Claus Zimmer
Department of Diagnostic and Interventional Neuroradiology, School of Medicine, Klinikum rechts der Isar, Technical University of Munich, Germany
MLV$^2$-Net: Rater-Based Majority-Label Voting for Consistent Meningeal Lymphatic Vessel Segmentation
Nov 13, 2024



Abstract:Meningeal lymphatic vessels (MLVs) are responsible for the drainage of waste products from the human brain. An impairment in their functionality has been associated with aging as well as brain disorders like multiple sclerosis and Alzheimer's disease. However, MLVs have only recently been described for the first time in magnetic resonance imaging (MRI), and their ramified structure renders manual segmentation particularly difficult. Further, as there is no consistent notion of their appearance, human-annotated MLV structures contain a high inter-rater variability that most automatic segmentation methods cannot take into account. In this work, we propose a new rater-aware training scheme for the popular nnU-Net model, and we explore rater-based ensembling strategies for accurate and consistent segmentation of MLVs. This enables us to boost nnU-Net's performance while obtaining explicit predictions in different annotation styles and a rater-based uncertainty estimation. Our final model, MLV$^2$-Net, achieves a Dice similarity coefficient of 0.806 with respect to the human reference standard. The model further matches the human inter-rater reliability and replicates age-related associations with MLV volume.
ISLES 2024: The first longitudinal multimodal multi-center real-world dataset in (sub-)acute stroke
Aug 20, 2024
Abstract:Stroke remains a leading cause of global morbidity and mortality, placing a heavy socioeconomic burden. Over the past decade, advances in endovascular reperfusion therapy and the use of CT and MRI imaging for treatment guidance have significantly improved patient outcomes and are now standard in clinical practice. To develop machine learning algorithms that can extract meaningful and reproducible models of brain function for both clinical and research purposes from stroke images - particularly for lesion identification, brain health quantification, and prognosis - large, diverse, and well-annotated public datasets are essential. While only a few datasets with (sub-)acute stroke data were previously available, several large, high-quality datasets have recently been made publicly accessible. However, these existing datasets include only MRI data. In contrast, our dataset is the first to offer comprehensive longitudinal stroke data, including acute CT imaging with angiography and perfusion, follow-up MRI at 2-9 days, as well as acute and longitudinal clinical data up to a three-month outcome. The dataset includes a training dataset of n = 150 and a test dataset of n = 100 scans. Training data is publicly available, while test data will be used exclusively for model validation. We are making this dataset available as part of the 2024 edition of the Ischemic Stroke Lesion Segmentation (ISLES) challenge (https://www.isles-challenge.org/), which continuously aims to establish benchmark methods for acute and sub-acute ischemic stroke lesion segmentation, aiding in creating open stroke imaging datasets and evaluating cutting-edge image processing algorithms.
Approaching Peak Ground Truth
Dec 31, 2022

Abstract:Machine learning models are typically evaluated by computing similarity with reference annotations and trained by maximizing similarity with such. Especially in the bio-medical domain, annotations are subjective and suffer from low inter- and intra-rater reliability. Since annotations only reflect the annotation entity's interpretation of the real world, this can lead to sub-optimal predictions even though the model achieves high similarity scores. Here, the theoretical concept of Peak Ground Truth (PGT) is introduced. PGT marks the point beyond which an increase in similarity with the reference annotation stops translating to better Real World Model Performance (RWMP). Additionally, a quantitative technique to approximate PGT by computing inter- and intra-rater reliability is proposed. Finally, three categories of PGT-aware strategies to evaluate and improve model performance are reviewed.
ISLES 2022: A multi-center magnetic resonance imaging stroke lesion segmentation dataset
Jun 14, 2022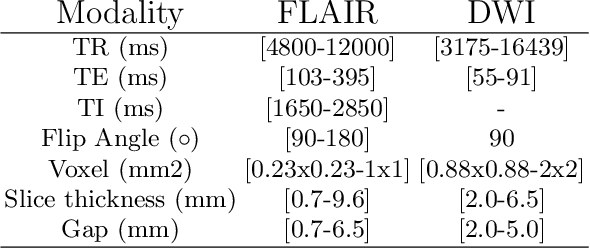
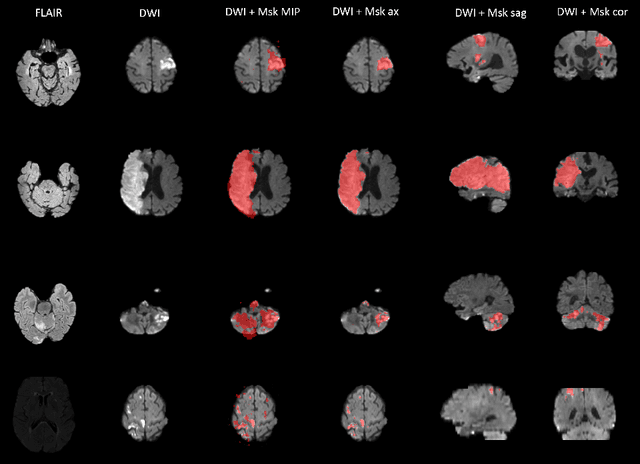

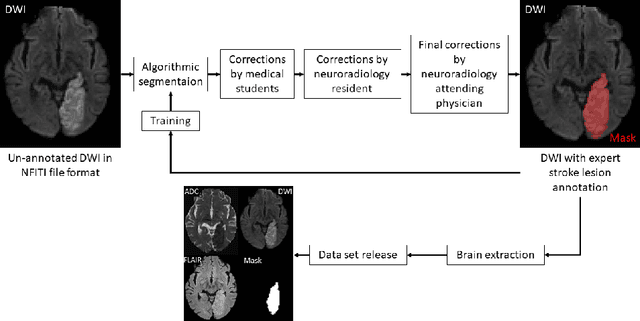
Abstract:Magnetic resonance imaging (MRI) is a central modality for stroke imaging. It is used upon patient admission to make treatment decisions such as selecting patients for intravenous thrombolysis or endovascular therapy. MRI is later used in the duration of hospital stay to predict outcome by visualizing infarct core size and location. Furthermore, it may be used to characterize stroke etiology, e.g. differentiation between (cardio)-embolic and non-embolic stroke. Computer based automated medical image processing is increasingly finding its way into clinical routine. Previous iterations of the Ischemic Stroke Lesion Segmentation (ISLES) challenge have aided in the generation of identifying benchmark methods for acute and sub-acute ischemic stroke lesion segmentation. Here we introduce an expert-annotated, multicenter MRI dataset for segmentation of acute to subacute stroke lesions. This dataset comprises 400 multi-vendor MRI cases with high variability in stroke lesion size, quantity and location. It is split into a training dataset of n=250 and a test dataset of n=150. All training data will be made publicly available. The test dataset will be used for model validation only and will not be released to the public. This dataset serves as the foundation of the ISLES 2022 challenge with the goal of finding algorithmic methods to enable the development and benchmarking of robust and accurate segmentation algorithms for ischemic stroke.
Deep Quality Estimation: Creating Surrogate Models for Human Quality Ratings
May 17, 2022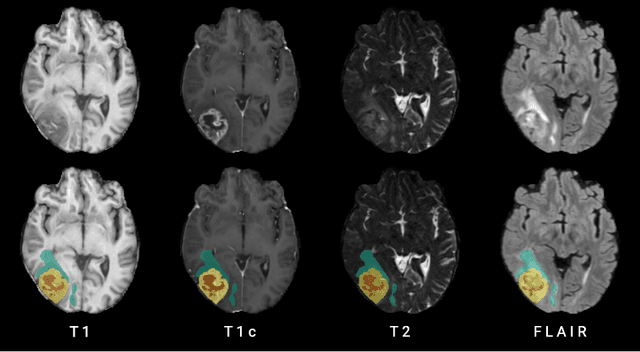
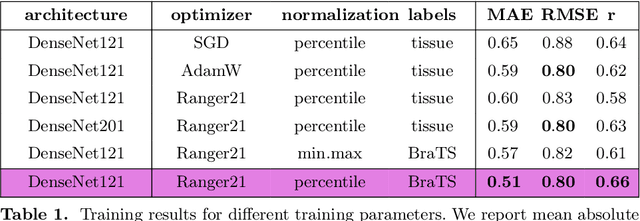
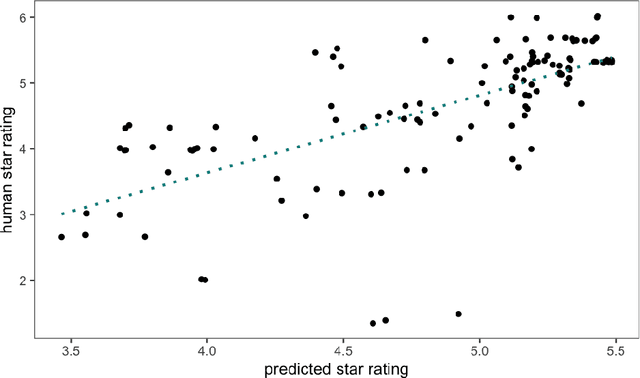
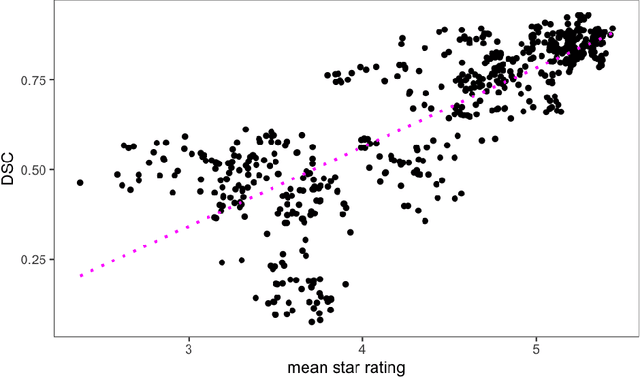
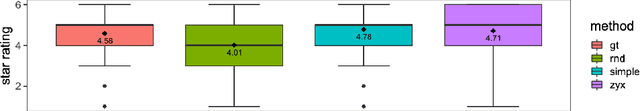
Abstract:Human ratings are abstract representations of segmentation quality. To approximate human quality ratings on scarce expert data, we train surrogate quality estimation models. We evaluate on a complex multi-class segmentation problem, specifically glioma segmentation following the BraTS annotation protocol. The training data features quality ratings from 15 expert neuroradiologists on a scale ranging from 1 to 6 stars for various computer-generated and manual 3D annotations. Even though the networks operate on 2D images and with scarce training data, we can approximate segmentation quality within a margin of error comparable to human intra-rater reliability. Segmentation quality prediction has broad applications. While an understanding of segmentation quality is imperative for successful clinical translation of automatic segmentation quality algorithms, it can play an essential role in training new segmentation models. Due to the split-second inference times, it can be directly applied within a loss function or as a fully-automatic dataset curation mechanism in a federated learning setting.
blob loss: instance imbalance aware loss functions for semantic segmentation
May 17, 2022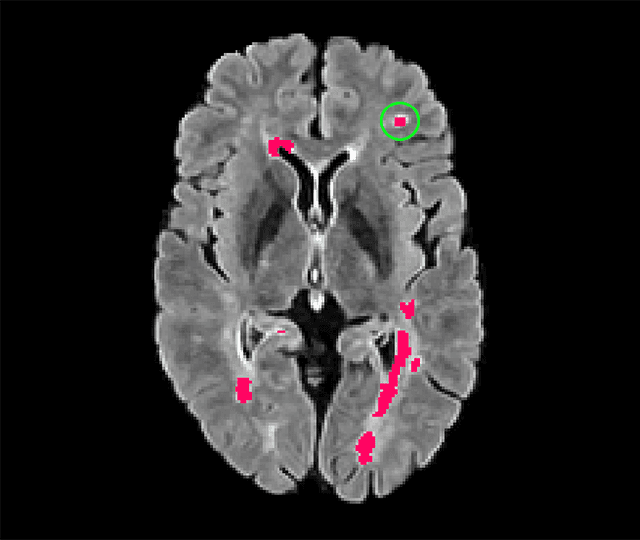
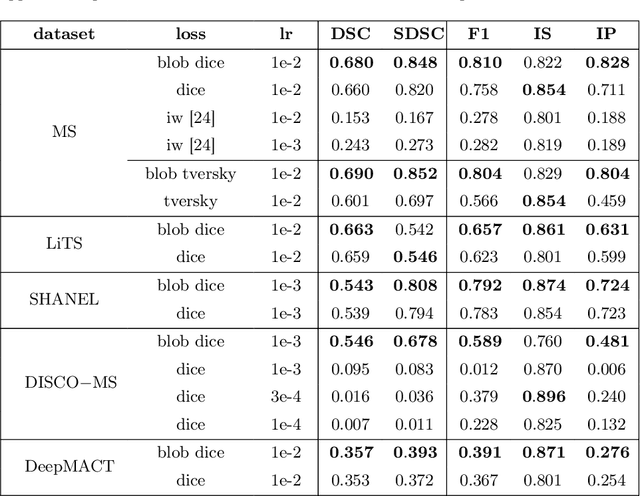

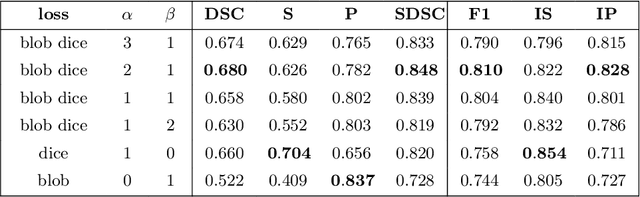
Abstract:Deep convolutional neural networks have proven to be remarkably effective in semantic segmentation tasks. Most popular loss functions were introduced targeting improved volumetric scores, such as the Sorensen Dice coefficient. By design, DSC can tackle class imbalance; however, it does not recognize instance imbalance within a class. As a result, a large foreground instance can dominate minor instances and still produce a satisfactory Sorensen Dice coefficient. Nevertheless, missing out on instances will lead to poor detection performance. This represents a critical issue in applications such as disease progression monitoring. For example, it is imperative to locate and surveil small-scale lesions in the follow-up of multiple sclerosis patients. We propose a novel family of loss functions, nicknamed blob loss, primarily aimed at maximizing instance-level detection metrics, such as F1 score and sensitivity. Blob loss is designed for semantic segmentation problems in which the instances are the connected components within a class. We extensively evaluate a DSC-based blob loss in five complex 3D semantic segmentation tasks featuring pronounced instance heterogeneity in terms of texture and morphology. Compared to soft Dice loss, we achieve 5 percent improvement for MS lesions, 3 percent improvement for liver tumor, and an average 2 percent improvement for Microscopy segmentation tasks considering F1 score.
A Deep Learning Approach to Predicting Collateral Flow in Stroke Patients Using Radiomic Features from Perfusion Images
Oct 24, 2021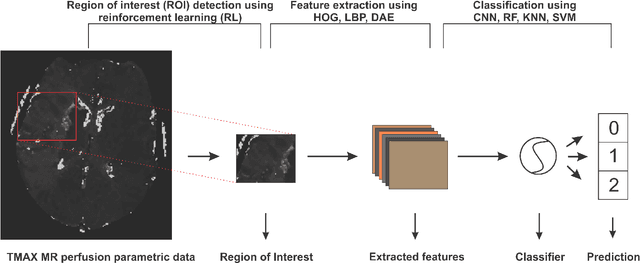
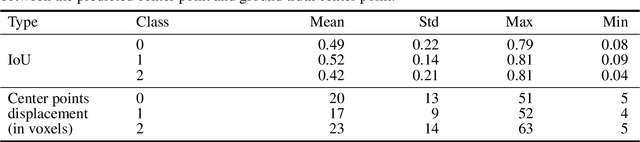
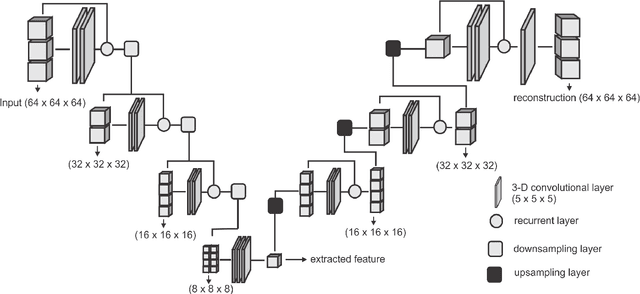
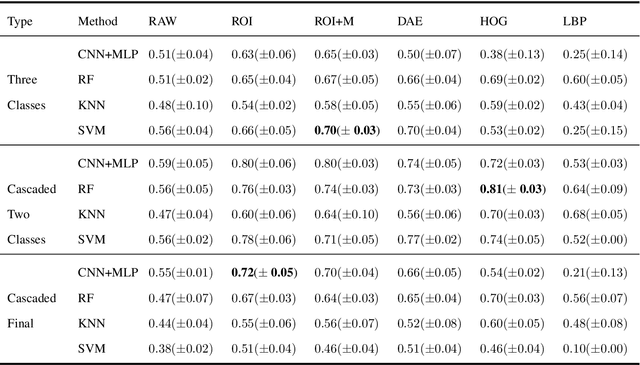
Abstract:Collateral circulation results from specialized anastomotic channels which are capable of providing oxygenated blood to regions with compromised blood flow caused by ischemic injuries. The quality of collateral circulation has been established as a key factor in determining the likelihood of a favorable clinical outcome and goes a long way to determine the choice of stroke care model - that is the decision to transport or treat eligible patients immediately. Though there exist several imaging methods and grading criteria for quantifying collateral blood flow, the actual grading is mostly done through manual inspection of the acquired images. This approach is associated with a number of challenges. First, it is time-consuming - the clinician needs to scan through several slices of images to ascertain the region of interest before deciding on what severity grade to assign to a patient. Second, there is a high tendency for bias and inconsistency in the final grade assigned to a patient depending on the experience level of the clinician. We present a deep learning approach to predicting collateral flow grading in stroke patients based on radiomic features extracted from MR perfusion data. First, we formulate a region of interest detection task as a reinforcement learning problem and train a deep learning network to automatically detect the occluded region within the 3D MR perfusion volumes. Second, we extract radiomic features from the obtained region of interest through local image descriptors and denoising auto-encoders. Finally, we apply a convolutional neural network and other machine learning classifiers to the extracted radiomic features to automatically predict the collateral flow grading of the given patient volume as one of three severity classes - no flow (0), moderate flow (1), and good flow (2)...
A Computed Tomography Vertebral Segmentation Dataset with Anatomical Variations and Multi-Vendor Scanner Data
Mar 10, 2021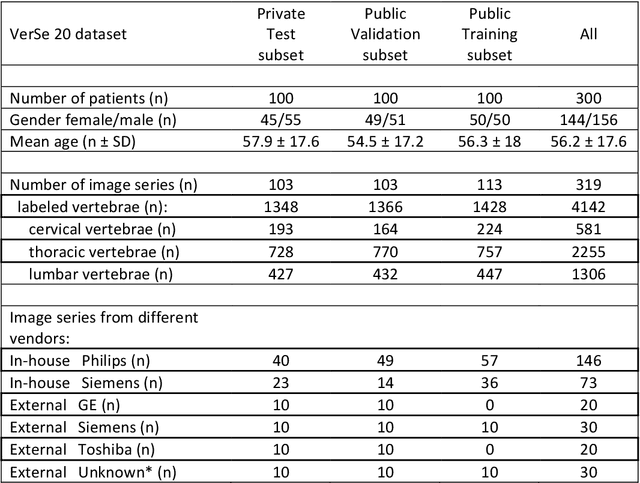
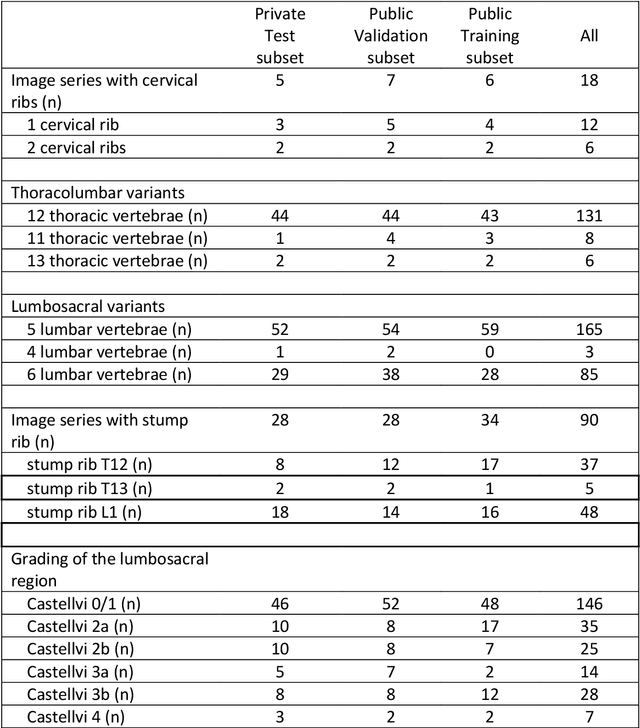
Abstract:With the advent of deep learning algorithms, fully automated radiological image analysis is within reach. In spine imaging, several atlas- and shape-based as well as deep learning segmentation algorithms have been proposed, allowing for subsequent automated analysis of morphology and pathology. The first Large Scale Vertebrae Segmentation Challenge (VerSe 2019) showed that these perform well on normal anatomy, but fail in variants not frequently present in the training dataset. Building on that experience, we report on the largely increased VerSe 2020 dataset and results from the second iteration of the VerSe challenge (MICCAI 2020, Lima, Peru). VerSe 2020 comprises annotated spine computed tomography (CT) images from 300 subjects with 4142 fully visualized and annotated vertebrae, collected across multiple centres from four different scanner manufacturers, enriched with cases that exhibit anatomical variants such as enumeration abnormalities (n=77) and transitional vertebrae (n=161). Metadata includes vertebral labelling information, voxel-level segmentation masks obtained with a human-machine hybrid algorithm and anatomical ratings, to enable the development and benchmarking of robust and accurate segmentation algorithms.
Are we using appropriate segmentation metrics? Identifying correlates of human expert perception for CNN training beyond rolling the DICE coefficient
Mar 10, 2021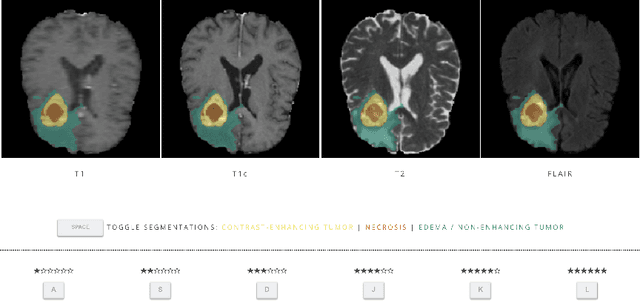

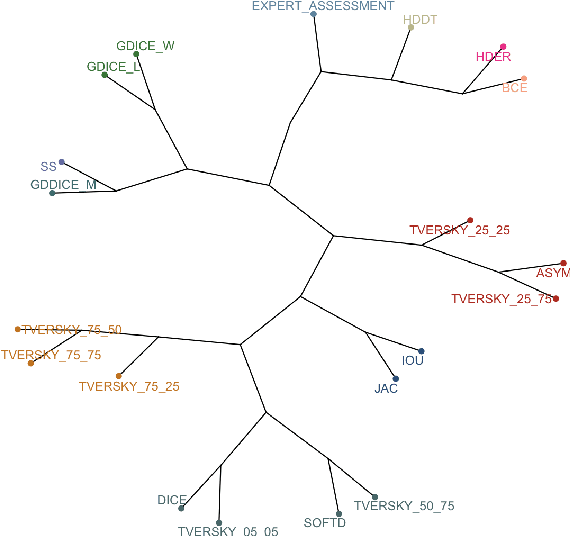
Abstract:In this study, we explore quantitative correlates of qualitative human expert perception. We discover that current quality metrics and loss functions, considered for biomedical image segmentation tasks, correlate moderately with segmentation quality assessment by experts, especially for small yet clinically relevant structures, such as the enhancing tumor in brain glioma. We propose a method employing classical statistics and experimental psychology to create complementary compound loss functions for modern deep learning methods, towards achieving a better fit with human quality assessment. When training a CNN for delineating adult brain tumor in MR images, all four proposed loss candidates outperform the established baselines on the clinically important and hardest to segment enhancing tumor label, while maintaining performance for other label channels.
DeepVesselNet: Vessel Segmentation, Centerline Prediction, and Bifurcation Detection in 3-D Angiographic Volumes
Mar 25, 2018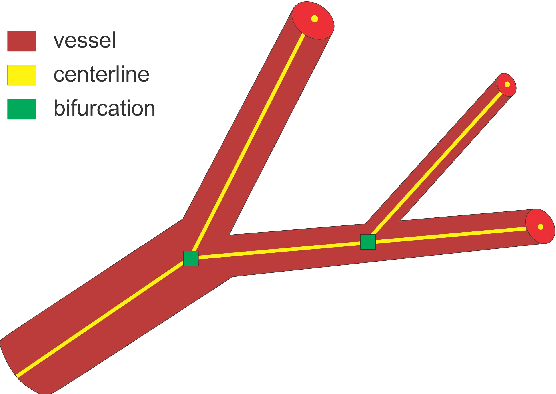
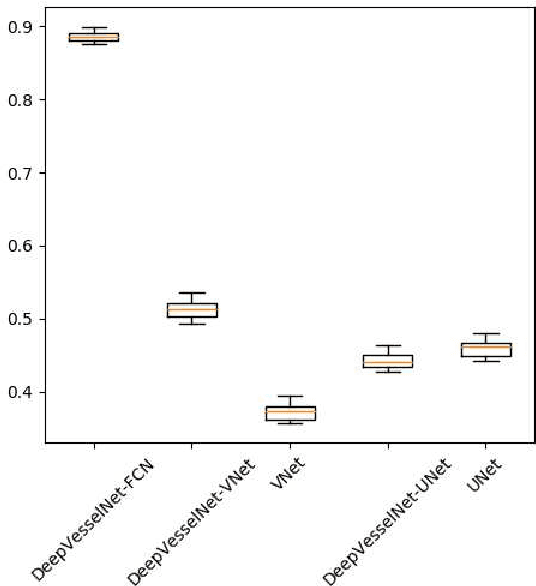
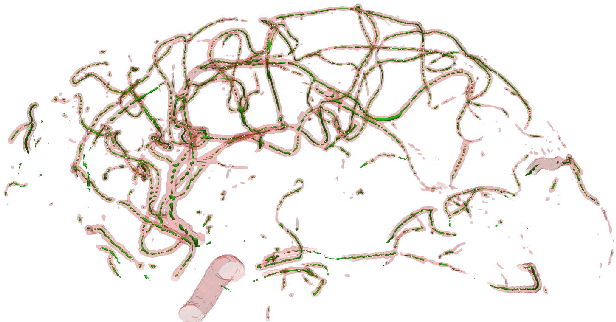
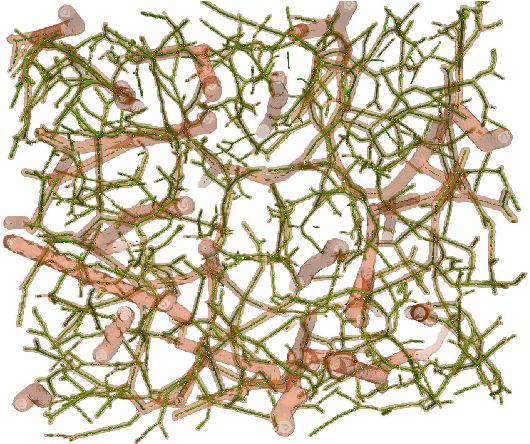
Abstract:We present DeepVesselNet, an architecture tailored to the challenges to be addressed when extracting vessel networks and corresponding features in 3-D angiography using deep learning. We discuss the problems of low execution speed and high memory requirements associated with full 3-D convolutional networks, high class imbalance arising from low percentage (less than 3%) of vessel voxels, and unavailability of accurately annotated training data - and offer solutions that are the building blocks of DeepVesselNet. First, we formulate 2-D orthogonal cross-hair filters which make use of 3-D context information. Second, we introduce a class balancing cross-entropy score with false positive rate correction to handle the high class imbalance and high false positive rate problems associated with existing loss functions. Finally, we generate synthetic dataset using a computational angiogenesis model, capable of generating vascular networks under physiological constraints on local network structure and topology, and use these data for transfer learning. DeepVesselNet is optimized for segmenting vessels, predicting centerlines, and localizing bifurcations. We test the performance on a range of angiographic volumes including clinical Time-of-Flight MRA data of the human brain, as well as synchrotron radiation X-ray tomographic microscopy scans of the rat brain. Our experiments show that, by replacing 3-D filters with 2-D orthogonal cross-hair filters in our network, speed is improved by 23% while accuracy is maintained. Our class balancing metric is crucial for training the network and pre-training with synthetic data helps in early convergence of the training process.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge