Earl Lawrence
Out-of-distribution transfer of PDE foundation models to material dynamics under extreme loading
Mar 04, 2026Abstract:Most PDE foundation models are pretrained and fine-tuned on fluid-centric benchmarks. Their utility under extreme-loading material dynamics remains unclear. We benchmark out-of-distribution transfer on two discontinuity-dominated regimes in which shocks, evolving interfaces, and fracture produce highly non-smooth fields: shock-driven multi-material interface dynamics (perturbed layered interface or PLI) and dynamic fracture/failure evolution (FRAC). We formulate the downstream task as terminal-state prediction, i.e., learning a long-horizon map that predicts the final state directly from the first snapshot without intermediate supervision. Using a unified training and evaluation protocol, we evaluate two open-source pretrained PDE foundation models, POSEIDON and MORPH, and compare fine-tuning from pretrained weights against training from scratch across training-set sizes to quantify sample efficiency under distribution shift.
PDE foundation model-accelerated inverse estimation of system parameters in inertial confinement fusion
Mar 04, 2026Abstract:PDE foundation models are typically pretrained on large, diverse corpora of PDE datasets and can be adapted to new settings with limited task-specific data. However, most downstream evaluations focus on forward problems, such as autoregressive rollout prediction. In this work, we study an inverse problem in inertial confinement fusion (ICF): estimating system parameters (inputs) from multi-modal, snapshot-style observations (outputs). Using the open JAG benchmark, which provides hyperspectral X-ray images and scalar observables per simulation, we finetune the PDE foundation model and train a lightweight task-specific head to jointly reconstruct hyperspectral images and regress system parameters. The fine-tuned model achieves accurate hyperspectral reconstruction (test MSE 1.2e-3) and strong parameter-estimation performance (up to R^2=0.995). Data-scaling experiments (5%-100% of the training set) show consistent improvements in both reconstruction and regression losses as the amount of training data increases, with the largest marginal gains in the low-data regime. Finally, finetuning from pretrained MORPH weights outperforms training the same architecture from scratch, demonstrating that foundation-model initialization improves sample efficiency for data-limited inverse problems in ICF.
Loss Landscape Geometry and the Learning of Symmetries: Or, What Influence Functions Reveal About Robust Generalization
Jan 28, 2026Abstract:We study how neural emulators of partial differential equation solution operators internalize physical symmetries by introducing an influence-based diagnostic that measures the propagation of parameter updates between symmetry-related states, defined as the metric-weighted overlap of loss gradients evaluated along group orbits. This quantity probes the local geometry of the learned loss landscape and goes beyond forward-pass equivariance tests by directly assessing whether learning dynamics couple physically equivalent configurations. Applying our diagnostic to autoregressive fluid flow emulators, we show that orbit-wise gradient coherence provides the mechanism for learning to generalize over symmetry transformations and indicates when training selects a symmetry compatible basin. The result is a novel technique for evaluating if surrogate models have internalized symmetry properties of the known solution operator.
A Partitioned Sparse Variational Gaussian Process for Fast, Distributed Spatial Modeling
Jul 22, 2025Abstract:The next generation of Department of Energy supercomputers will be capable of exascale computation. For these machines, far more computation will be possible than that which can be saved to disk. As a result, users will be unable to rely on post-hoc access to data for uncertainty quantification and other statistical analyses and there will be an urgent need for sophisticated machine learning algorithms which can be trained in situ. Algorithms deployed in this setting must be highly scalable, memory efficient and capable of handling data which is distributed across nodes as spatially contiguous partitions. One suitable approach involves fitting a sparse variational Gaussian process (SVGP) model independently and in parallel to each spatial partition. The resulting model is scalable, efficient and generally accurate, but produces the undesirable effect of constructing discontinuous response surfaces due to the disagreement between neighboring models at their shared boundary. In this paper, we extend this idea by allowing for a small amount of communication between neighboring spatial partitions which encourages better alignment of the local models, leading to smoother spatial predictions and a better fit in general. Due to our decentralized communication scheme, the proposed extension remains highly scalable and adds very little overhead in terms of computation (and none, in terms of memory). We demonstrate this Partitioned SVGP (PSVGP) approach for the Energy Exascale Earth System Model (E3SM) and compare the results to the independent SVGP case.
Dynamic Data Assimilation of MPAS-O and the Global Drifter Dataset
Jan 11, 2023



Abstract:In this study, we propose a new method for combining in situ buoy measurements with Earth system models (ESMs) to improve the accuracy of temperature predictions in the ocean. The technique utilizes the dynamics and modes identified in ESMs to improve the accuracy of buoy measurements while still preserving features such as seasonality. Using this technique, errors in localized temperature predictions made by the MPAS-O model can be corrected. We demonstrate that our approach improves accuracy compared to other interpolation and data assimilation methods. We apply our method to assimilate the Model for Prediction Across Scales Ocean component (MPAS-O) with the Global Drifter Program's in-situ ocean buoy dataset.
Fast emulation of density functional theory simulations using approximate Gaussian processes
Aug 24, 2022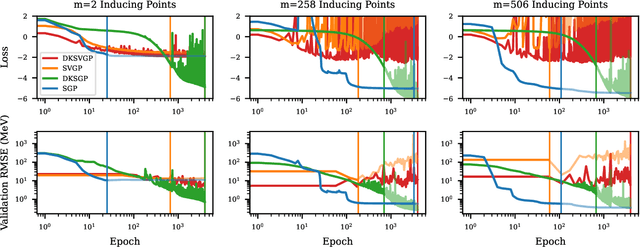
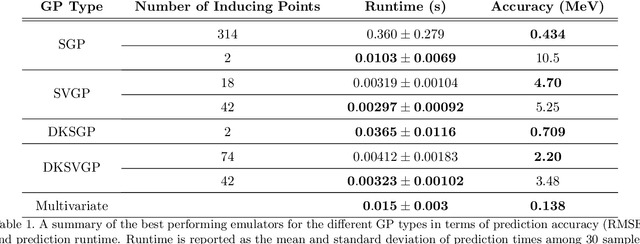
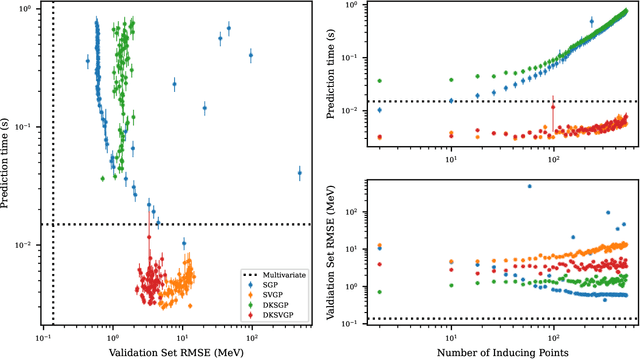
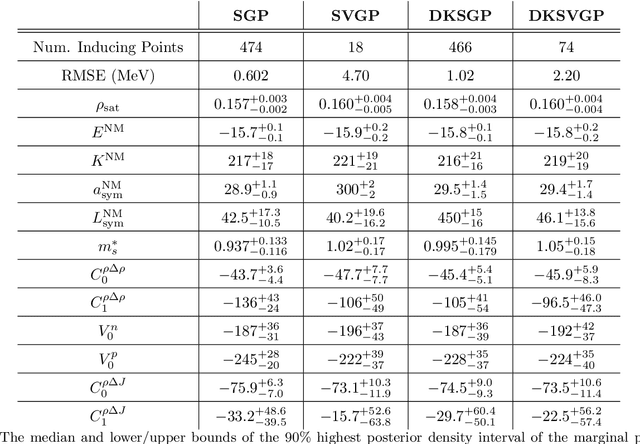
Abstract:Fitting a theoretical model to experimental data in a Bayesian manner using Markov chain Monte Carlo typically requires one to evaluate the model thousands (or millions) of times. When the model is a slow-to-compute physics simulation, Bayesian model fitting becomes infeasible. To remedy this, a second statistical model that predicts the simulation output -- an "emulator" -- can be used in lieu of the full simulation during model fitting. A typical emulator of choice is the Gaussian process (GP), a flexible, non-linear model that provides both a predictive mean and variance at each input point. Gaussian process regression works well for small amounts of training data ($n < 10^3$), but becomes slow to train and use for prediction when the data set size becomes large. Various methods can be used to speed up the Gaussian process in the medium-to-large data set regime ($n > 10^5$), trading away predictive accuracy for drastically reduced runtime. This work examines the accuracy-runtime trade-off of several approximate Gaussian process models -- the sparse variational GP, stochastic variational GP, and deep kernel learned GP -- when emulating the predictions of density functional theory (DFT) models. Additionally, we use the emulators to calibrate, in a Bayesian manner, the DFT model parameters using observed data, resolving the computational barrier imposed by the data set size, and compare calibration results to previous work. The utility of these calibrated DFT models is to make predictions, based on observed data, about the properties of experimentally unobserved nuclides of interest e.g. super-heavy nuclei.
Relationship-aware Multivariate Sampling Strategy for Scientific Simulation Data
Aug 31, 2020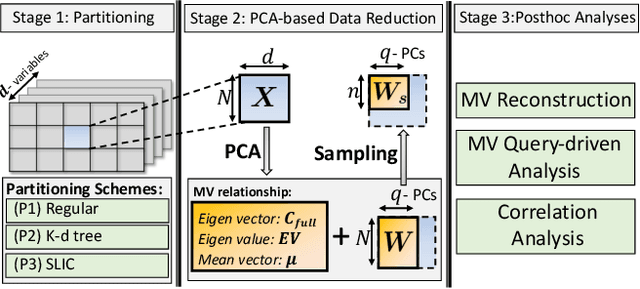
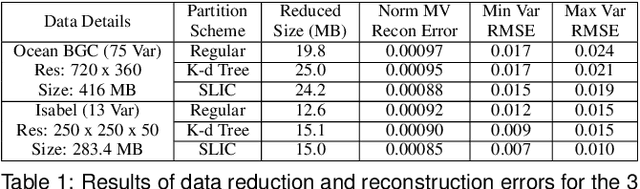
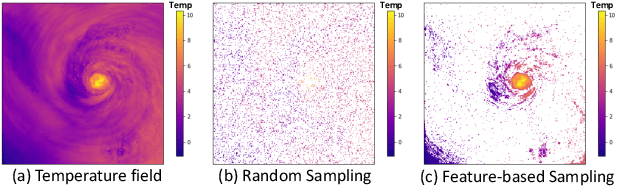
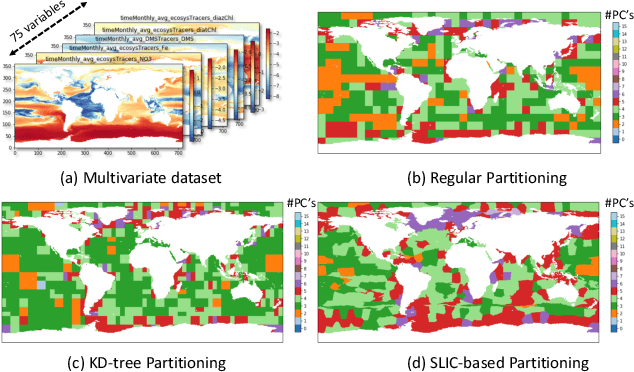
Abstract:With the increasing computational power of current supercomputers, the size of data produced by scientific simulations is rapidly growing. To reduce the storage footprint and facilitate scalable post-hoc analyses of such scientific data sets, various data reduction/summarization methods have been proposed over the years. Different flavors of sampling algorithms exist to sample the high-resolution scientific data, while preserving important data properties required for subsequent analyses. However, most of these sampling algorithms are designed for univariate data and cater to post-hoc analyses of single variables. In this work, we propose a multivariate sampling strategy which preserves the original variable relationships and enables different multivariate analyses directly on the sampled data. Our proposed strategy utilizes principal component analysis to capture the variance of multivariate data and can be built on top of any existing state-of-the-art sampling algorithms for single variables. In addition, we also propose variants of different data partitioning schemes (regular and irregular) to efficiently model the local multivariate relationships. Using two real-world multivariate data sets, we demonstrate the efficacy of our proposed multivariate sampling strategy with respect to its data reduction capabilities as well as the ease of performing efficient post-hoc multivariate analyses.
Scaled Vecchia approximation for fast computer-model emulation
May 29, 2020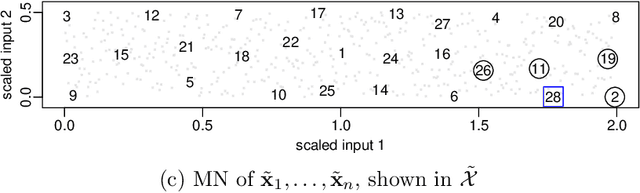
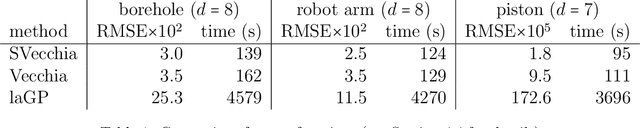


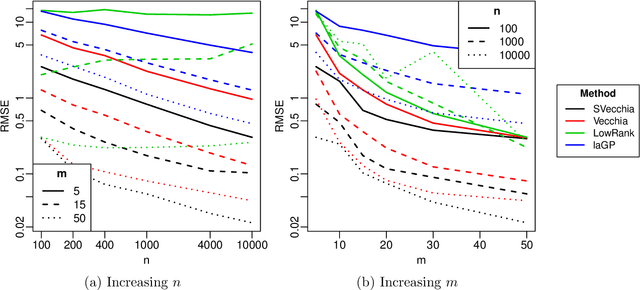
Abstract:Many scientific phenomena are studied using computer experiments consisting of multiple runs of a computer model while varying the input settings. Gaussian processes (GPs) are a popular tool for the analysis of computer experiments, enabling interpolation between input settings, but direct GP inference is computationally infeasible for large datasets. We adapt and extend a powerful class of GP methods from spatial statistics to enable the scalable analysis and emulation of large computer experiments. Specifically, we apply Vecchia's ordered conditional approximation in a transformed input space, with each input scaled according to how strongly it relates to the computer-model response. The scaling is learned from the data, by estimating parameters in the GP covariance function using Fisher scoring. Our methods are highly scalable, enabling estimation, joint prediction and simulation in near-linear time in the number of model runs. In several numerical examples, our approach substantially outperformed existing methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge