"Time": models, code, and papers
Towards unraveling calibration biases in medical image analysis
May 09, 2023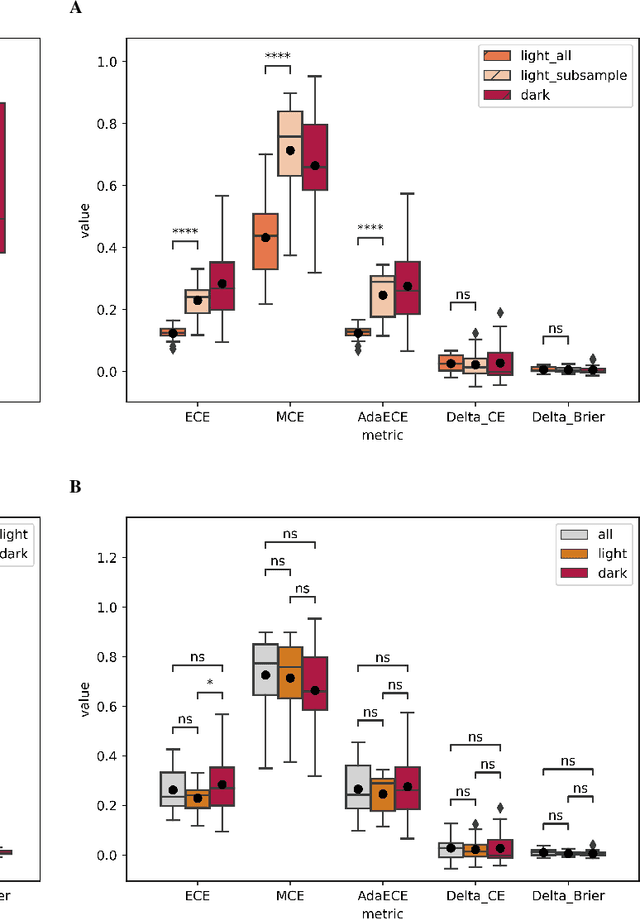
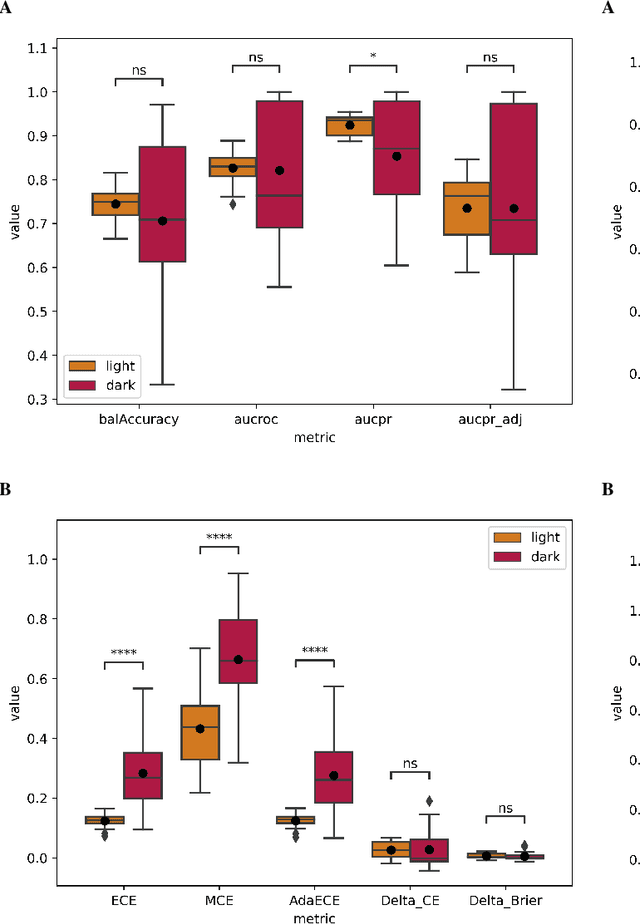
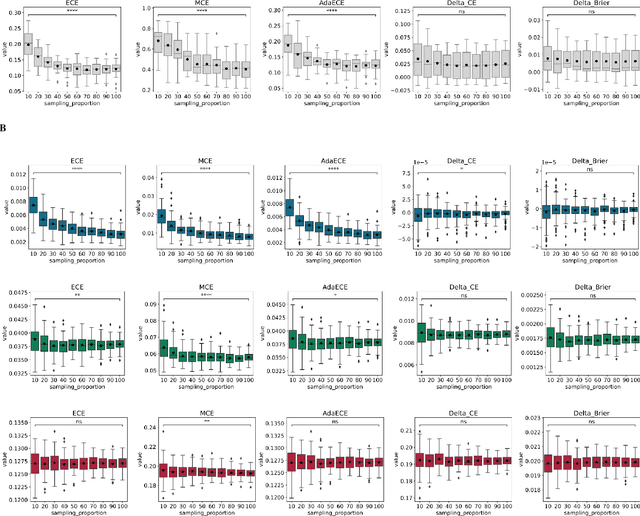
In recent years the development of artificial intelligence (AI) systems for automated medical image analysis has gained enormous momentum. At the same time, a large body of work has shown that AI systems can systematically and unfairly discriminate against certain populations in various application scenarios. These two facts have motivated the emergence of algorithmic fairness studies in this field. Most research on healthcare algorithmic fairness to date has focused on the assessment of biases in terms of classical discrimination metrics such as AUC and accuracy. Potential biases in terms of model calibration, however, have only recently begun to be evaluated. This is especially important when working with clinical decision support systems, as predictive uncertainty is key for health professionals to optimally evaluate and combine multiple sources of information. In this work we study discrimination and calibration biases in models trained for automatic detection of malignant dermatological conditions from skin lesions images. Importantly, we show how several typically employed calibration metrics are systematically biased with respect to sample sizes, and how this can lead to erroneous fairness analysis if not taken into consideration. This is of particular relevance to fairness studies, where data imbalance results in drastic sample size differences between demographic sub-groups, which, if not taken into account, can act as confounders.
T-SciQ: Teaching Multimodal Chain-of-Thought Reasoning via Large Language Model Signals for Science Question Answering
May 09, 2023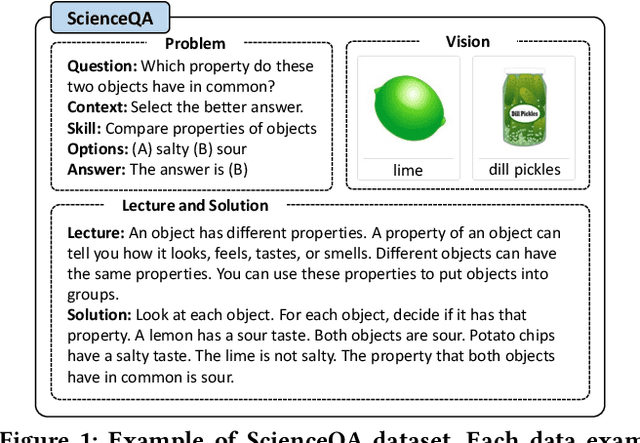
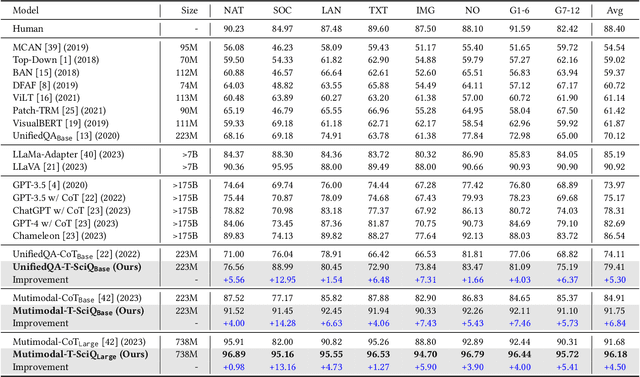
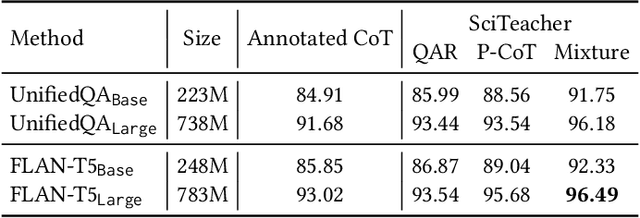
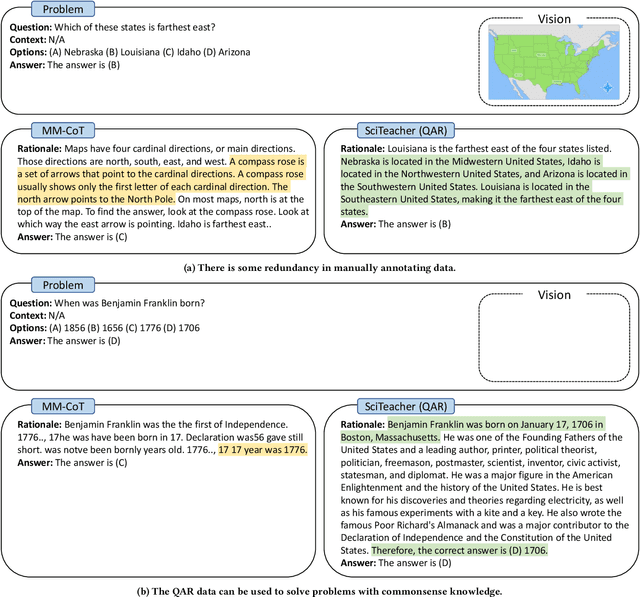
Large Language Models (LLMs) have recently demonstrated exceptional performance in various Natural Language Processing (NLP) tasks. They have also shown the ability to perform chain-of-thought (CoT) reasoning to solve complex problems. Recent studies have explored CoT reasoning in complex multimodal scenarios, such as the science question answering task, by fine-tuning multimodal models with high-quality human-annotated CoT rationales. However, collecting high-quality COT rationales is usually time-consuming and costly. Besides, the annotated rationales are hardly accurate due to the redundant information involved or the essential information missed. To address these issues, we propose a novel method termed \emph{T-SciQ} that aims at teaching science question answering with LLM signals. The T-SciQ approach generates high-quality CoT rationales as teaching signals and is advanced to train much smaller models to perform CoT reasoning in complex modalities. Additionally, we introduce a novel data mixing strategy to produce more effective teaching data samples for simple and complex science question answer problems. Extensive experimental results show that our T-SciQ method achieves a new state-of-the-art performance on the ScienceQA benchmark, with an accuracy of 96.18%. Moreover, our approach outperforms the most powerful fine-tuned baseline by 4.5%.
Neuralizer: General Neuroimage Analysis without Re-Training
May 09, 2023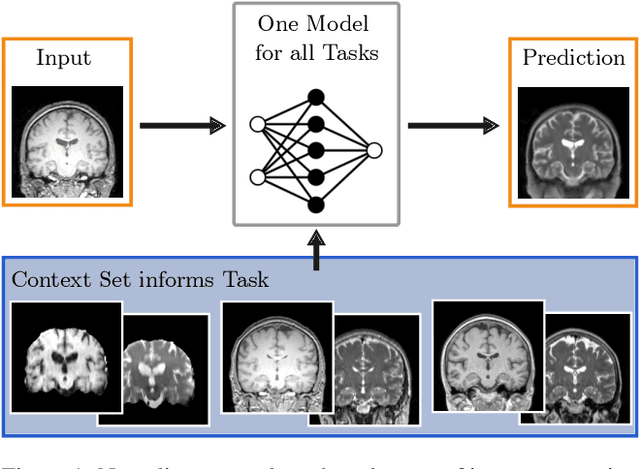
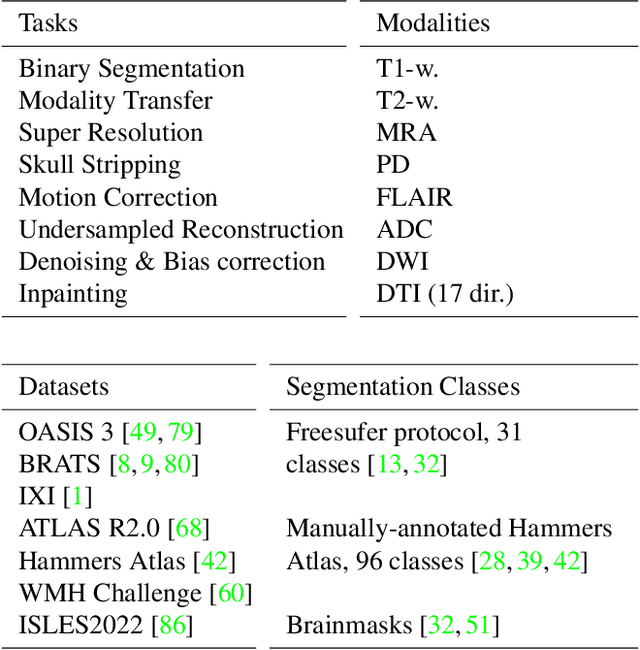
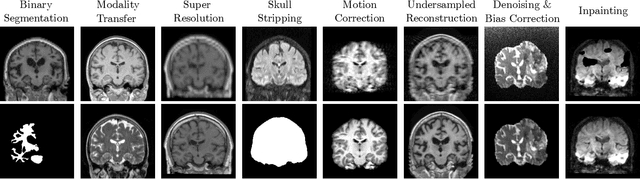

Neuroimage processing tasks like segmentation, reconstruction, and registration are central to the study of neuroscience. Robust deep learning strategies and architectures used to solve these tasks are often similar. Yet, when presented with a new task or a dataset with different visual characteristics, practitioners most often need to train a new model, or fine-tune an existing one. This is a time-consuming process that poses a substantial barrier for the thousands of neuroscientists and clinical researchers who often lack the resources or machine-learning expertise to train deep learning models. In practice, this leads to a lack of adoption of deep learning, and neuroscience tools being dominated by classical frameworks. We introduce Neuralizer, a single model that generalizes to previously unseen neuroimaging tasks and modalities without the need for re-training or fine-tuning. Tasks do not have to be known a priori, and generalization happens in a single forward pass during inference. The model can solve processing tasks across multiple image modalities, acquisition methods, and datasets, and generalize to tasks and modalities it has not been trained on. Our experiments on coronal slices show that when few annotated subjects are available, our multi-task network outperforms task-specific baselines without training on the task.
Optimizing Privacy, Utility and Efficiency in Constrained Multi-Objective Federated Learning
May 09, 2023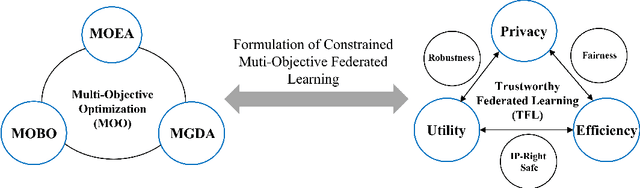
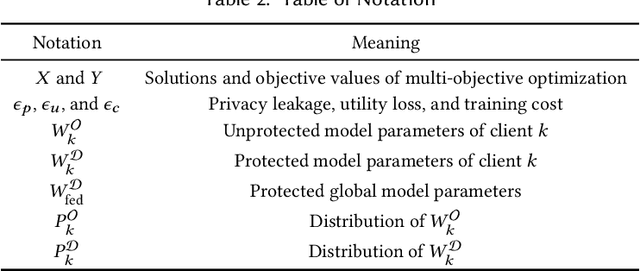


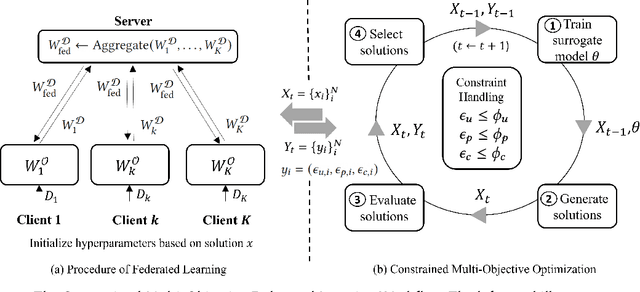
Conventionally, federated learning aims to optimize a single objective, typically the utility. However, for a federated learning system to be trustworthy, it needs to simultaneously satisfy multiple/many objectives, such as maximizing model performance, minimizing privacy leakage and training cost, and being robust to malicious attacks. Multi-Objective Optimization (MOO) aiming to optimize multiple conflicting objectives at the same time is quite suitable for solving the optimization problem of Trustworthy Federated Learning (TFL). In this paper, we unify MOO and TFL by formulating the problem of constrained multi-objective federated learning (CMOFL). Under this formulation, existing MOO algorithms can be adapted to TFL straightforwardly. Different from existing CMOFL works focusing on utility, efficiency, fairness, and robustness, we consider optimizing privacy leakage along with utility loss and training cost, the three primary objectives of a TFL system. We develop two improved CMOFL algorithms based on NSGA-II and PSL, respectively, for effectively and efficiently finding Pareto optimal solutions, and we provide theoretical analysis on their convergence. We design specific measurements of privacy leakage, utility loss, and training cost for three privacy protection mechanisms: Randomization, BatchCrypt (An efficient version of homomorphic encryption), and Sparsification. Empirical experiments conducted under each of the three protection mechanisms demonstrate the effectiveness of our proposed algorithms.
RATs-NAS: Redirection of Adjacent Trails on GCN for Neural Architecture Search
May 09, 2023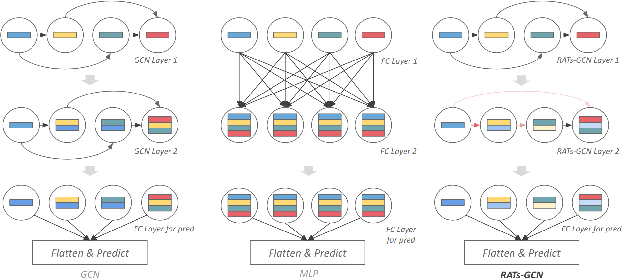
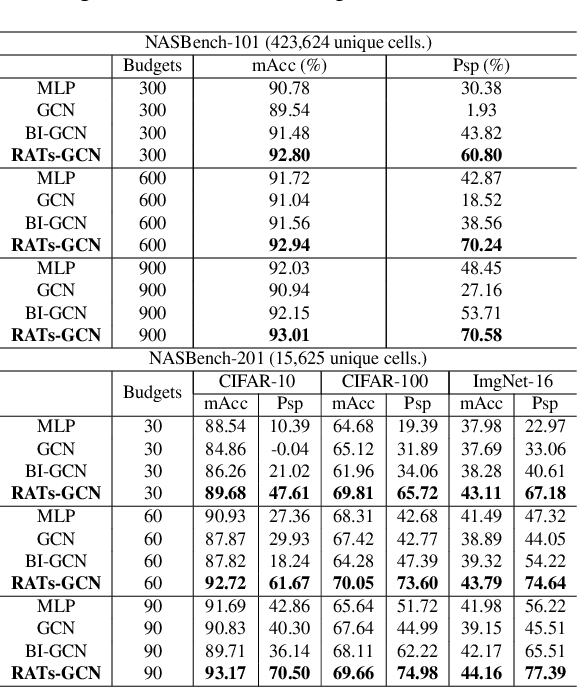
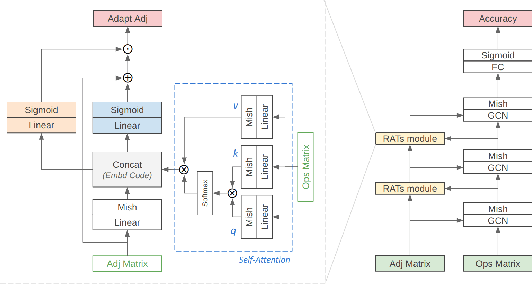
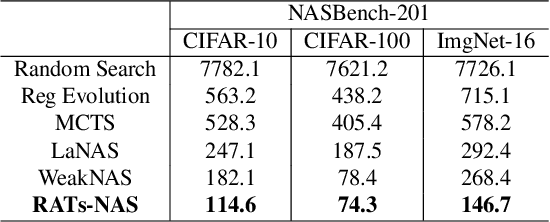
Various hand-designed CNN architectures have been developed, such as VGG, ResNet, DenseNet, etc., and achieve State-of-the-Art (SoTA) levels on different tasks. Neural Architecture Search (NAS) now focuses on automatically finding the best CNN architecture to handle the above tasks. However, the verification of a searched architecture is very time-consuming and makes predictor-based methods become an essential and important branch of NAS. Two commonly used techniques to build predictors are graph-convolution networks (GCN) and multilayer perceptron (MLP). In this paper, we consider the difference between GCN and MLP on adjacent operation trails and then propose the Redirected Adjacent Trails NAS (RATs-NAS) to quickly search for the desired neural network architecture. The RATs-NAS consists of two components: the Redirected Adjacent Trails GCN (RATs-GCN) and the Predictor-based Search Space Sampling (P3S) module. RATs-GCN can change trails and their strengths to search for a better neural network architecture. P3S can rapidly focus on tighter intervals of FLOPs in the search space. Based on our observations on cell-based NAS, we believe that architectures with similar FLOPs will perform similarly. Finally, the RATs-NAS consisting of RATs-GCN and P3S beats WeakNAS, Arch-Graph, and others by a significant margin on three sub-datasets of NASBench-201.
DifFIQA: Face Image Quality Assessment Using Denoising Diffusion Probabilistic Models
May 09, 2023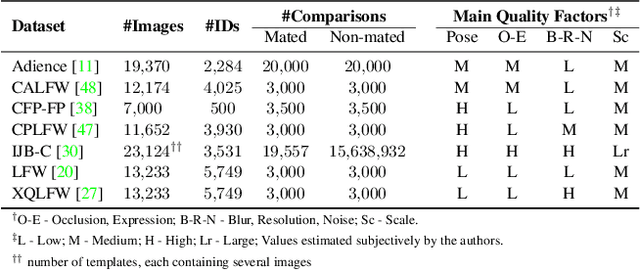
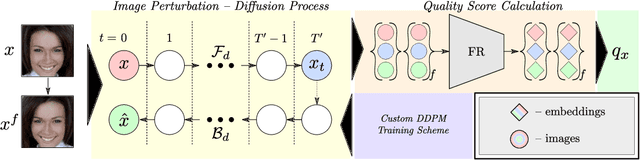
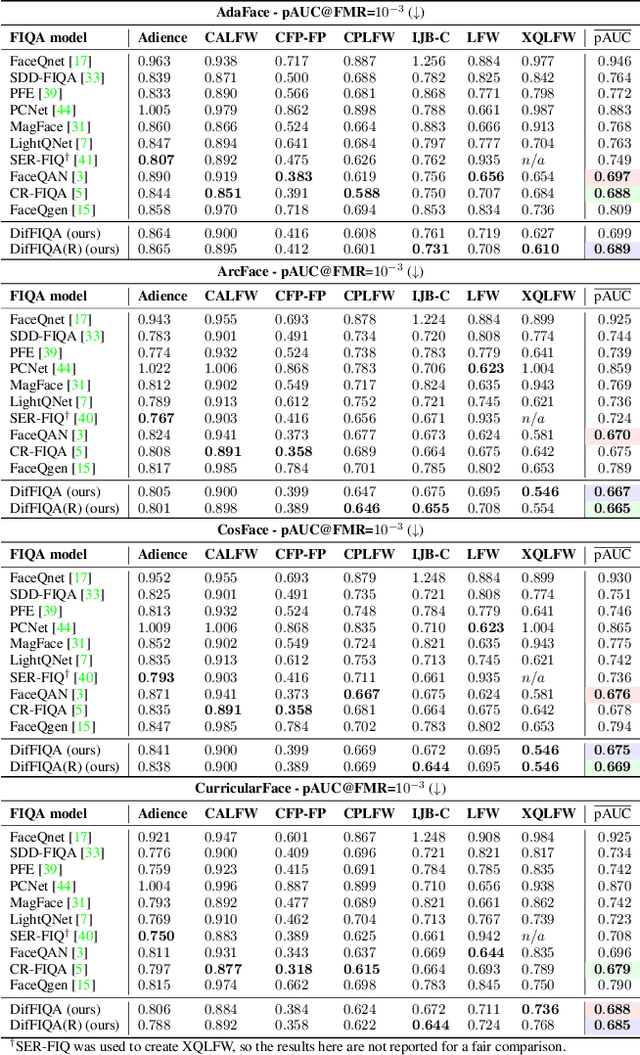
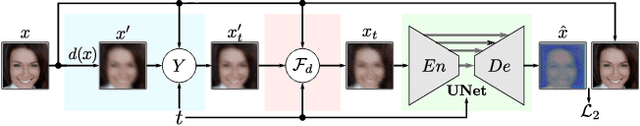
Modern face recognition (FR) models excel in constrained scenarios, but often suffer from decreased performance when deployed in unconstrained (real-world) environments due to uncertainties surrounding the quality of the captured facial data. Face image quality assessment (FIQA) techniques aim to mitigate these performance degradations by providing FR models with sample-quality predictions that can be used to reject low-quality samples and reduce false match errors. However, despite steady improvements, ensuring reliable quality estimates across facial images with diverse characteristics remains challenging. In this paper, we present a powerful new FIQA approach, named DifFIQA, which relies on denoising diffusion probabilistic models (DDPM) and ensures highly competitive results. The main idea behind the approach is to utilize the forward and backward processes of DDPMs to perturb facial images and quantify the impact of these perturbations on the corresponding image embeddings for quality prediction. Because the diffusion-based perturbations are computationally expensive, we also distill the knowledge encoded in DifFIQA into a regression-based quality predictor, called DifFIQA(R), that balances performance and execution time. We evaluate both models in comprehensive experiments on 7 datasets, with 4 target FR models and against 10 state-of-the-art FIQA techniques with highly encouraging results. The source code will be made publicly available.
Structured Sentiment Analysis as Transition-based Dependency Parsing
May 09, 2023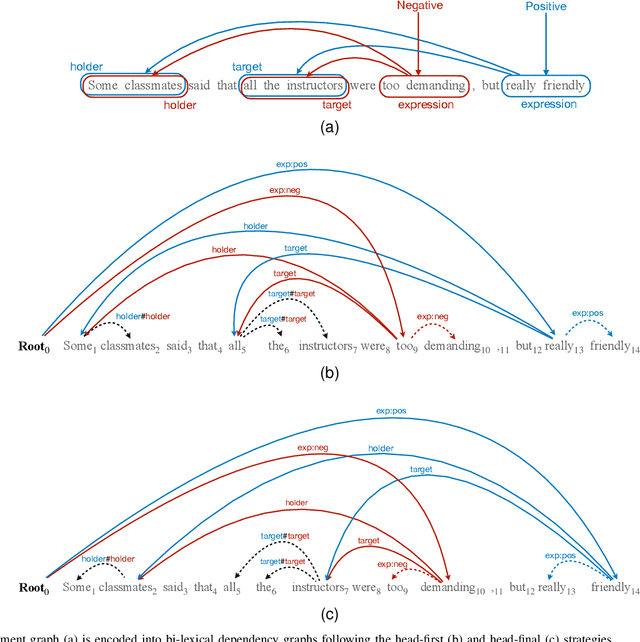

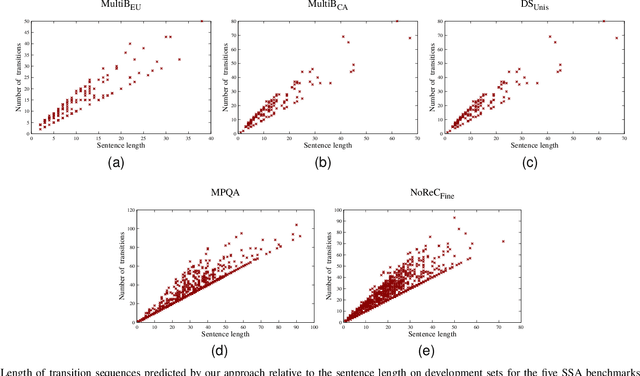
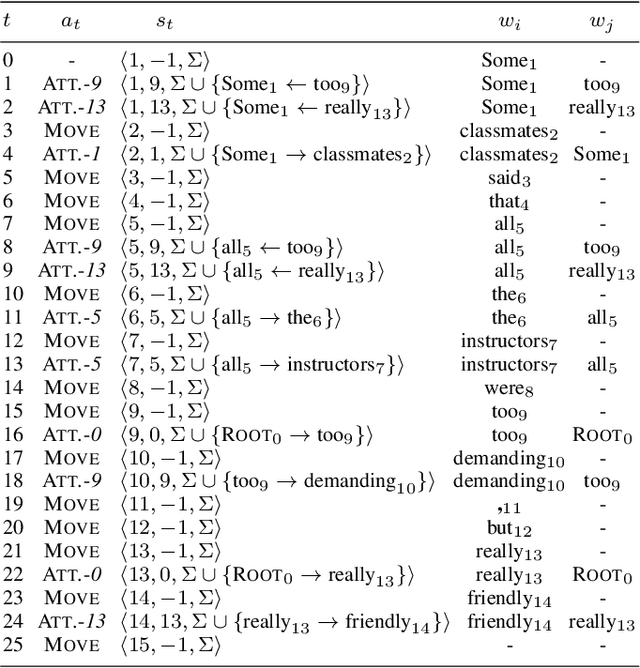
Structured sentiment analysis (SSA) aims to automatically extract people's opinions from a text in natural language and adequately represent that information in a graph structure. One of the most accurate methods for performing SSA was recently proposed and consists of approaching it as a dependency parsing task. Although we can find in the literature how transition-based algorithms excel in dependency parsing in terms of accuracy and efficiency, all proposed attempts to tackle SSA following that approach were based on graph-based models. In this article, we present the first transition-based method to address SSA as dependency parsing. Specifically, we design a transition system that processes the input text in a left-to-right pass, incrementally generating the graph structure containing all identified opinions. To effectively implement our final transition-based model, we resort to a Pointer Network architecture as a backbone. From an extensive evaluation, we demonstrate that our model offers the best performance to date in practically all cases among prior dependency-based methods, and surpass recent task-specific techniques on the most challenging datasets. We additionally include an in-depth analysis and empirically prove that the overall time-complexity cost of our approach is quadratic in the sentence length, being more efficient than top-performing graph-based parsers.
Dynamic Sparse Training with Structured Sparsity
May 03, 2023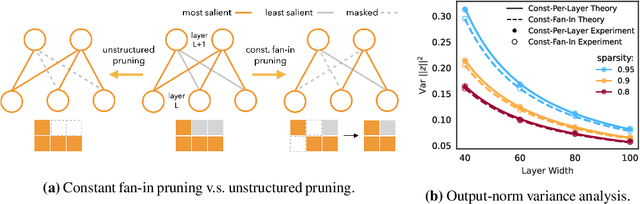
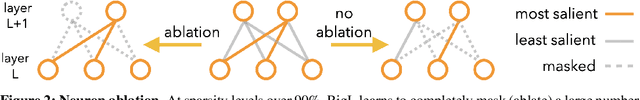
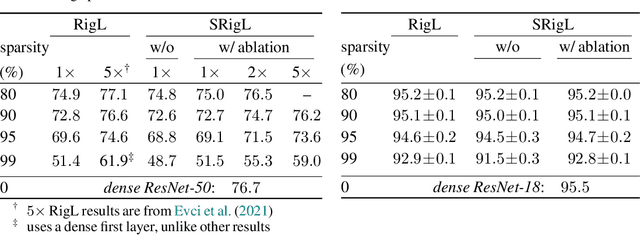
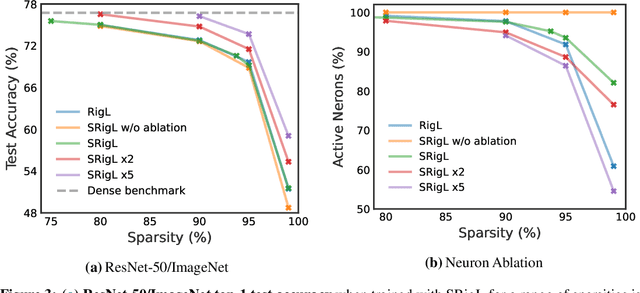
DST methods achieve state-of-the-art results in sparse neural network training, matching the generalization of dense models while enabling sparse training and inference. Although the resulting models are highly sparse and theoretically cheaper to train, achieving speedups with unstructured sparsity on real-world hardware is challenging. In this work we propose a DST method to learn a variant of structured N:M sparsity, the acceleration of which in general is commonly supported in commodity hardware. Furthermore, we motivate with both a theoretical analysis and empirical results, the generalization performance of our specific N:M sparsity (constant fan-in), present a condensed representation with a reduced parameter and memory footprint, and demonstrate reduced inference time compared to dense models with a naive PyTorch CPU implementation of the condensed representation Our source code is available at https://github.com/calgaryml/condensed-sparsity
Exploring the Protein Sequence Space with Global Generative Models
May 03, 2023Recent advancements in specialized large-scale architectures for training image and language have profoundly impacted the field of computer vision and natural language processing (NLP). Language models, such as the recent ChatGPT and GPT4 have demonstrated exceptional capabilities in processing, translating, and generating human languages. These breakthroughs have also been reflected in protein research, leading to the rapid development of numerous new methods in a short time, with unprecedented performance. Language models, in particular, have seen widespread use in protein research, as they have been utilized to embed proteins, generate novel ones, and predict tertiary structures. In this book chapter, we provide an overview of the use of protein generative models, reviewing 1) language models for the design of novel artificial proteins, 2) works that use non-Transformer architectures, and 3) applications in directed evolution approaches.
Cellular EXchange Imaging (CEXI): Evaluation of a diffusion model including water exchange in cells using numerical phantoms of permeable spheres
Apr 12, 2023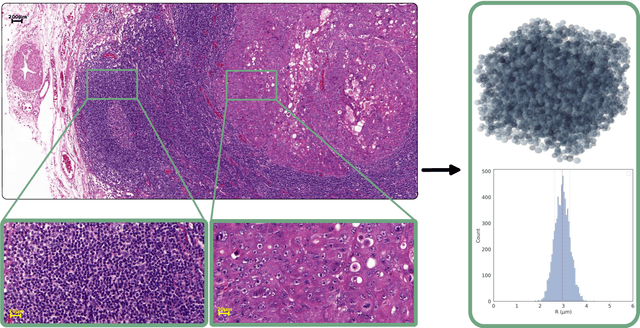

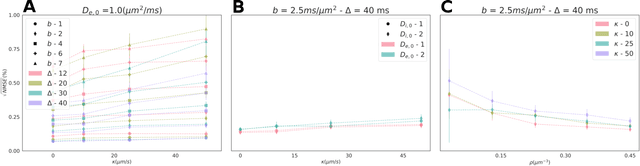
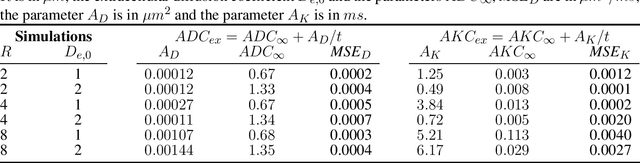
Purpose: Biophysical models of diffusion MRI have been developed to characterize microstructure in various tissues, but existing models are not suitable for tissue composed of permeable spherical cells. In this study we introduce Cellular Exchange Imaging (CEXI), a model tailored for permeable spherical cells, and compares its performance to a related Ball \& Sphere (BS) model that neglects permeability. Methods: We generated DW-MRI signals using Monte-Carlo simulations with a PGSE sequence in numerical substrates made of spherical cells and their extracellular space for a range of membrane permeability. From these signals, the properties of the substrates were inferred using both BS and CEXI models. Results: CEXI outperformed the impermeable model by providing more stable estimates cell size and intracellular volume fraction that were diffusion time-independent. Notably, CEXI accurately estimated the exchange time for low to moderate permeability levels previously reported in other studies ($\kappa<25\mu m/s$). However, in highly permeable substrates ($\kappa=50\mu m/s$), the estimated parameters were less stable, particularly the diffusion coefficients. Conclusion: This study highlights the importance of modeling the exchange time to accurately quantify microstructure properties in permeable cellular substrates. Future studies should evaluate CEXI in clinical applications such as lymph nodes, investigate exchange time as a potential biomarker of tumor severity, and develop more appropriate tissue models that account for anisotropic diffusion and highly permeable membranes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge