large language models (LLMs) can enhance productivity as conversational agents, pre-trained open-source LLMs lack domain-specific knowledge for Electronic Design Automation (EDA). In a Retrieval-Augmented Generation (RAG) context, LLMs rely on external context but may still produce inaccurate responses. Retrieval-Augmented Fine-Tuning (RAFT) improves LLM performance, but acquiring labeled question/answer (Q/A) data in EDA is difficult. To address this, we propose using synthetic Q/A datasets to enhance LLMs with RAFT. Our results show that RAFT with synthetic data significantly boosts LLM performance for RAG-based EDA tasks. We also investigate the impact of using real user questions as Retrieval-Augmented Few-Shot (RAFS) examples for synthetic data generation. Additionally, we implement secure access control to ensure sensitive information is only accessible to authorized personnel. Finally, we assess the risk of data leakage and unintended memorization during fine-tuning with synthetic data, providing practical insights.
Modern Hopfield Networks meet Encoded Neural Representations -- Addressing Practical Considerations
Sep 24, 2024



Abstract:Content-addressable memories such as Modern Hopfield Networks (MHN) have been studied as mathematical models of auto-association and storage/retrieval in the human declarative memory, yet their practical use for large-scale content storage faces challenges. Chief among them is the occurrence of meta-stable states, particularly when handling large amounts of high dimensional content. This paper introduces Hopfield Encoding Networks (HEN), a framework that integrates encoded neural representations into MHNs to improve pattern separability and reduce meta-stable states. We show that HEN can also be used for retrieval in the context of hetero association of images with natural language queries, thus removing the limitation of requiring access to partial content in the same domain. Experimental results demonstrate substantial reduction in meta-stable states and increased storage capacity while still enabling perfect recall of a significantly larger number of inputs advancing the practical utility of associative memory networks for real-world tasks.
Self-Regulated Data-Free Knowledge Amalgamation for Text Classification
Jun 16, 2024



Abstract:Recently, there has been a growing availability of pre-trained text models on various model repositories. These models greatly reduce the cost of training new models from scratch as they can be fine-tuned for specific tasks or trained on large datasets. However, these datasets may not be publicly accessible due to the privacy, security, or intellectual property issues. In this paper, we aim to develop a lightweight student network that can learn from multiple teacher models without accessing their original training data. Hence, we investigate Data-Free Knowledge Amalgamation (DFKA), a knowledge-transfer task that combines insights from multiple pre-trained teacher models and transfers them effectively to a compact student network. To accomplish this, we propose STRATANET, a modeling framework comprising: (a) a steerable data generator that produces text data tailored to each teacher and (b) an amalgamation module that implements a self-regulative strategy using confidence estimates from the teachers' different layers to selectively integrate their knowledge and train a versatile student. We evaluate our method on three benchmark text classification datasets with varying labels or domains. Empirically, we demonstrate that the student model learned using our STRATANET outperforms several baselines significantly under data-driven and data-free constraints.
VHDL-Eval: A Framework for Evaluating Large Language Models in VHDL Code Generation
Jun 06, 2024



Abstract:With the unprecedented advancements in Large Language Models (LLMs), their application domains have expanded to include code generation tasks across various programming languages. While significant progress has been made in enhancing LLMs for popular programming languages, there exists a notable gap in comprehensive evaluation frameworks tailored for Hardware Description Languages (HDLs), particularly VHDL. This paper addresses this gap by introducing a comprehensive evaluation framework designed specifically for assessing LLM performance in VHDL code generation task. We construct a dataset for evaluating LLMs on VHDL code generation task. This dataset is constructed by translating a collection of Verilog evaluation problems to VHDL and aggregating publicly available VHDL problems, resulting in a total of 202 problems. To assess the functional correctness of the generated VHDL code, we utilize a curated set of self-verifying testbenches specifically designed for those aggregated VHDL problem set. We conduct an initial evaluation of different LLMs and their variants, including zero-shot code generation, in-context learning (ICL), and Parameter-efficient fine-tuning (PEFT) methods. Our findings underscore the considerable challenges faced by existing LLMs in VHDL code generation, revealing significant scope for improvement. This study emphasizes the necessity of supervised fine-tuning code generation models specifically for VHDL, offering potential benefits to VHDL designers seeking efficient code generation solutions.
Ask-EDA: A Design Assistant Empowered by LLM, Hybrid RAG and Abbreviation De-hallucination
Jun 03, 2024Abstract:Electronic design engineers are challenged to find relevant information efficiently for a myriad of tasks within design construction, verification and technology development. Large language models (LLM) have the potential to help improve productivity by serving as conversational agents that effectively function as subject-matter experts. In this paper we demonstrate Ask-EDA, a chat agent designed to serve as a 24x7 expert available to provide guidance to design engineers. Ask-EDA leverages LLM, hybrid retrieval augmented generation (RAG) and abbreviation de-hallucination (ADH) techniques to deliver more relevant and accurate responses. We curated three evaluation datasets, namely q2a-100, cmds-100 and abbr-100. Each dataset is tailored to assess a distinct aspect: general design question answering, design command handling and abbreviation resolution. We demonstrated that hybrid RAG offers over a 40% improvement in Recall on the q2a-100 dataset and over a 60% improvement on the cmds-100 dataset compared to not using RAG, while ADH yields over a 70% enhancement in Recall on the abbr-100 dataset. The evaluation results show that Ask-EDA can effectively respond to design-related inquiries.
TAI-GAN: A Temporally and Anatomically Informed Generative Adversarial Network for early-to-late frame conversion in dynamic cardiac PET inter-frame motion correction
Feb 14, 2024



Abstract:Inter-frame motion in dynamic cardiac positron emission tomography (PET) using rubidium-82 (82-Rb) myocardial perfusion imaging impacts myocardial blood flow (MBF) quantification and the diagnosis accuracy of coronary artery diseases. However, the high cross-frame distribution variation due to rapid tracer kinetics poses a considerable challenge for inter-frame motion correction, especially for early frames where intensity-based image registration techniques often fail. To address this issue, we propose a novel method called Temporally and Anatomically Informed Generative Adversarial Network (TAI-GAN) that utilizes an all-to-one mapping to convert early frames into those with tracer distribution similar to the last reference frame. The TAI-GAN consists of a feature-wise linear modulation layer that encodes channel-wise parameters generated from temporal information and rough cardiac segmentation masks with local shifts that serve as anatomical information. Our proposed method was evaluated on a clinical 82-Rb PET dataset, and the results show that our TAI-GAN can produce converted early frames with high image quality, comparable to the real reference frames. After TAI-GAN conversion, the motion estimation accuracy and subsequent myocardial blood flow (MBF) quantification with both conventional and deep learning-based motion correction methods were improved compared to using the original frames.
TAI-GAN: Temporally and Anatomically Informed GAN for early-to-late frame conversion in dynamic cardiac PET motion correction
Aug 23, 2023Abstract:The rapid tracer kinetics of rubidium-82 ($^{82}$Rb) and high variation of cross-frame distribution in dynamic cardiac positron emission tomography (PET) raise significant challenges for inter-frame motion correction, particularly for the early frames where conventional intensity-based image registration techniques are not applicable. Alternatively, a promising approach utilizes generative methods to handle the tracer distribution changes to assist existing registration methods. To improve frame-wise registration and parametric quantification, we propose a Temporally and Anatomically Informed Generative Adversarial Network (TAI-GAN) to transform the early frames into the late reference frame using an all-to-one mapping. Specifically, a feature-wise linear modulation layer encodes channel-wise parameters generated from temporal tracer kinetics information, and rough cardiac segmentations with local shifts serve as the anatomical information. We validated our proposed method on a clinical $^{82}$Rb PET dataset and found that our TAI-GAN can produce converted early frames with high image quality, comparable to the real reference frames. After TAI-GAN conversion, motion estimation accuracy and clinical myocardial blood flow (MBF) quantification were improved compared to using the original frames. Our code is published at https://github.com/gxq1998/TAI-GAN.
Segmentation-free PVC for Cardiac SPECT using a Densely-connected Multi-dimensional Dynamic Network
Jun 24, 2022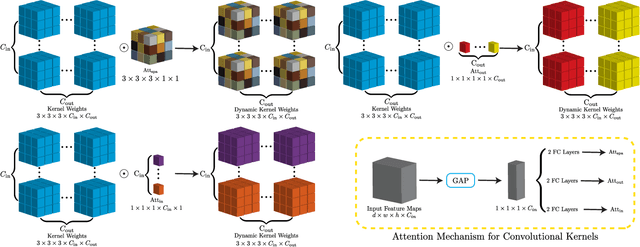
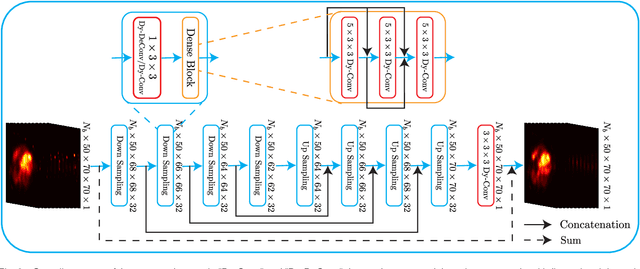
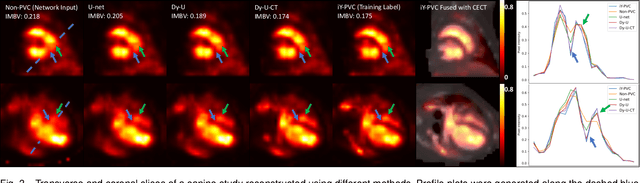

Abstract:In nuclear imaging, limited resolution causes partial volume effects (PVEs) that affect image sharpness and quantitative accuracy. Partial volume correction (PVC) methods incorporating high-resolution anatomical information from CT or MRI have been demonstrated to be effective. However, such anatomical-guided methods typically require tedious image registration and segmentation steps. Accurately segmented organ templates are also hard to obtain, particularly in cardiac SPECT imaging, due to the lack of hybrid SPECT/CT scanners with high-end CT and associated motion artifacts. Slight mis-registration/mis-segmentation would result in severe degradation in image quality after PVC. In this work, we develop a deep-learning-based method for fast cardiac SPECT PVC without anatomical information and associated organ segmentation. The proposed network involves a densely-connected multi-dimensional dynamic mechanism, allowing the convolutional kernels to be adapted based on the input images, even after the network is fully trained. Intramyocardial blood volume (IMBV) is introduced as an additional clinical-relevant loss function for network optimization. The proposed network demonstrated promising performance on 28 canine studies acquired on a GE Discovery NM/CT 570c dedicated cardiac SPECT scanner with a 64-slice CT using Technetium-99m-labeled red blood cells. This work showed that the proposed network with densely-connected dynamic mechanism produced superior results compared with the same network without such mechanism. Results also showed that the proposed network without anatomical information could produce images with statistically comparable IMBV measurements to the images generated by anatomical-guided PVC methods, which could be helpful in clinical translation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge