"Information": models, code, and papers
Choosing the Number of Topics in LDA Models -- A Monte Carlo Comparison of Selection Criteria
Dec 28, 2022
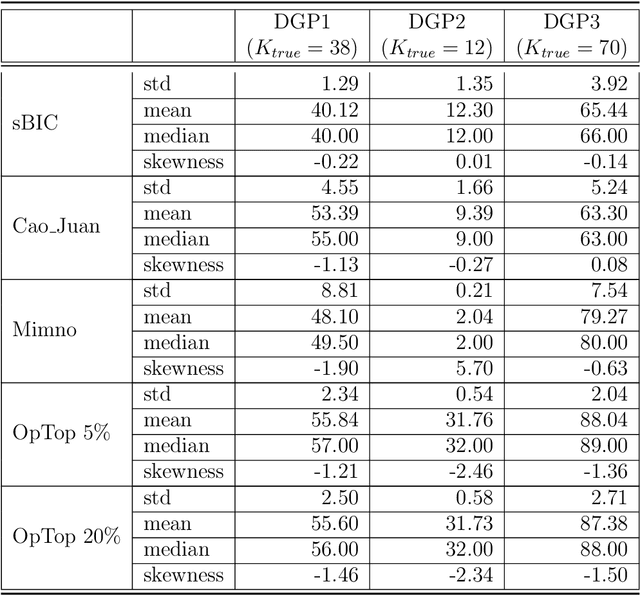
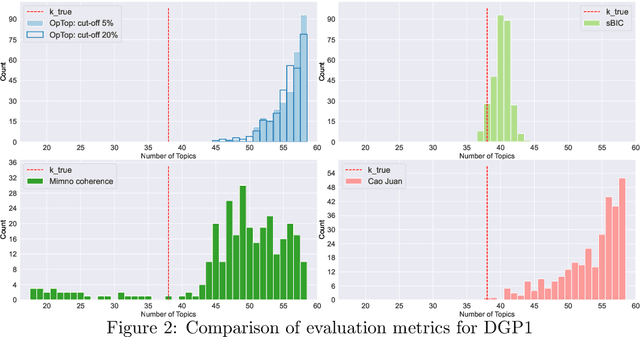
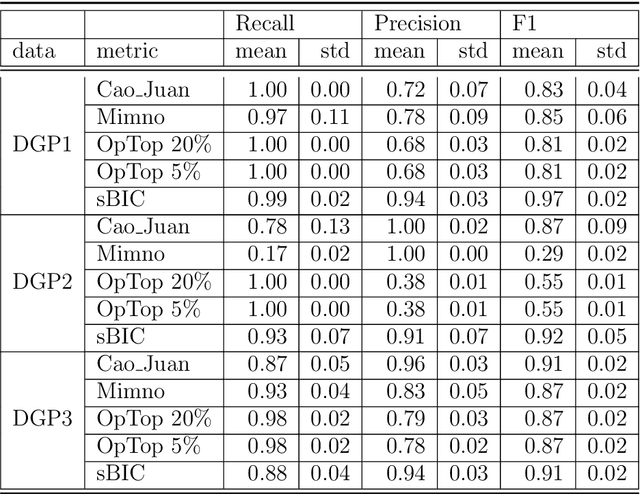
Selecting the number of topics in LDA models is considered to be a difficult task, for which alternative approaches have been proposed. The performance of the recently developed singular Bayesian information criterion (sBIC) is evaluated and compared to the performance of alternative model selection criteria. The sBIC is a generalization of the standard BIC that can be implemented to singular statistical models. The comparison is based on Monte Carlo simulations and carried out for several alternative settings, varying with respect to the number of topics, the number of documents and the size of documents in the corpora. Performance is measured using different criteria which take into account the correct number of topics, but also whether the relevant topics from the DGPs are identified. Practical recommendations for LDA model selection in applications are derived.
Plumber: A Modular Framework to Create Information Extraction Pipelines
Jun 03, 2022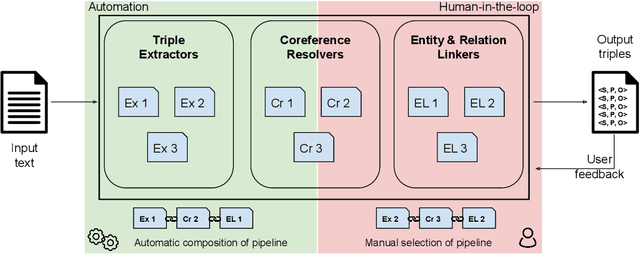
Information Extraction (IE) tasks are commonly studied topics in various domains of research. Hence, the community continuously produces multiple techniques, solutions, and tools to perform such tasks. However, running those tools and integrating them within existing infrastructure requires time, expertise, and resources. One pertinent task here is triples extraction and linking, where structured triples are extracted from a text and aligned to an existing Knowledge Graph (KG). In this paper, we present PLUMBER, the first framework that allows users to manually and automatically create suitable IE pipelines from a community-created pool of tools to perform triple extraction and alignment on unstructured text. Our approach provides an interactive medium to alter the pipelines and perform IE tasks. A short video to show the working of the framework for different use-cases is available online under: https://www.youtube.com/watch?v=XC9rJNIUv8g
TFormer: A throughout fusion transformer for multi-modal skin lesion diagnosis
Nov 21, 2022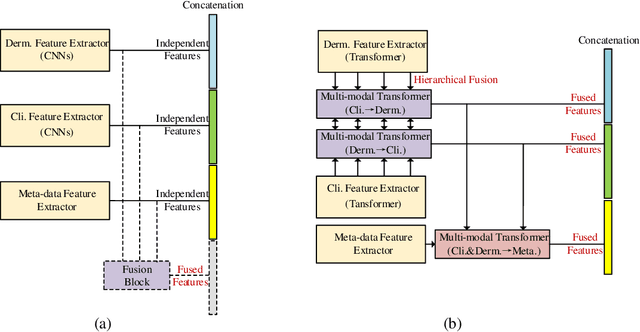
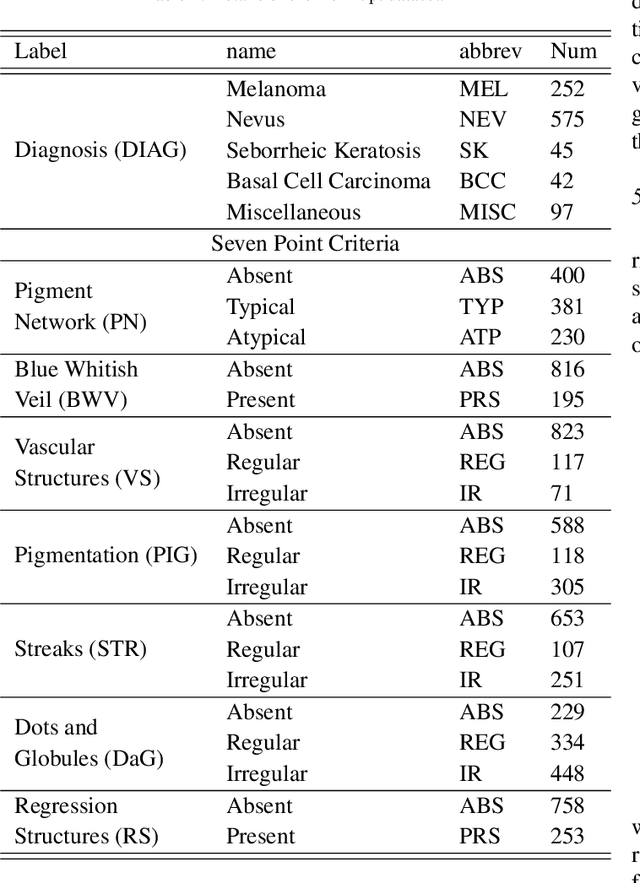
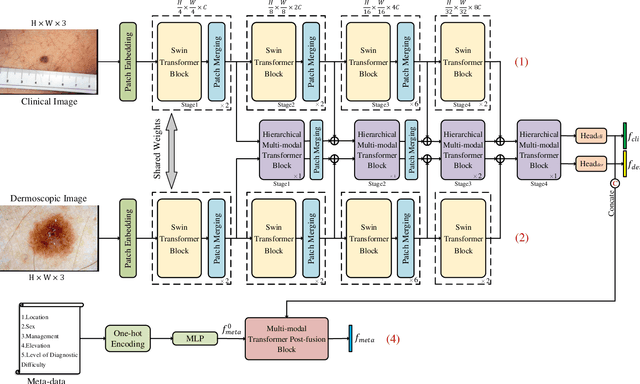

Multi-modal skin lesion diagnosis (MSLD) has achieved remarkable success by modern computer-aided diagnosis technology based on deep convolutions. However, the information aggregation across modalities in MSLD remains challenging due to severity unaligned spatial resolution (dermoscopic image and clinical image) and heterogeneous data (dermoscopic image and patients' meta-data). Limited by the intrinsic local attention, most recent MSLD pipelines using pure convolutions struggle to capture representative features in shallow layers, thus the fusion across different modalities is usually done at the end of the pipelines, even at the last layer, leading to an insufficient information aggregation. To tackle the issue, we introduce a pure transformer-based method, which we refer to as ``Throughout Fusion Transformer (TFormer)", for sufficient information intergration in MSLD. Different from the existing approaches with convolutions, the proposed network leverages transformer as feature extraction backbone, bringing more representative shallow features. We then carefully design a stack of dual-branch hierarchical multi-modal transformer (HMT) blocks to fuse information across different image modalities in a stage-by-stage way. With the aggregated information of image modalities, a multi-modal transformer post-fusion (MTP) block is designed to integrate features across image and non-image data. Such a strategy that information of the image modalities is firstly fused then the heterogeneous ones enables us to better divide and conquer the two major challenges while ensuring inter-modality dynamics are effectively modeled. Experiments conducted on the public Derm7pt dataset validate the superiority of the proposed method. Our TFormer outperforms other state-of-the-art methods. Ablation experiments also suggest the effectiveness of our designs.
GNN-SL: Sequence Labeling Based on Nearest Examples via GNN
Dec 12, 2022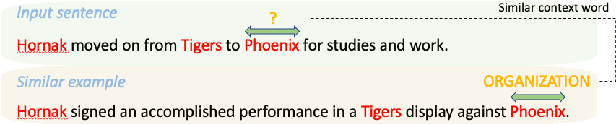
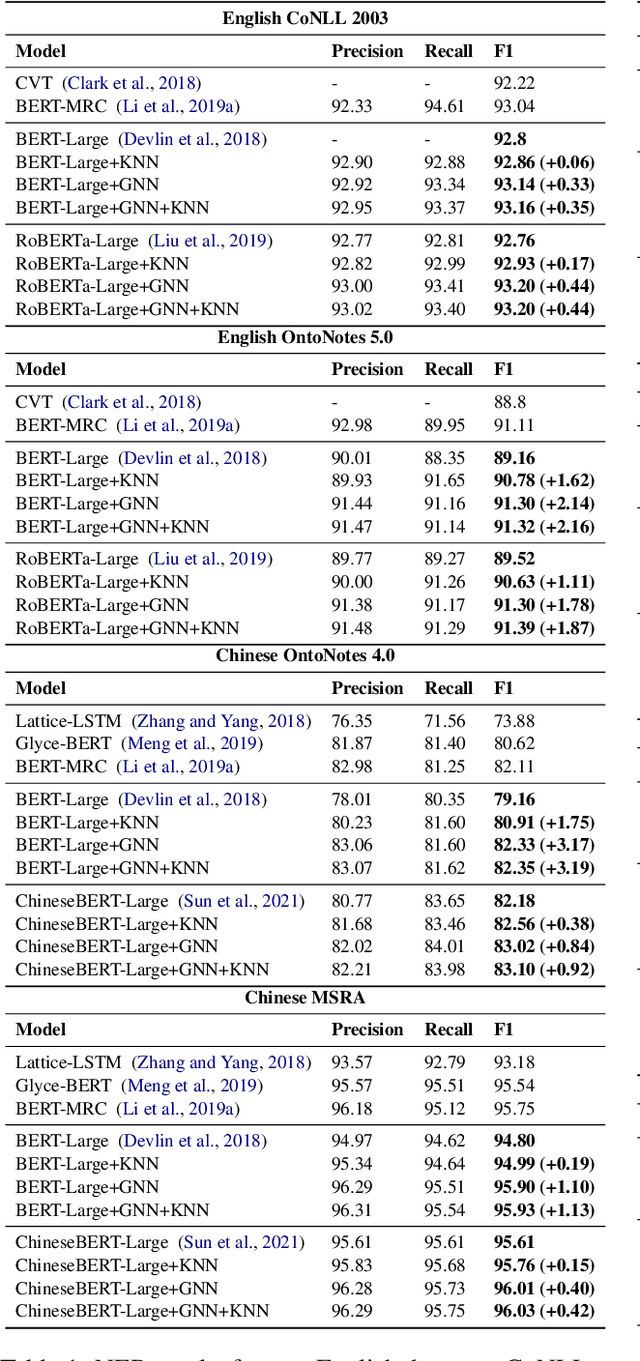
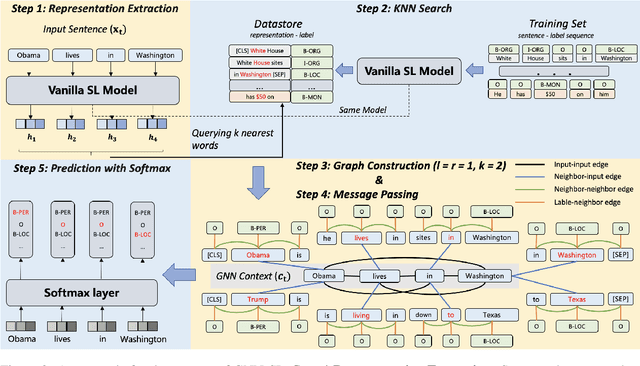
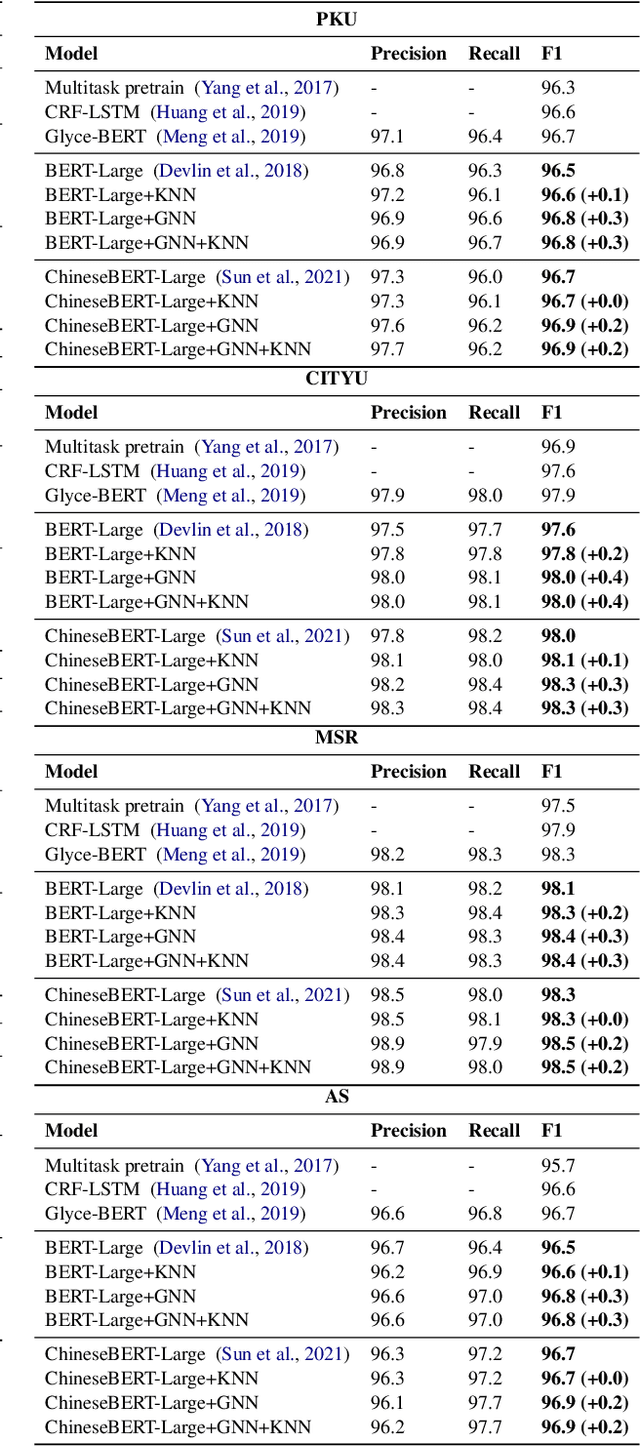
To better handle long-tail cases in the sequence labeling (SL) task, in this work, we introduce graph neural networks sequence labeling (GNN-SL), which augments the vanilla SL model output with similar tagging examples retrieved from the whole training set. Since not all the retrieved tagging examples benefit the model prediction, we construct a heterogeneous graph, and leverage graph neural networks (GNNs) to transfer information between the retrieved tagging examples and the input word sequence. The augmented node which aggregates information from neighbors is used to do prediction. This strategy enables the model to directly acquire similar tagging examples and improves the general quality of predictions. We conduct a variety of experiments on three typical sequence labeling tasks: Named Entity Recognition (NER), Part of Speech Tagging (POS), and Chinese Word Segmentation (CWS) to show the significant performance of our GNN-SL. Notably, GNN-SL achieves SOTA results of 96.9 (+0.2) on PKU, 98.3 (+0.4) on CITYU, 98.5 (+0.2) on MSR, and 96.9 (+0.2) on AS for the CWS task, and results comparable to SOTA performances on NER datasets, and POS datasets.
CbwLoss: Constrained Bidirectional Weighted Loss for Self-supervised Learning of Depth and Pose
Dec 12, 2022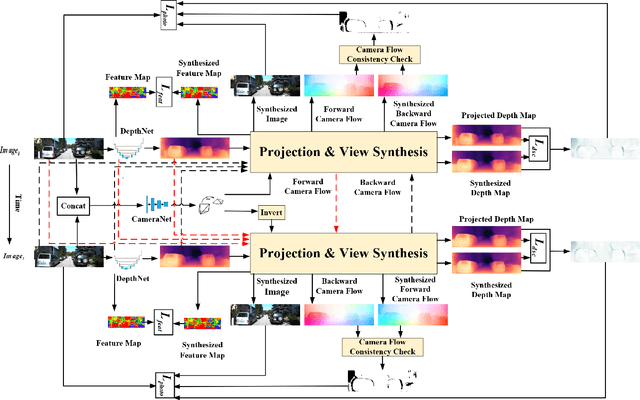
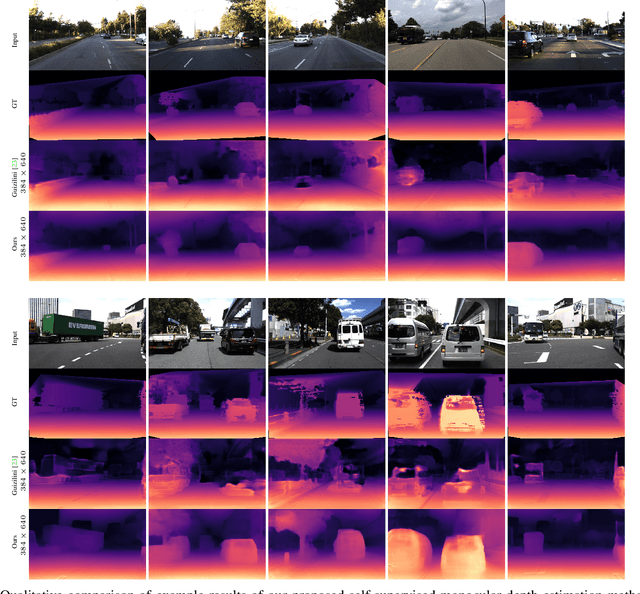
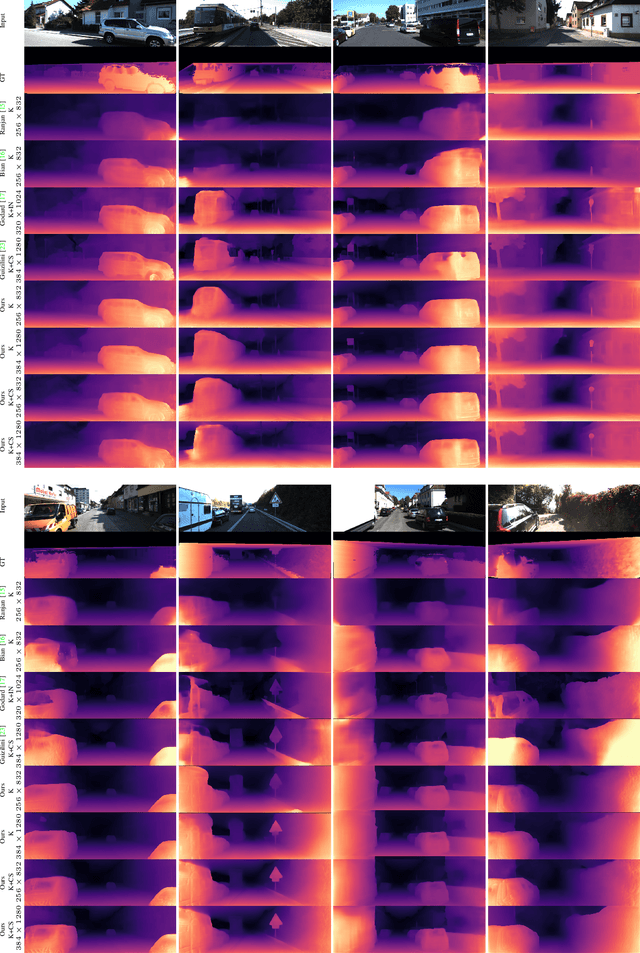
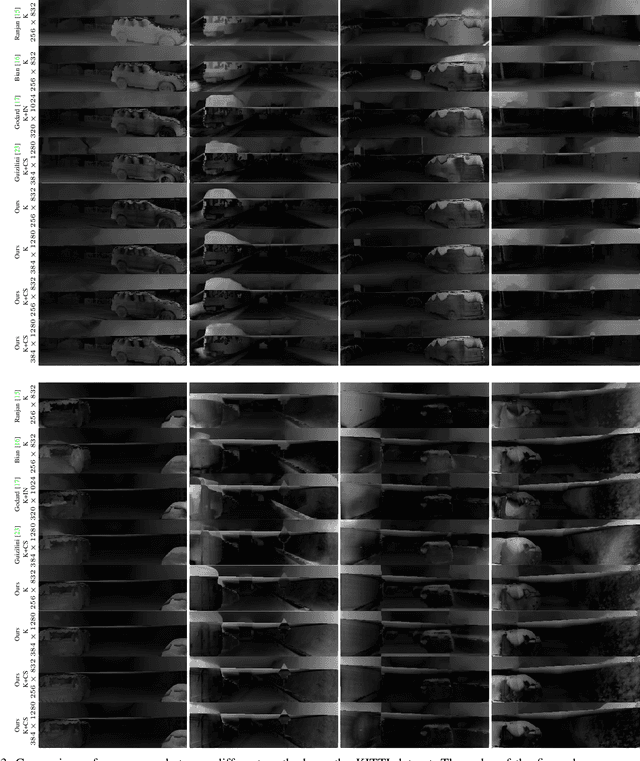

Photometric differences are widely used as supervision signals to train neural networks for estimating depth and camera pose from unlabeled monocular videos. However, this approach is detrimental for model optimization because occlusions and moving objects in a scene violate the underlying static scenario assumption. In addition, pixels in textureless regions or less discriminative pixels hinder model training. To solve these problems, in this paper, we deal with moving objects and occlusions utilizing the difference of the flow fields and depth structure generated by affine transformation and view synthesis, respectively. Secondly, we mitigate the effect of textureless regions on model optimization by measuring differences between features with more semantic and contextual information without adding networks. In addition, although the bidirectionality component is used in each sub-objective function, a pair of images are reasoned about only once, which helps reduce overhead. Extensive experiments and visual analysis demonstrate the effectiveness of the proposed method, which outperform existing state-of-the-art self-supervised methods under the same conditions and without introducing additional auxiliary information.
L-SeqSleepNet: Whole-cycle Long Sequence Modelling for Automatic Sleep Staging
Jan 09, 2023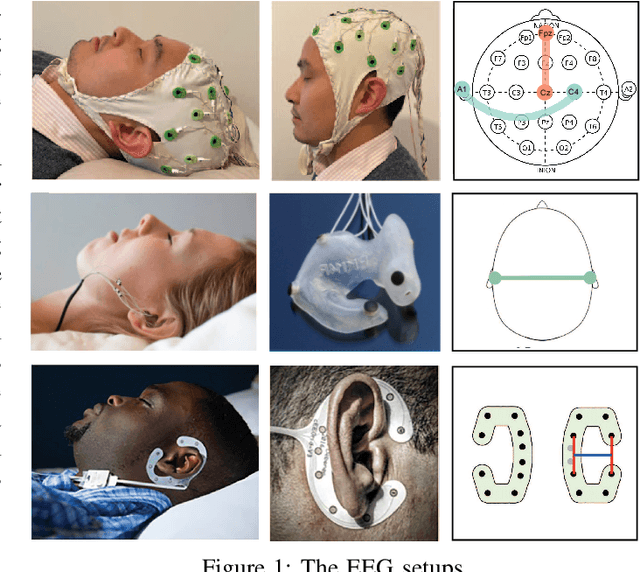
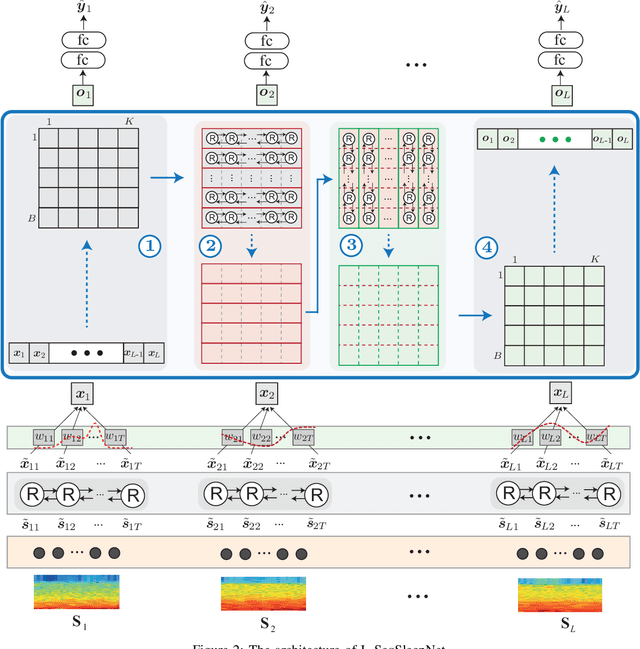
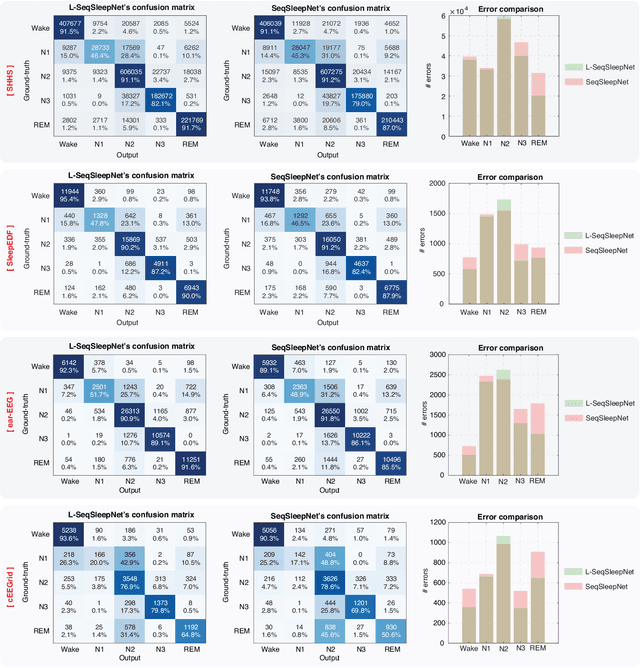
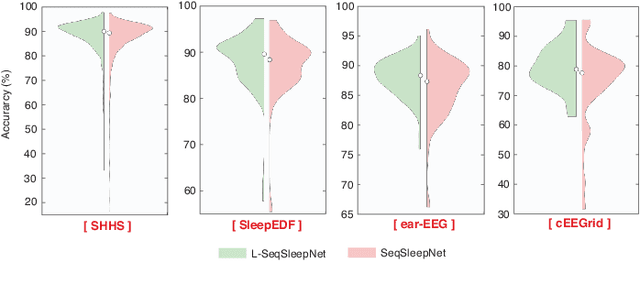
Human sleep is cyclical with a period of approximately 90 minutes, implying long temporal dependency in the sleep data. Yet, exploring this long-term dependency when developing sleep staging models has remained untouched. In this work, we show that while encoding the logic of a whole sleep cycle is crucial to improve sleep staging performance, the sequential modelling approach in existing state-of-the-art deep learning models are inefficient for that purpose. We then introduce a method for efficient long sequence modelling and propose a new deep learning model, L-SeqSleepNet, incorporating this method to take into account whole-cycle sleep information for sleep staging. Evaluating L-SeqSleepNet on a set of four distinct databases of various sizes, we demonstrate state-of-the-art performance obtained by the model over three different EEG setups, including scalp EEG in conventional Polysomnography (PSG), in-ear EEG, and around-the-ear EEG (cEEGrid), even with a single-EEG channel input. Our analyses also show that L-SeqSleepNet is able to remedy the effect of N2 sleep (the major class in terms of classification) to bring down errors in other sleep stages and that the network largely reduces exceptionally high errors seen in many subjects. Finally, the computation time only grows at a sub-linear rate when the sequence length increases.
Distributed Filtering with Value of Information Censoring
Apr 01, 2022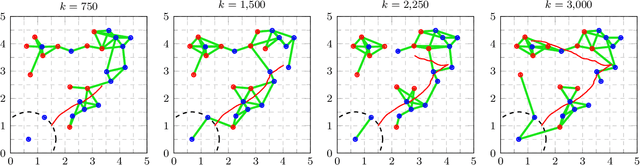
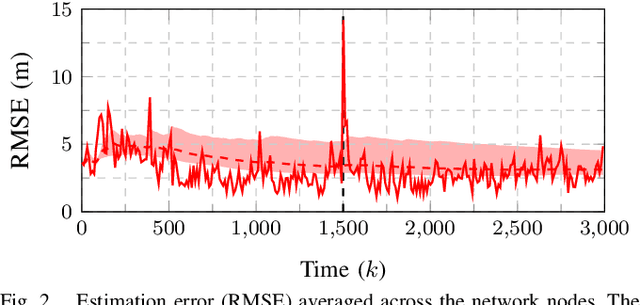
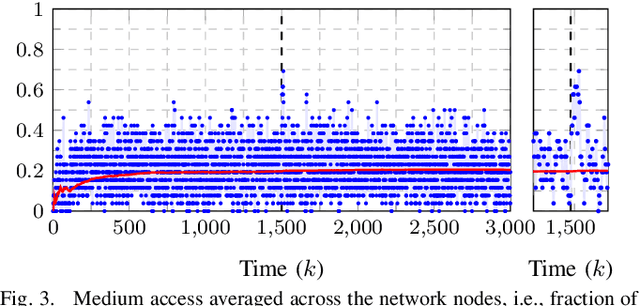
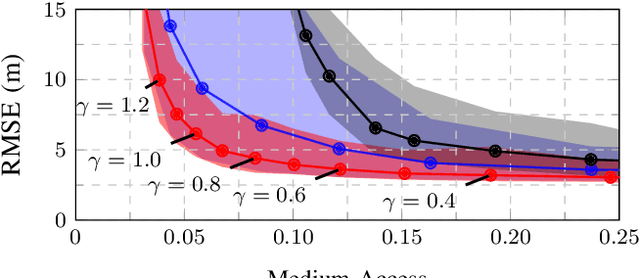
This work presents a distributed estimation algorithm that efficiently uses the available communication resources. The approach is based on Bayesian filtering that is distributed across a network by using the logarithmic opinion pool operator. Communication efficiency is achieved by having only agents with high Value of Information (VoI) share their estimates, and the algorithm provides a tunable trade-off between communication resources and estimation error. Under linear-Gaussian models the algorithm takes the form of a censored distributed Information filter, which guarantees the consistency of agent estimates. Importantly, consistent estimates are shown to play a crucial role in enabling the large reductions in communication usage provided by the VoI censoring approach. We verify the performance of the proposed method via complex simulations in a dynamic network topology and by experimental validation over a real ad-hoc wireless communication network. The results show the validity of using the proposed method to drastically reduce the communication costs of distributed estimation tasks.
JEMMA: An Extensible Java Dataset for ML4Code Applications
Dec 18, 2022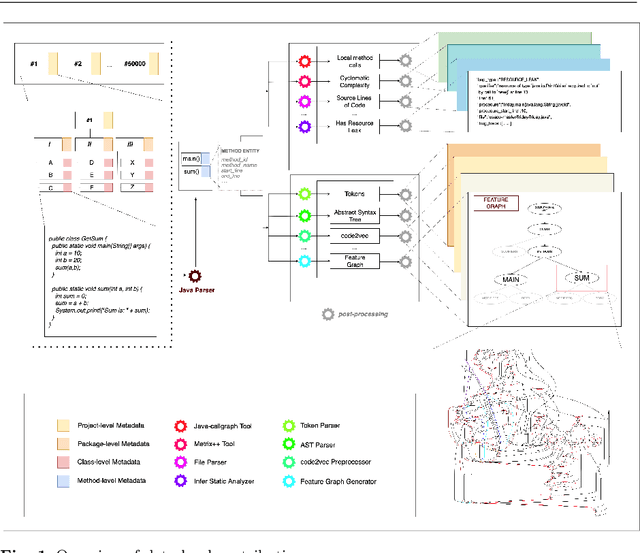
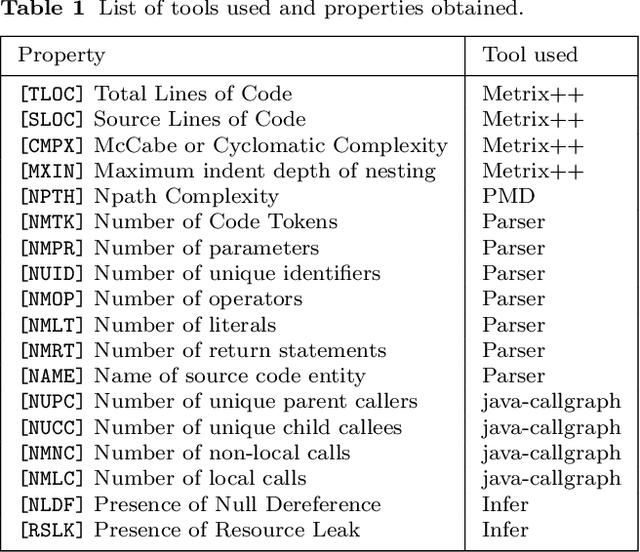

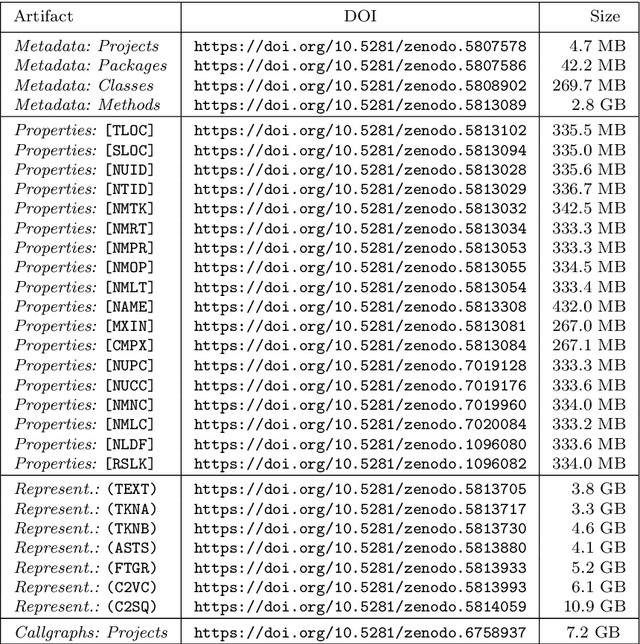
Machine Learning for Source Code (ML4Code) is an active research field in which extensive experimentation is needed to discover how to best use source code's richly structured information. With this in mind, we introduce JEMMA, an Extensible Java Dataset for ML4Code Applications, which is a large-scale, diverse, and high-quality dataset targeted at ML4Code. Our goal with JEMMA is to lower the barrier to entry in ML4Code by providing the building blocks to experiment with source code models and tasks. JEMMA comes with a considerable amount of pre-processed information such as metadata, representations (e.g., code tokens, ASTs, graphs), and several properties (e.g., metrics, static analysis results) for 50,000 Java projects from the 50KC dataset, with over 1.2 million classes and over 8 million methods. JEMMA is also extensible allowing users to add new properties and representations to the dataset, and evaluate tasks on them. Thus, JEMMA becomes a workbench that researchers can use to experiment with novel representations and tasks operating on source code. To demonstrate the utility of the dataset, we also report results from two empirical studies on our data, ultimately showing that significant work lies ahead in the design of context-aware source code models that can reason over a broader network of source code entities in a software project, the very task that JEMMA is designed to help with.
OTFS-SCMA: A Downlink NOMA Scheme for Massive Connectivity in High Mobility Channels
Jan 03, 2023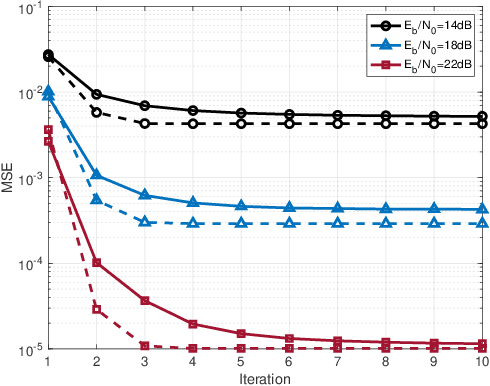
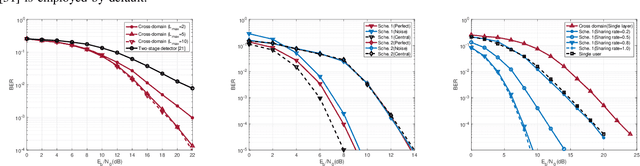
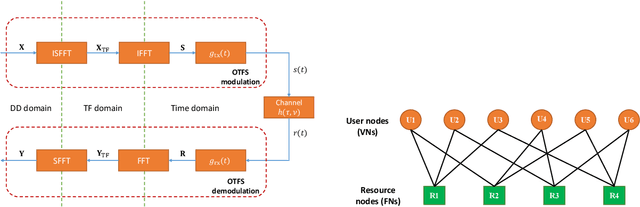
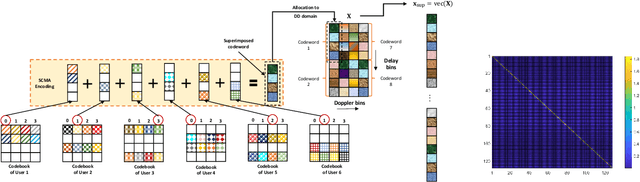
This paper studies a downlink system that combines orthogonal-time-frequency-space (OTFS) modulation and sparse code multiple access (SCMA) to support massive connectivity in high-mobility environments. We propose a cross-domain receiver for the considered OTFS-SCMA system which efficiently carries out OTFS symbol estimation and SCMA decoding in a joint manner. This is done by iteratively passing the extrinsic information between the time domain and the delay-Doppler (DD) domain via the corresponding unitary transformation to ensure the principal orthogonality of errors from each domain. We show that the proposed OTFS-SCMA detection algorithm exists at a fixed point in the state evolution when it converges. To further enhance the error performance of the proposed OTFS-SCMA system, we investigate the cooperation between downlink users to exploit the diversity gains and develop a distributed cooperative detection (DCD) algorithm with the aid of belief consensus. Our numerical results demonstrate the effectiveness and convergence of the proposed algorithm and show an increased spectral efficiency compared to the conventional OTFS transmission.
Generative appearance replay for continual unsupervised domain adaptation
Jan 03, 2023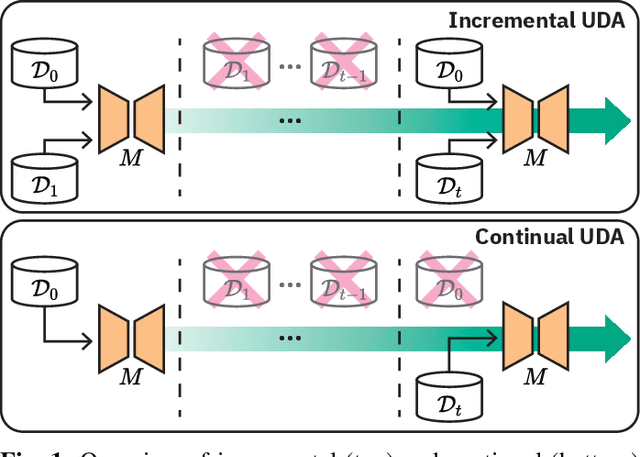
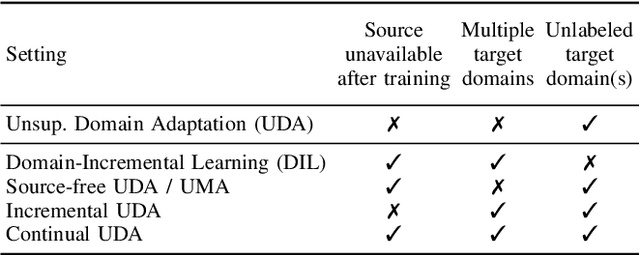
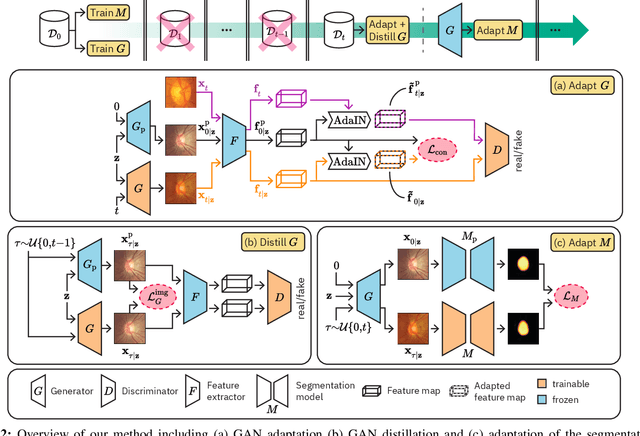
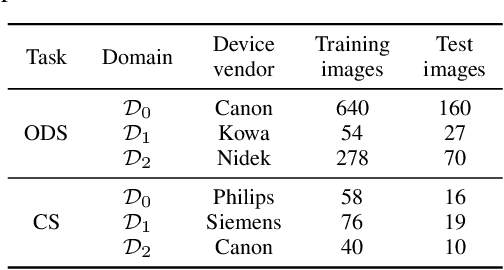
Deep learning models can achieve high accuracy when trained on large amounts of labeled data. However, real-world scenarios often involve several challenges: Training data may become available in installments, may originate from multiple different domains, and may not contain labels for training. Certain settings, for instance medical applications, often involve further restrictions that prohibit retention of previously seen data due to privacy regulations. In this work, to address such challenges, we study unsupervised segmentation in continual learning scenarios that involve domain shift. To that end, we introduce GarDA (Generative Appearance Replay for continual Domain Adaptation), a generative-replay based approach that can adapt a segmentation model sequentially to new domains with unlabeled data. In contrast to single-step unsupervised domain adaptation (UDA), continual adaptation to a sequence of domains enables leveraging and consolidation of information from multiple domains. Unlike previous approaches in incremental UDA, our method does not require access to previously seen data, making it applicable in many practical scenarios. We evaluate GarDA on two datasets with different organs and modalities, where it substantially outperforms existing techniques.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge