Bernhard Kainz
Biomedical Image Analysis Group, Department of Computing, Imperial College London
Zero-Shot Anomaly Detection with Pre-trained Segmentation Models
Jun 15, 2023Abstract:This technical report outlines our submission to the zero-shot track of the Visual Anomaly and Novelty Detection (VAND) 2023 Challenge. Building on the performance of the WINCLIP framework, we aim to enhance the system's localization capabilities by integrating zero-shot segmentation models. In addition, we perform foreground instance segmentation which enables the model to focus on the relevant parts of the image, thus allowing the models to better identify small or subtle deviations. Our pipeline requires no external data or information, allowing for it to be directly applied to new datasets. Our team (Variance Vigilance Vanguard) ranked third in the zero-shot track of the VAND challenge, and achieve an average F1-max score of 81.5/24.2 at a sample/pixel level on the VisA dataset.
Quantifying Sample Anonymity in Score-Based Generative Models with Adversarial Fingerprinting
Jun 02, 2023



Abstract:Recent advances in score-based generative models have led to a huge spike in the development of downstream applications using generative models ranging from data augmentation over image and video generation to anomaly detection. Despite publicly available trained models, their potential to be used for privacy preserving data sharing has not been fully explored yet. Training diffusion models on private data and disseminating the models and weights rather than the raw dataset paves the way for innovative large-scale data-sharing strategies, particularly in healthcare, where safeguarding patients' personal health information is paramount. However, publishing such models without individual consent of, e.g., the patients from whom the data was acquired, necessitates guarantees that identifiable training samples will never be reproduced, thus protecting personal health data and satisfying the requirements of policymakers and regulatory bodies. This paper introduces a method for estimating the upper bound of the probability of reproducing identifiable training images during the sampling process. This is achieved by designing an adversarial approach that searches for anatomic fingerprints, such as medical devices or dermal art, which could potentially be employed to re-identify training images. Our method harnesses the learned score-based model to estimate the probability of the entire subspace of the score function that may be utilized for one-to-one reproduction of training samples. To validate our estimates, we generate anomalies containing a fingerprint and investigate whether generated samples from trained generative models can be uniquely mapped to the original training samples. Overall our results show that privacy-breaching images are reproduced at sampling time if the models were trained without care.
Realistic Data Enrichment for Robust Image Segmentation in Histopathology
Apr 19, 2023


Abstract:Poor performance of quantitative analysis in histopathological Whole Slide Images (WSI) has been a significant obstacle in clinical practice. Annotating large-scale WSIs manually is a demanding and time-consuming task, unlikely to yield the expected results when used for fully supervised learning systems. Rarely observed disease patterns and large differences in object scales are difficult to model through conventional patient intake. Prior methods either fall back to direct disease classification, which only requires learning a few factors per image, or report on average image segmentation performance, which is highly biased towards majority observations. Geometric image augmentation is commonly used to improve robustness for average case predictions and to enrich limited datasets. So far no method provided sampling of a realistic posterior distribution to improve stability, e.g. for the segmentation of imbalanced objects within images. Therefore, we propose a new approach, based on diffusion models, which can enrich an imbalanced dataset with plausible examples from underrepresented groups by conditioning on segmentation maps. Our method can simply expand limited clinical datasets making them suitable to train machine learning pipelines, and provides an interpretable and human-controllable way of generating histopathology images that are indistinguishable from real ones to human experts. We validate our findings on two datasets, one from the public domain and one from a Kidney Transplant study.
Pay Attention: Accuracy Versus Interpretability Trade-off in Fine-tuned Diffusion Models
Mar 31, 2023



Abstract:The recent progress of diffusion models in terms of image quality has led to a major shift in research related to generative models. Current approaches often fine-tune pre-trained foundation models using domain-specific text-to-image pairs. This approach is straightforward for X-ray image generation due to the high availability of radiology reports linked to specific images. However, current approaches hardly ever look at attention layers to verify whether the models understand what they are generating. In this paper, we discover an important trade-off between image fidelity and interpretability in generative diffusion models. In particular, we show that fine-tuning text-to-image models with learnable text encoder leads to a lack of interpretability of diffusion models. Finally, we demonstrate the interpretability of diffusion models by showing that keeping the language encoder frozen, enables diffusion models to achieve state-of-the-art phrase grounding performance on certain diseases for a challenging multi-label segmentation task, without any additional training. Code and models will be available at https://github.com/MischaD/chest-distillation.
Feature-Conditioned Cascaded Video Diffusion Models for Precise Echocardiogram Synthesis
Mar 23, 2023Abstract:Image synthesis is expected to provide value for the translation of machine learning methods into clinical practice. Fundamental problems like model robustness, domain transfer, causal modelling, and operator training become approachable through synthetic data. Especially, heavily operator-dependant modalities like Ultrasound imaging require robust frameworks for image and video generation. So far, video generation has only been possible by providing input data that is as rich as the output data, e.g., image sequence plus conditioning in, video out. However, clinical documentation is usually scarce and only single images are reported and stored, thus retrospective patient-specific analysis or the generation of rich training data becomes impossible with current approaches. In this paper, we extend elucidated diffusion models for video modelling to generate plausible video sequences from single images and arbitrary conditioning with clinical parameters. We explore this idea within the context of echocardiograms by looking into the variation of the Left Ventricle Ejection Fraction, the most essential clinical metric gained from these examinations. We use the publicly available EchoNet-Dynamic dataset for all our experiments. Our image to sequence approach achieves an $R^2$ score of 93%, which is 38 points higher than recently proposed sequence to sequence generation methods. Code and models will be available at: https://github.com/HReynaud/EchoDiffusion.
Confidence-Aware and Self-Supervised Image Anomaly Localisation
Mar 23, 2023Abstract:Universal anomaly detection still remains a challenging problem in machine learning and medical image analysis. It is possible to learn an expected distribution from a single class of normative samples, e.g., through epistemic uncertainty estimates, auto-encoding models, or from synthetic anomalies in a self-supervised way. The performance of self-supervised anomaly detection approaches is still inferior compared to methods that use examples from known unknown classes to shape the decision boundary. However, outlier exposure methods often do not identify unknown unknowns. Here we discuss an improved self-supervised single-class training strategy that supports the approximation of probabilistic inference with loosen feature locality constraints. We show that up-scaling of gradients with histogram-equalised images is beneficial for recently proposed self-supervision tasks. Our method is integrated into several out-of-distribution (OOD) detection models and we show evidence that our method outperforms the state-of-the-art on various benchmark datasets. Source code will be publicly available by the time of the conference.
Understanding metric-related pitfalls in image analysis validation
Feb 09, 2023Abstract:Validation metrics are key for the reliable tracking of scientific progress and for bridging the current chasm between artificial intelligence (AI) research and its translation into practice. However, increasing evidence shows that particularly in image analysis, metrics are often chosen inadequately in relation to the underlying research problem. This could be attributed to a lack of accessibility of metric-related knowledge: While taking into account the individual strengths, weaknesses, and limitations of validation metrics is a critical prerequisite to making educated choices, the relevant knowledge is currently scattered and poorly accessible to individual researchers. Based on a multi-stage Delphi process conducted by a multidisciplinary expert consortium as well as extensive community feedback, the present work provides the first reliable and comprehensive common point of access to information on pitfalls related to validation metrics in image analysis. Focusing on biomedical image analysis but with the potential of transfer to other fields, the addressed pitfalls generalize across application domains and are categorized according to a newly created, domain-agnostic taxonomy. To facilitate comprehension, illustrations and specific examples accompany each pitfall. As a structured body of information accessible to researchers of all levels of expertise, this work enhances global comprehension of a key topic in image analysis validation.
Zero-Shot Object Segmentation through Concept Distillation from Generative Image Foundation Models
Dec 29, 2022



Abstract:Curating datasets for object segmentation is a difficult task. With the advent of large-scale pre-trained generative models, conditional image generation has been given a significant boost in result quality and ease of use. In this paper, we present a novel method that enables the generation of general foreground-background segmentation models from simple textual descriptions, without requiring segmentation labels. We leverage and explore pre-trained latent diffusion models, to automatically generate weak segmentation masks for concepts and objects. The masks are then used to fine-tune the diffusion model on an inpainting task, which enables fine-grained removal of the object, while at the same time providing a synthetic foreground and background dataset. We demonstrate that using this method beats previous methods in both discriminative and generative performance and closes the gap with fully supervised training while requiring no pixel-wise object labels. We show results on the task of segmenting four different objects (humans, dogs, cars, birds).
ParaDime: A Framework for Parametric Dimensionality Reduction
Oct 10, 2022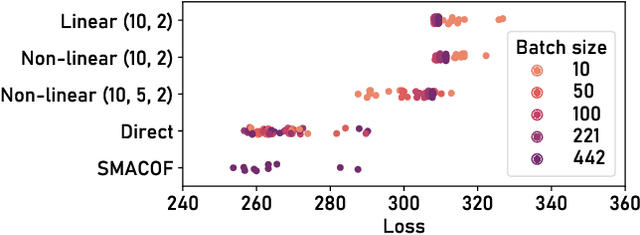
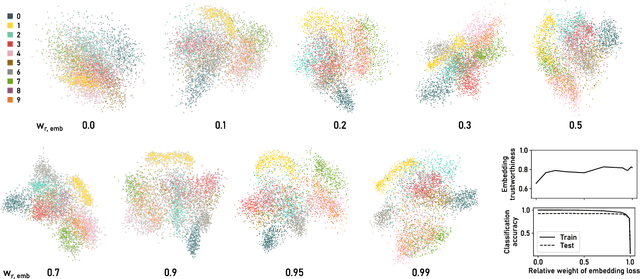
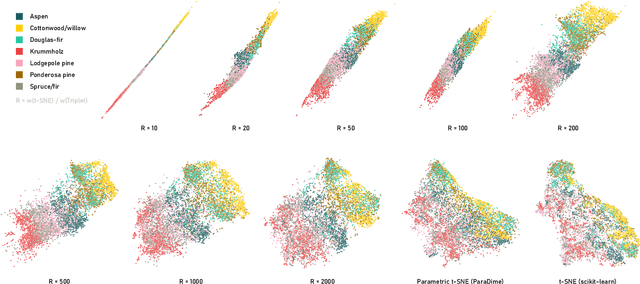
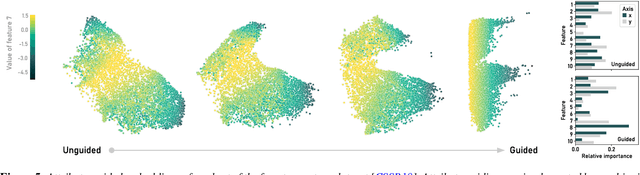
Abstract:ParaDime is a framework for parametric dimensionality reduction (DR). In parametric DR, neural networks are trained to embed high-dimensional data items in a low-dimensional space while minimizing an objective function. ParaDime builds on the idea that the objective functions of several modern DR techniques result from transformed inter-item relationships. It provides a common interface to specify the way these relations and transformations are computed and how they are used within the losses that govern the training process. Through this interface, ParaDime unifies parametric versions of DR techniques such as metric MDS, t-SNE, and UMAP. Furthermore, it allows users to fully customize each aspect of the DR process. We show how this ease of customization makes ParaDime suitable for experimenting with interesting techniques, such as hybrid classification/embedding models or supervised DR, which opens up new possibilities for visualizing high-dimensional data.
Self-Supervised 3D Human Pose Estimation in Static Video Via Neural Rendering
Oct 10, 2022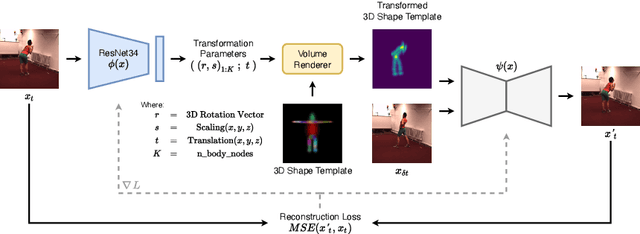
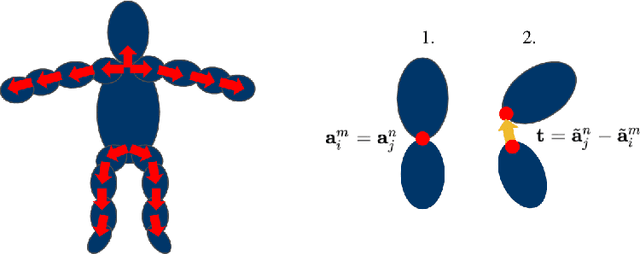
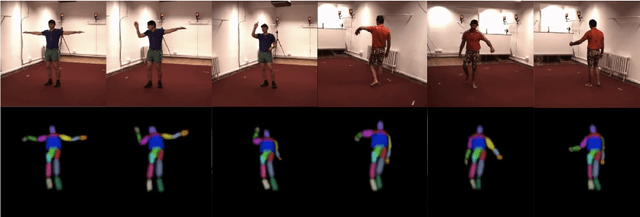
Abstract:Inferring 3D human pose from 2D images is a challenging and long-standing problem in the field of computer vision with many applications including motion capture, virtual reality, surveillance or gait analysis for sports and medicine. We present preliminary results for a method to estimate 3D pose from 2D video containing a single person and a static background without the need for any manual landmark annotations. We achieve this by formulating a simple yet effective self-supervision task: our model is required to reconstruct a random frame of a video given a frame from another timepoint and a rendered image of a transformed human shape template. Crucially for optimisation, our ray casting based rendering pipeline is fully differentiable, enabling end to end training solely based on the reconstruction task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge