Christos Davatzikos
from the iSTAGING consortium, for the ADNI
Extraction of Hierarchical Functional Connectivity Components in human brain using Adversarial Learning
Apr 20, 2021
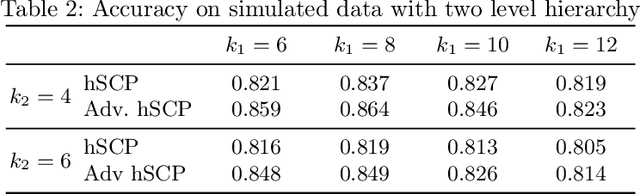
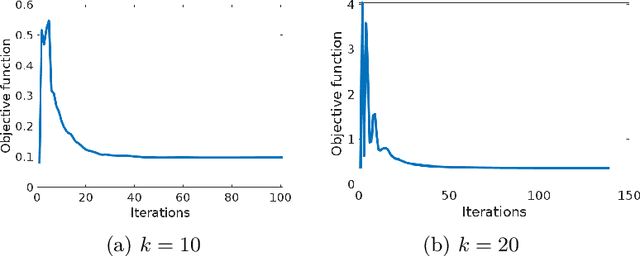
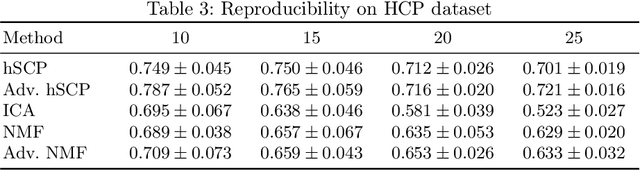
Abstract:The estimation of sparse hierarchical components reflecting patterns of the brain's functional connectivity from rsfMRI data can contribute to our understanding of the brain's functional organization, and can lead to biomarkers of diseases. However, inter-scanner variations and other confounding factors pose a challenge to the robust and reproducible estimation of functionally-interpretable brain networks, and especially to reproducible biomarkers. Moreover, the brain is believed to be organized hierarchically, and hence single-scale decompositions miss this hierarchy. The paper aims to use current advancements in adversarial learning to estimate interpretable hierarchical patterns in the human brain using rsfMRI data, which are robust to "adversarial effects" such as inter-scanner variations. We write the estimation problem as a minimization problem and solve it using alternating updates. Extensive experiments on simulation and a real-world dataset show high reproducibility of the components compared to other well-known methods.
Embracing the Disharmony in Heterogeneous Medical Data
Mar 23, 2021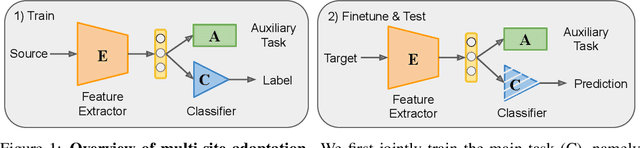
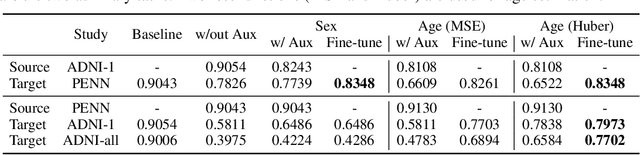
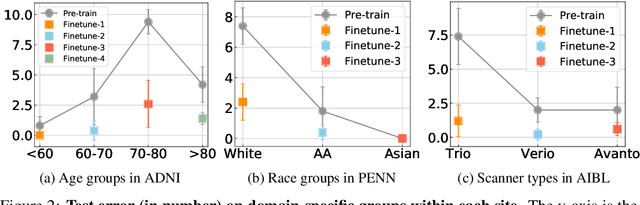
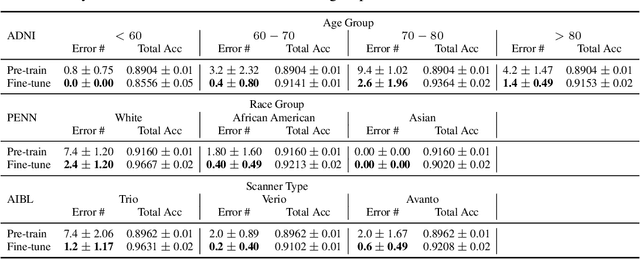
Abstract:Heterogeneity in medical imaging data is often tackled, in the context of machine learning, using domain invariance, i.e. deriving models that are robust to domain shifts, which can be both within domain (e.g. demographics) and across domains (e.g. scanner/protocol characteristics). However this approach can be detrimental to performance because it necessitates averaging across intra-class variability and reduces discriminatory power of learned models, in order to achieve better intra- and inter-domain generalization. This paper instead embraces the heterogeneity and treats it as a multi-task learning problem to explicitly adapt trained classifiers to both inter-site and intra-site heterogeneity. We demonstrate that the error of a base classifier on challenging 3D brain magnetic resonance imaging (MRI) datasets can be reduced by 2-3 times, in certain tasks, by adapting to the specific demographics of the patients, and different acquisition protocols. Learning the characteristics of domain shifts is achieved via auxiliary learning tasks leveraging commonly available data and variables, e.g. demographics. In our experiments, we use gender classification and age regression as auxiliary tasks helping the network weights trained on a source site adapt to data from a target site; we show that this approach improves classification accuracy by 5-30 % across different datasets on the main classification tasks, e.g. disease classification.
GaNDLF: A Generally Nuanced Deep Learning Framework for Scalable End-to-End Clinical Workflows in Medical Imaging
Feb 26, 2021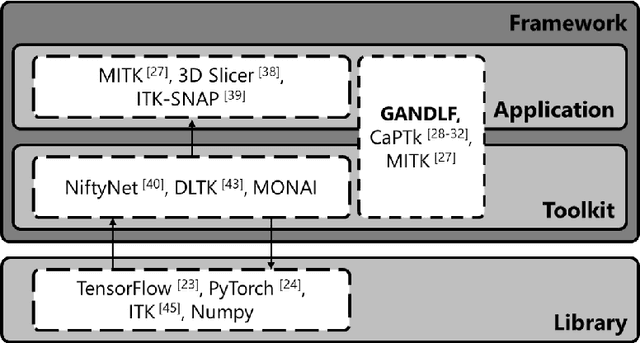
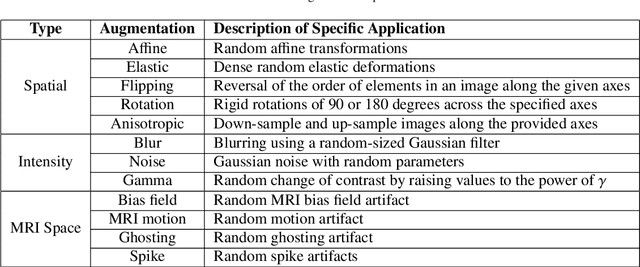

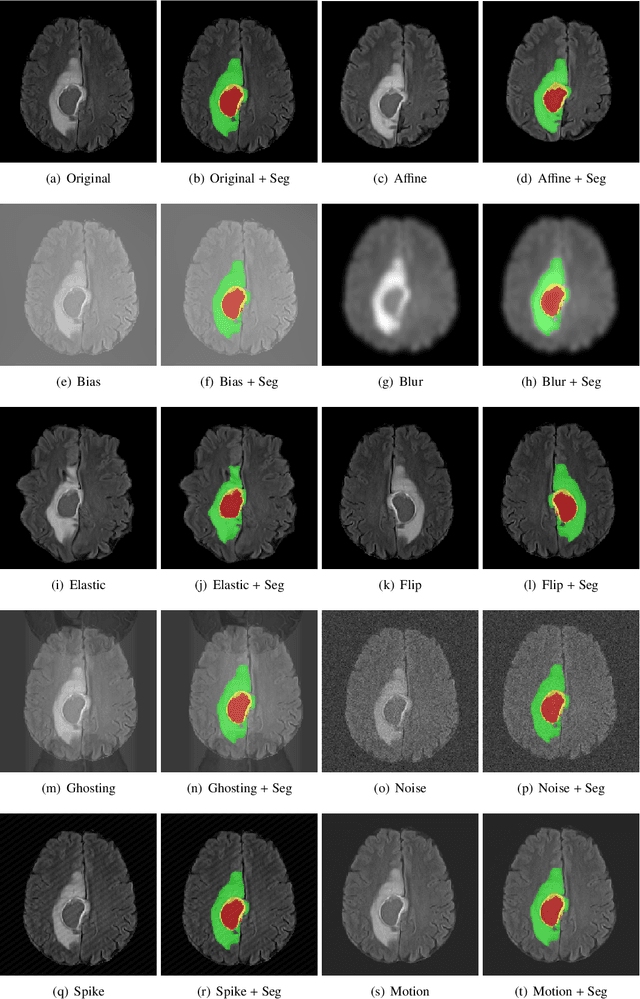
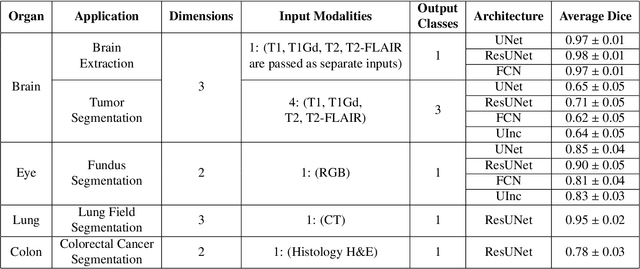
Abstract:Deep Learning (DL) has greatly highlighted the potential impact of optimized machine learning in both the scientific and clinical communities. The advent of open-source DL libraries from major industrial entities, such as TensorFlow (Google), PyTorch (Facebook), and MXNet (Apache), further contributes to DL promises on the democratization of computational analytics. However, increased technical and specialized background is required to develop DL algorithms, and the variability of implementation details hinders their reproducibility. Towards lowering the barrier and making the mechanism of DL development, training, and inference more stable, reproducible, and scalable, without requiring an extensive technical background, this manuscript proposes the \textbf{G}ener\textbf{a}lly \textbf{N}uanced \textbf{D}eep \textbf{L}earning \textbf{F}ramework (GaNDLF). With built-in support for $k$-fold cross-validation, data augmentation, multiple modalities and output classes, and multi-GPU training, as well as the ability to work with both radiographic and histologic imaging, GaNDLF aims to provide an end-to-end solution for all DL-related tasks, to tackle problems in medical imaging and provide a robust application framework for deployment in clinical workflows.
Disentangling brain heterogeneity via semi-supervised deep-learning and MRI: dimensional representations of Alzheimer's Disease
Feb 24, 2021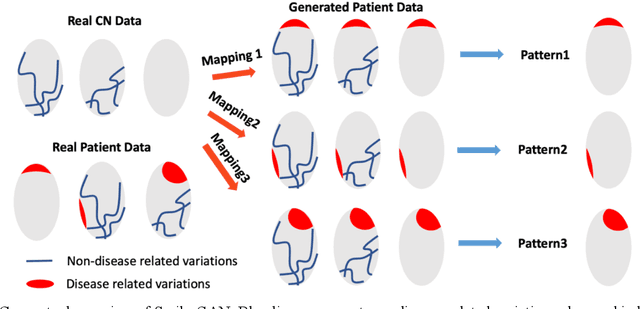
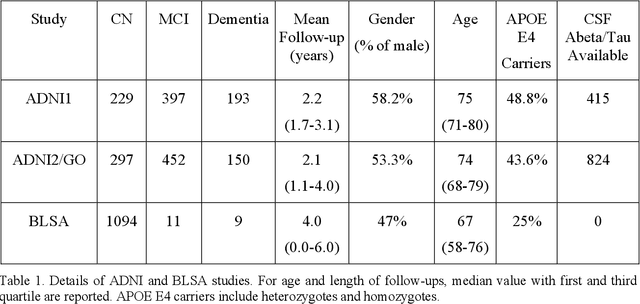
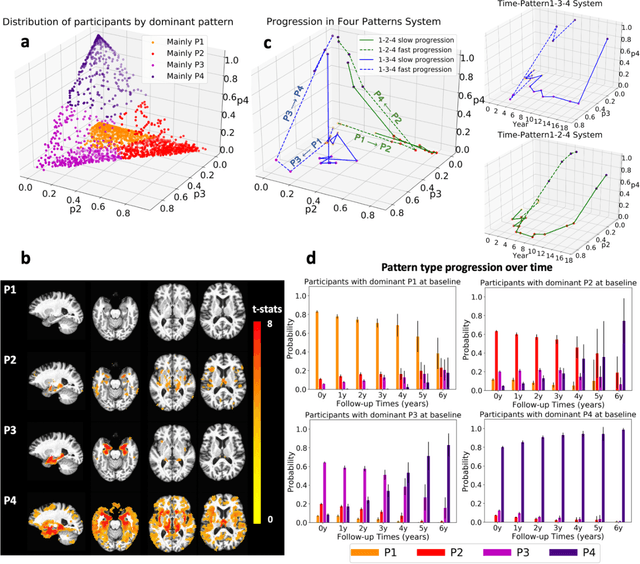
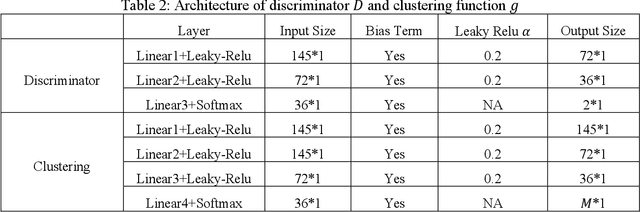
Abstract:Heterogeneity of brain diseases is a challenge for precision diagnosis/prognosis. We describe and validate Smile-GAN (SeMI-supervised cLustEring-Generative Adversarial Network), a novel semi-supervised deep-clustering method, which dissects neuroanatomical heterogeneity, enabling identification of disease subtypes via their imaging signatures relative to controls. When applied to MRIs (2 studies; 2,832 participants; 8,146 scans) including cognitively normal individuals and those with cognitive impairment and dementia, Smile-GAN identified 4 neurodegenerative patterns/axes: P1, normal anatomy and highest cognitive performance; P2, mild/diffuse atrophy and more prominent executive dysfunction; P3, focal medial temporal atrophy and relatively greater memory impairment; P4, advanced neurodegeneration. Further application to longitudinal data revealed two distinct progression pathways: P1$\rightarrow$P2$\rightarrow$P4 and P1$\rightarrow$P3$\rightarrow$P4. Baseline expression of these patterns predicted the pathway and rate of future neurodegeneration. Pattern expression offered better yet complementary performance in predicting clinical progression, compared to amyloid/tau. These deep-learning derived biomarkers offer promise for precision diagnostics and targeted clinical trial recruitment.
Medical Image Harmonization Using Deep Learning Based Canonical Mapping: Toward Robust and Generalizable Learning in Imaging
Oct 11, 2020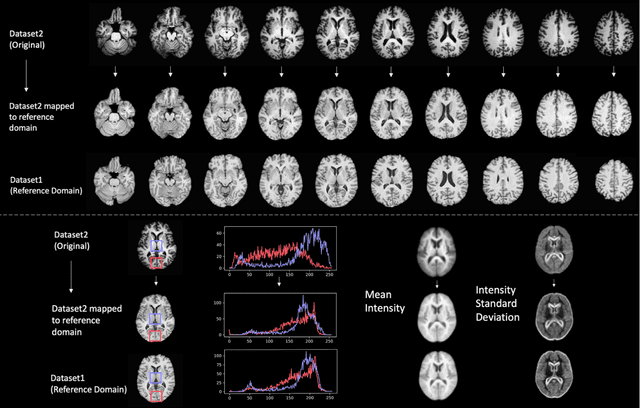
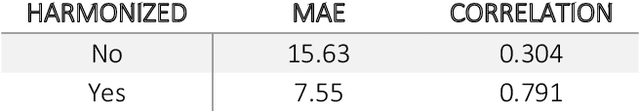
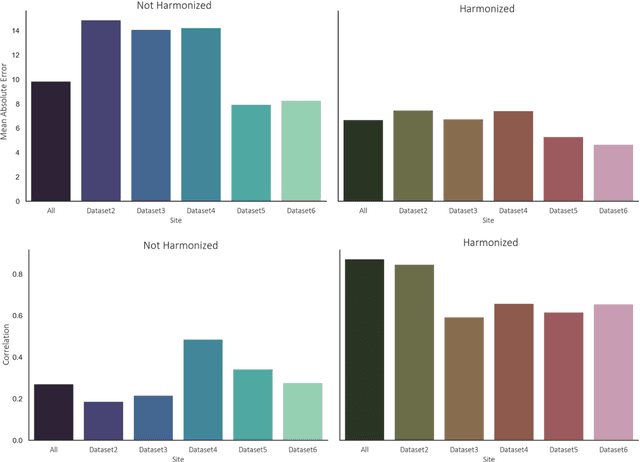
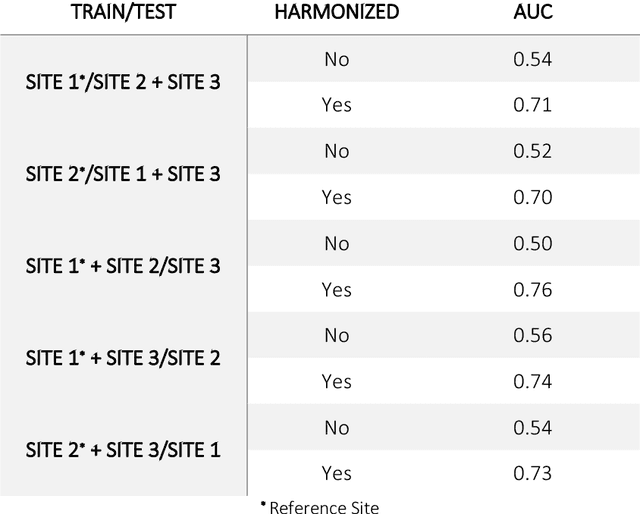
Abstract:Conventional and deep learning-based methods have shown great potential in the medical imaging domain, as means for deriving diagnostic, prognostic, and predictive biomarkers, and by contributing to precision medicine. However, these methods have yet to see widespread clinical adoption, in part due to limited generalization performance across various imaging devices, acquisition protocols, and patient populations. In this work, we propose a new paradigm in which data from a diverse range of acquisition conditions are "harmonized" to a common reference domain, where accurate model learning and prediction can take place. By learning an unsupervised image to image canonical mapping from diverse datasets to a reference domain using generative deep learning models, we aim to reduce confounding data variation while preserving semantic information, thereby rendering the learning task easier in the reference domain. We test this approach on two example problems, namely MRI-based brain age prediction and classification of schizophrenia, leveraging pooled cohorts of neuroimaging MRI data spanning 9 sites and 9701 subjects. Our results indicate a substantial improvement in these tasks in out-of-sample data, even when training is restricted to a single site.
A Deep Network for Joint Registration and Reconstruction of Images with Pathologies
Aug 17, 2020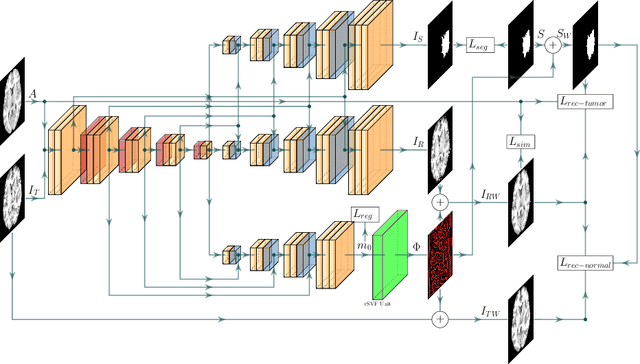
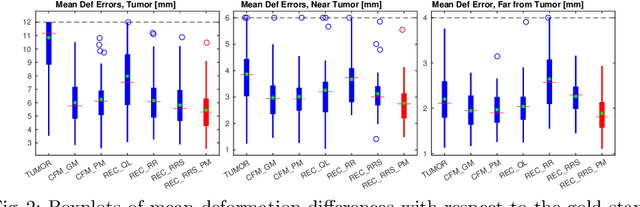
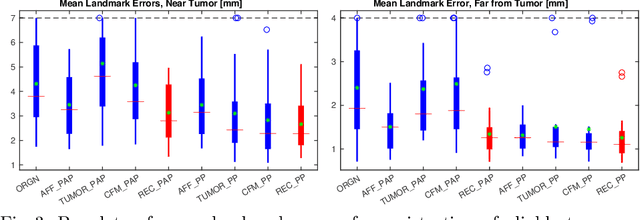
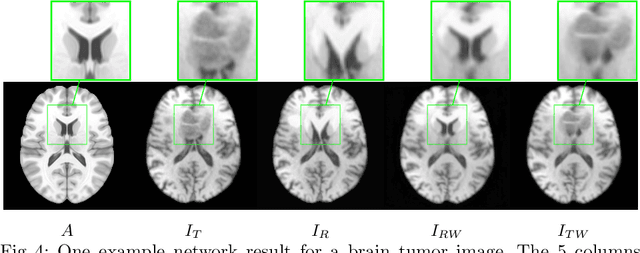
Abstract:Registration of images with pathologies is challenging due to tissue appearance changes and missing correspondences caused by the pathologies. Moreover, mass effects as observed for brain tumors may displace tissue, creating larger deformations over time than what is observed in a healthy brain. Deep learning models have successfully been applied to image registration to offer dramatic speed up and to use surrogate information (e.g., segmentations) during training. However, existing approaches focus on learning registration models using images from healthy patients. They are therefore not designed for the registration of images with strong pathologies for example in the context of brain tumors, and traumatic brain injuries. In this work, we explore a deep learning approach to register images with brain tumors to an atlas. Our model learns an appearance mapping from images with tumors to the atlas, while simultaneously predicting the transformation to atlas space. Using separate decoders, the network disentangles the tumor mass effect from the reconstruction of quasi-normal images. Results on both synthetic and real brain tumor scans show that our approach outperforms cost function masking for registration to the atlas and that reconstructed quasi-normal images can be used for better longitudinal registrations.
A review of deep learning in medical imaging: Image traits, technology trends, case studies with progress highlights, and future promises
Aug 02, 2020



Abstract:Since its renaissance, deep learning has been widely used in various medical imaging tasks and has achieved remarkable success in many medical imaging applications, thereby propelling us into the so-called artificial intelligence (AI) era. It is known that the success of AI is mostly attributed to the availability of big data with annotations for a single task and the advances in high performance computing. However, medical imaging presents unique challenges that confront deep learning approaches. In this survey paper, we first highlight both clinical needs and technical challenges in medical imaging and describe how emerging trends in deep learning are addressing these issues. We cover the topics of network architecture, sparse and noisy labels, federating learning, interpretability, uncertainty quantification, etc. Then, we present several case studies that are commonly found in clinical practice, including digital pathology and chest, brain, cardiovascular, and abdominal imaging. Rather than presenting an exhaustive literature survey, we instead describe some prominent research highlights related to these case study applications. We conclude with a discussion and presentation of promising future directions.
MAGIC: Multi-scale Heterogeneity Analysis and Clustering for Brain Diseases
Jul 10, 2020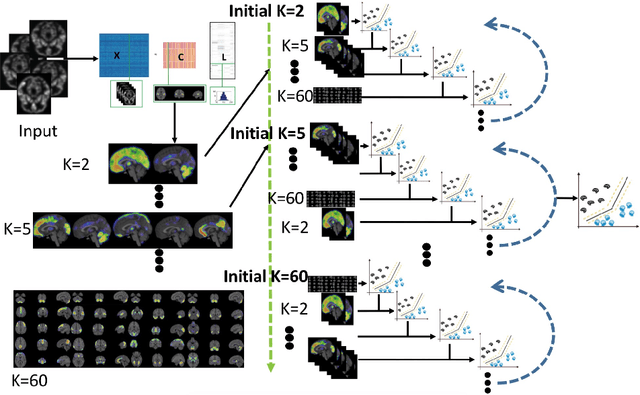
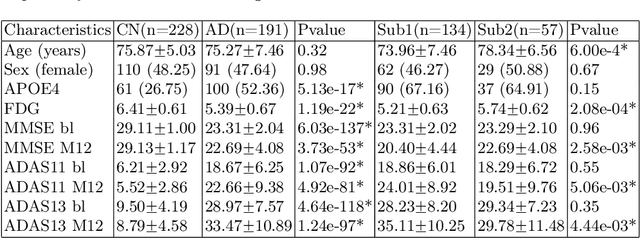
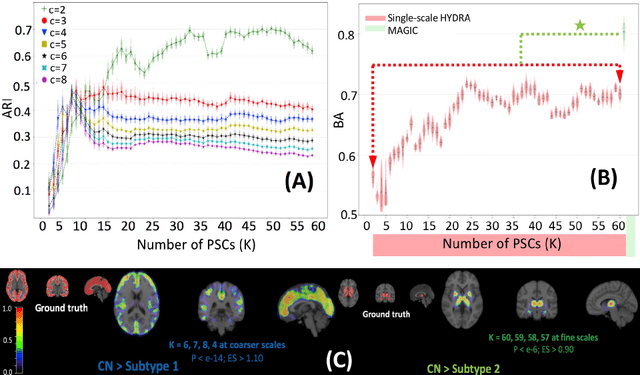
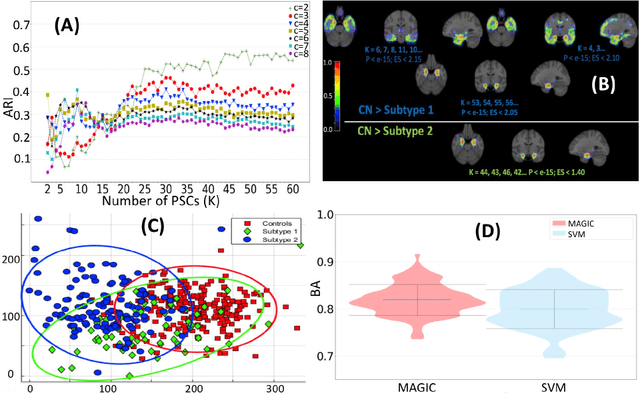
Abstract:There is a growing amount of clinical, anatomical and functional evidence for the heterogeneous presentation of neuropsychiatric and neurodegenerative diseases such as schizophrenia and Alzheimers Disease (AD). Elucidating distinct subtypes of diseases allows a better understanding of neuropathogenesis and enables the possibility of developing targeted treatment programs. Recent semi-supervised clustering techniques have provided a data-driven way to understand disease heterogeneity. However, existing methods do not take into account that subtypes of the disease might present themselves at different spatial scales across the brain. Here, we introduce a novel method, MAGIC, to uncover disease heterogeneity by leveraging multi-scale clustering. We first extract multi-scale patterns of structural covariance (PSCs) followed by a semi-supervised clustering with double cyclic block-wise optimization across different scales of PSCs. We validate MAGIC using simulated heterogeneous neuroanatomical data and demonstrate its clinical potential by exploring the heterogeneity of AD using T1 MRI scans of 228 cognitively normal (CN) and 191 patients. Our results indicate two main subtypes of AD with distinct atrophy patterns that consist of both fine-scale atrophy in the hippocampus as well as large-scale atrophy in cortical regions. The evidence for the heterogeneity is further corroborated by the clinical evaluation of two subtypes, which indicates that there is a subpopulation of AD patients that tend to be younger and decline faster in cognitive performance relative to the other subpopulation, which tends to be older and maintains a relatively steady decline in cognitive abilities.
Smile-GANs: Semi-supervised clustering via GANs for dissecting brain disease heterogeneity from medical images
Jun 27, 2020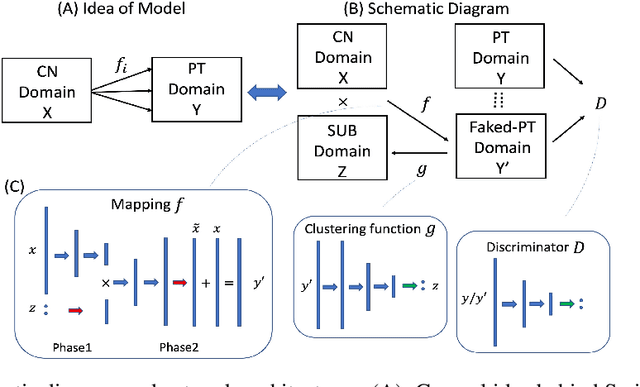

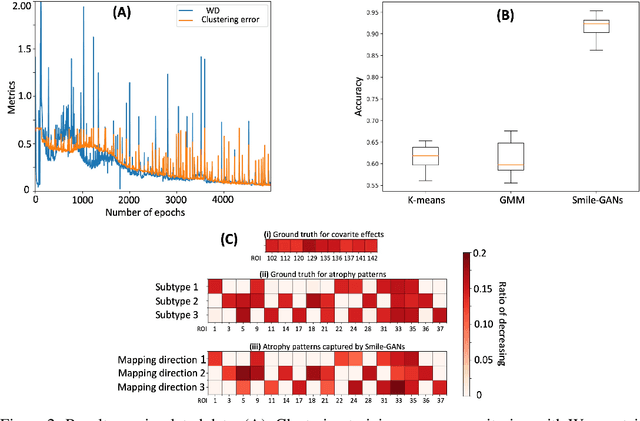
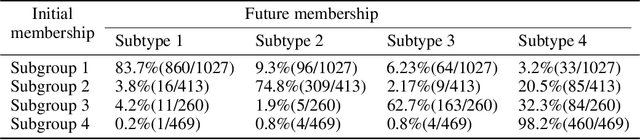
Abstract:Machine learning methods applied to complex biomedical data has enabled the construction of disease signatures of diagnostic/prognostic value. However, less attention has been given to understanding disease heterogeneity. Semi-supervised clustering methods can address this problem by estimating multiple transformations from a (e.g. healthy) control (CN) group to a patient (PT) group, seeking to capture the heterogeneity of underlying pathlogic processes. Herein, we propose a novel method, Smile-GANs (SeMi-supervIsed cLustEring via GANs), for semi-supervised clustering, and apply it to brain MRI scans. Smile-GANs first learns multiple distinct mappings by generating PT from CN, with each mapping characterizing one relatively distinct pathological pattern. Moreover, a clustering model is trained interactively with mapping functions to assign PT into corresponding subtype memberships. Using relaxed assumptions on PT/CN data distribution and imposing mapping non-linearity, Smile-GANs captures heterogeneous differences in distribution between the CN and PT domains. We first validate Smile-GANs using simulated data, subsequently on real data, by demonstrating its potential in characterizing heterogeneity in Alzheimer's Disease (AD) and its prodromal phases. The model was first trained using baseline MRIs from the ADNI2 database and then applied to longitudinal data from ADNI1 and BLSA. Four robust subtypes with distinct neuroanatomical patterns were discovered: 1) normal brain, 2) diffuse atrophy atypical of AD, 3) focal medial temporal lobe atrophy, 4) typical-AD. Further longitudinal analyses discover two distinct progressive pathways from prodromal to full AD: i) subtypes 1 - 2 - 4, and ii) subtypes 1 - 3 - 4. Although demonstrated on an important biomedical problem, Smile-GANs is general and can find application in many biomedical and other domains.
DeepMRSeg: A convolutional deep neural network for anatomy and abnormality segmentation on MR images
Jul 03, 2019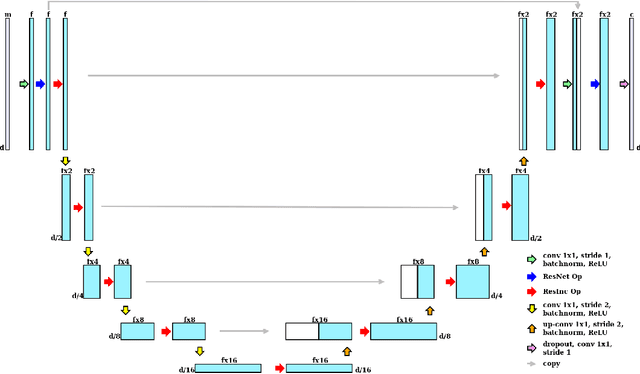
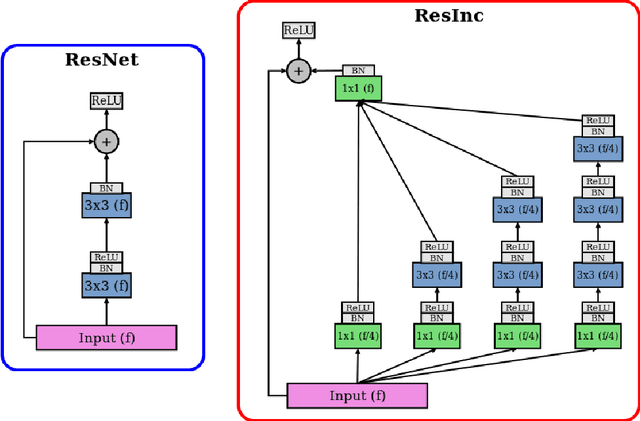
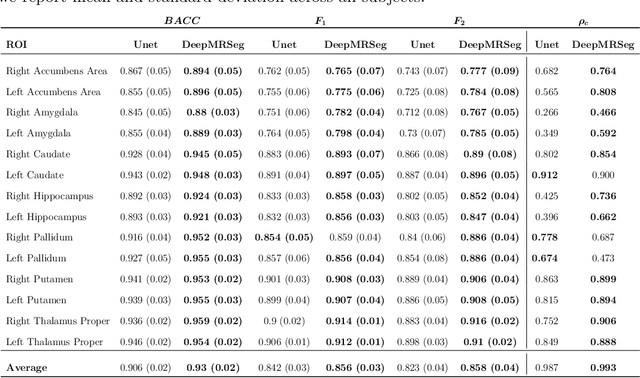
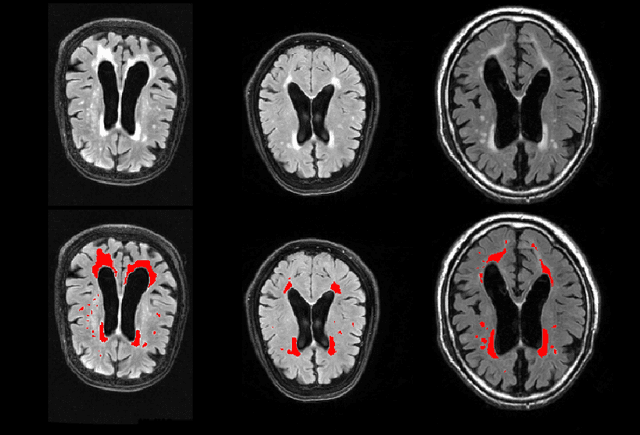
Abstract:Segmentation has been a major task in neuroimaging. A large number of automated methods have been developed for segmenting healthy and diseased brain tissues. In recent years, deep learning techniques have attracted a lot of attention as a result of their high accuracy in different segmentation problems. We present a new deep learning based segmentation method, DeepMRSeg, that can be applied in a generic way to a variety of segmentation tasks. The proposed architecture combines recent advances in the field of biomedical image segmentation and computer vision. We use a modified UNet architecture that takes advantage of multiple convolution filter sizes to achieve multi-scale feature extraction adaptive to the desired segmentation task. Importantly, our method operates on minimally processed raw MRI scan. We validated our method on a wide range of segmentation tasks, including white matter lesion segmentation, segmentation of deep brain structures and hippocampus segmentation. We provide code and pre-trained models to allow researchers apply our method on their own datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge