Daniel Rueckert
on behalf of the PINNACLE consortium
Approaching Peak Ground Truth
Dec 31, 2022

Abstract:Machine learning models are typically evaluated by computing similarity with reference annotations and trained by maximizing similarity with such. Especially in the bio-medical domain, annotations are subjective and suffer from low inter- and intra-rater reliability. Since annotations only reflect the annotation entity's interpretation of the real world, this can lead to sub-optimal predictions even though the model achieves high similarity scores. Here, the theoretical concept of Peak Ground Truth (PGT) is introduced. PGT marks the point beyond which an increase in similarity with the reference annotation stops translating to better Real World Model Performance (RWMP). Additionally, a quantitative technique to approximate PGT by computing inter- and intra-rater reliability is proposed. Finally, three categories of PGT-aware strategies to evaluate and improve model performance are reviewed.
Neural Implicit k-Space for Binning-free Non-Cartesian Cardiac MR Imaging
Dec 16, 2022Abstract:In this work, we propose a novel image reconstruction framework that directly learns a neural implicit representation in k-space for ECG-triggered non-Cartesian Cardiac Magnetic Resonance Imaging (CMR). While existing methods bin acquired data from neighboring time points to reconstruct one phase of the cardiac motion, our framework allows for a continuous, binning-free, and subject-specific k-space representation.We assign a unique coordinate that consists of time, coil index, and frequency domain location to each sampled k-space point. We then learn the subject-specific mapping from these unique coordinates to k-space intensities using a multi-layer perceptron with frequency domain regularization. During inference, we obtain a complete k-space for Cartesian coordinates and an arbitrary temporal resolution. A simple inverse Fourier transform recovers the image, eliminating the need for density compensation and costly non-uniform Fourier transforms for non-Cartesian data. This novel imaging framework was tested on 42 radially sampled datasets from 6 subjects. The proposed method outperforms other techniques qualitatively and quantitatively using data from four and one heartbeat(s) and 30 cardiac phases. Our results for one heartbeat reconstruction of 50 cardiac phases show improved artifact removal and spatio-temporal resolution, leveraging the potential for real-time CMR.
A Domain-specific Perceptual Metric via Contrastive Self-supervised Representation: Applications on Natural and Medical Images
Dec 03, 2022



Abstract:Quantifying the perceptual similarity of two images is a long-standing problem in low-level computer vision. The natural image domain commonly relies on supervised learning, e.g., a pre-trained VGG, to obtain a latent representation. However, due to domain shift, pre-trained models from the natural image domain might not apply to other image domains, such as medical imaging. Notably, in medical imaging, evaluating the perceptual similarity is exclusively performed by specialists trained extensively in diverse medical fields. Thus, medical imaging remains devoid of task-specific, objective perceptual measures. This work answers the question: Is it necessary to rely on supervised learning to obtain an effective representation that could measure perceptual similarity, or is self-supervision sufficient? To understand whether recent contrastive self-supervised representation (CSR) may come to the rescue, we start with natural images and systematically evaluate CSR as a metric across numerous contemporary architectures and tasks and compare them with existing methods. We find that in the natural image domain, CSR behaves on par with the supervised one on several perceptual tests as a metric, and in the medical domain, CSR better quantifies perceptual similarity concerning the experts' ratings. We also demonstrate that CSR can significantly improve image quality in two image synthesis tasks. Finally, our extensive results suggest that perceptuality is an emergent property of CSR, which can be adapted to many image domains without requiring annotations.
How Do Input Attributes Impact the Privacy Loss in Differential Privacy?
Nov 18, 2022



Abstract:Differential privacy (DP) is typically formulated as a worst-case privacy guarantee over all individuals in a database. More recently, extensions to individual subjects or their attributes, have been introduced. Under the individual/per-instance DP interpretation, we study the connection between the per-subject gradient norm in DP neural networks and individual privacy loss and introduce a novel metric termed the Privacy Loss-Input Susceptibility (PLIS), which allows one to apportion the subject's privacy loss to their input attributes. We experimentally show how this enables the identification of sensitive attributes and of subjects at high risk of data reconstruction.
The Role of Local Alignment and Uniformity in Image-Text Contrastive Learning on Medical Images
Nov 14, 2022Abstract:Image-text contrastive learning has proven effective for pretraining medical image models. When targeting localized downstream tasks like semantic segmentation or object detection, additional local contrastive losses that align image regions with sentences have shown promising results. We study how local contrastive losses are related to global (per-sample) contrastive losses and which effects they have on localized medical downstream tasks. Based on a theoretical comparison, we propose to remove some components of local losses and replace others by a novel distribution prior which enforces uniformity of representations within each sample. We empirically study this approach on chest X-ray tasks and find it to be very effective, outperforming methods without local losses on 12 of 18 tasks.
Exploiting segmentation labels and representation learning to forecast therapy response of PDAC patients
Nov 08, 2022


Abstract:The prediction of pancreatic ductal adenocarcinoma therapy response is a clinically challenging and important task in this high-mortality tumour entity. The training of neural networks able to tackle this challenge is impeded by a lack of large datasets and the difficult anatomical localisation of the pancreas. Here, we propose a hybrid deep neural network pipeline to predict tumour response to initial chemotherapy which is based on the Response Evaluation Criteria in Solid Tumors (RECIST) score, a standardised method for cancer response evaluation by clinicians as well as tumour markers, and clinical evaluation of the patients. We leverage a combination of representation transfer from segmentation to classification, as well as localisation and representation learning. Our approach yields a remarkably data-efficient method able to predict treatment response with a ROC-AUC of 63.7% using only 477 datasets in total.
Automated analysis of diabetic retinopathy using vessel segmentation maps as inductive bias
Oct 28, 2022Abstract:Recent studies suggest that early stages of diabetic retinopathy (DR) can be diagnosed by monitoring vascular changes in the deep vascular complex. In this work, we investigate a novel method for automated DR grading based on optical coherence tomography angiography (OCTA) images. Our work combines OCTA scans with their vessel segmentations, which then serve as inputs to task specific networks for lesion segmentation, image quality assessment and DR grading. For this, we generate synthetic OCTA images to train a segmentation network that can be directly applied on real OCTA data. We test our approach on MICCAI 2022's DR analysis challenge (DRAC). In our experiments, the proposed method performs equally well as the baseline model.
Generalised Likelihood Ratio Testing Adversaries through the Differential Privacy Lens
Oct 24, 2022Abstract:Differential Privacy (DP) provides tight upper bounds on the capabilities of optimal adversaries, but such adversaries are rarely encountered in practice. Under the hypothesis testing/membership inference interpretation of DP, we examine the Gaussian mechanism and relax the usual assumption of a Neyman-Pearson-Optimal (NPO) adversary to a Generalized Likelihood Test (GLRT) adversary. This mild relaxation leads to improved privacy guarantees, which we express in the spirit of Gaussian DP and $(\varepsilon, \delta)$-DP, including composition and sub-sampling results. We evaluate our results numerically and find them to match the theoretical upper bounds.
Label Noise-Robust Learning using a Confidence-Based Sieving Strategy
Oct 11, 2022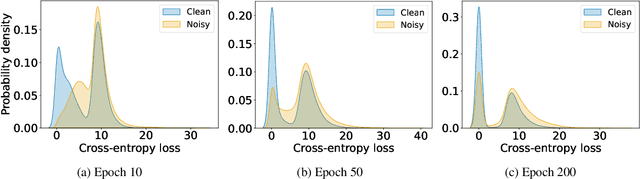
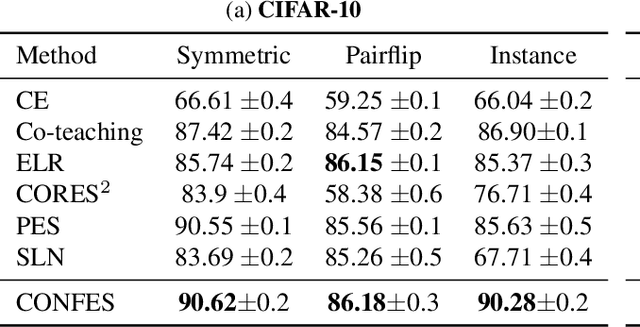
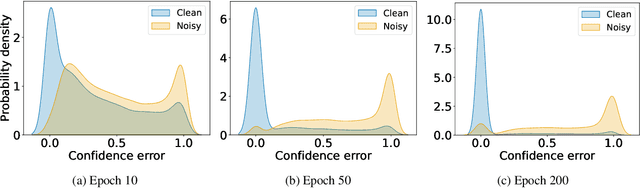
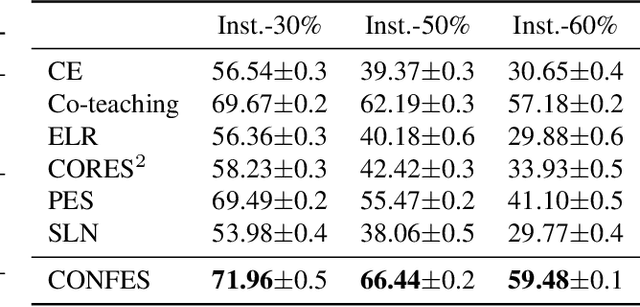
Abstract:In learning tasks with label noise, boosting model robustness against overfitting is a pivotal challenge because the model eventually memorizes labels including the noisy ones. Identifying the samples with corrupted labels and preventing the model from learning them is a promising approach to address this challenge. Per-sample training loss is a previously studied metric that considers samples with small loss as clean samples on which the model should be trained. In this work, we first demonstrate the ineffectiveness of this small-loss trick. Then, we propose a novel discriminator metric called confidence error and a sieving strategy called CONFES to effectively differentiate between the clean and noisy samples. We experimentally illustrate the superior performance of our proposed approach compared to recent studies on various settings such as synthetic and real-world label noise.
Kernel Normalized Convolutional Networks for Privacy-Preserving Machine Learning
Sep 30, 2022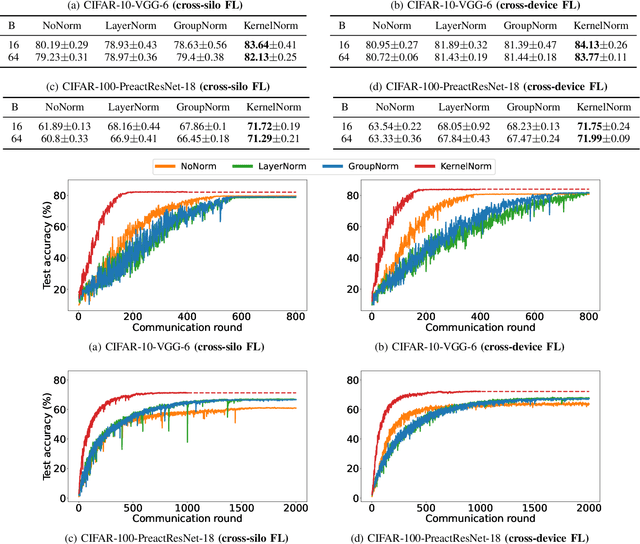
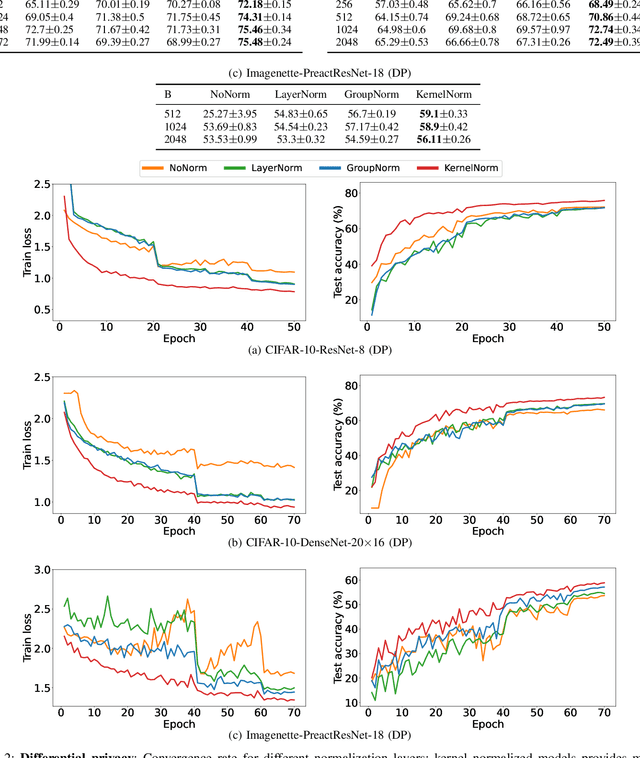
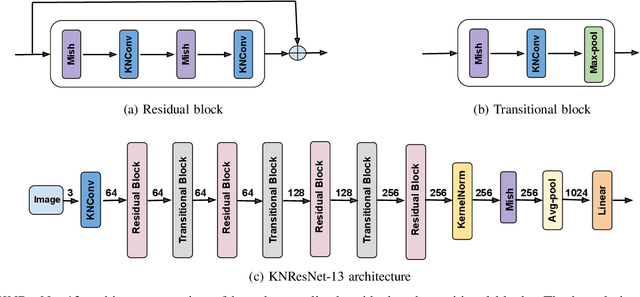

Abstract:Normalization is an important but understudied challenge in privacy-related application domains such as federated learning (FL) and differential privacy (DP). While the unsuitability of batch normalization for FL and DP has already been shown, the impact of the other normalization methods on the performance of federated or differentially private models is not well-known. To address this, we draw a performance comparison among layer normalization (LayerNorm), group normalization (GroupNorm), and the recently proposed kernel normalization (KernelNorm) in FL and DP settings. Our results indicate LayerNorm and GroupNorm provide no performance gain compared to the baseline (i.e. no normalization) for shallow models, but they considerably enhance performance of deeper models. KernelNorm, on the other hand, significantly outperforms its competitors in terms of accuracy and convergence rate (or communication efficiency) for both shallow and deeper models. Given these key observations, we propose a kernel normalized ResNet architecture called KNResNet-13 for differentially private learning environments. Using the proposed architecture, we provide new state-of-the-art accuracy values on the CIFAR-10 and Imagenette datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge