"Image": models, code, and papers
A Deep Network for Joint Registration and Reconstruction of Images with Pathologies
Aug 17, 2020
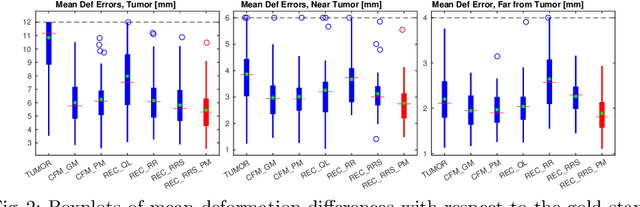
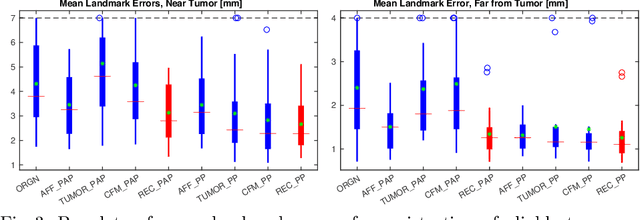
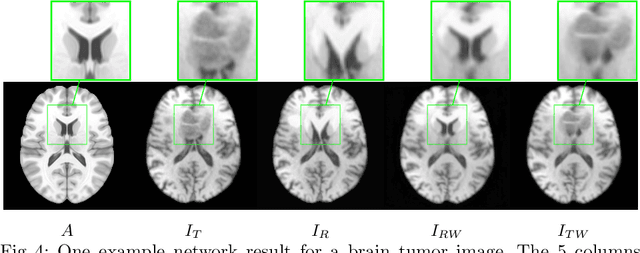
Registration of images with pathologies is challenging due to tissue appearance changes and missing correspondences caused by the pathologies. Moreover, mass effects as observed for brain tumors may displace tissue, creating larger deformations over time than what is observed in a healthy brain. Deep learning models have successfully been applied to image registration to offer dramatic speed up and to use surrogate information (e.g., segmentations) during training. However, existing approaches focus on learning registration models using images from healthy patients. They are therefore not designed for the registration of images with strong pathologies for example in the context of brain tumors, and traumatic brain injuries. In this work, we explore a deep learning approach to register images with brain tumors to an atlas. Our model learns an appearance mapping from images with tumors to the atlas, while simultaneously predicting the transformation to atlas space. Using separate decoders, the network disentangles the tumor mass effect from the reconstruction of quasi-normal images. Results on both synthetic and real brain tumor scans show that our approach outperforms cost function masking for registration to the atlas and that reconstructed quasi-normal images can be used for better longitudinal registrations.
Multi-Stream Networks and Ground-Truth Generation for Crowd Counting
Mar 11, 2020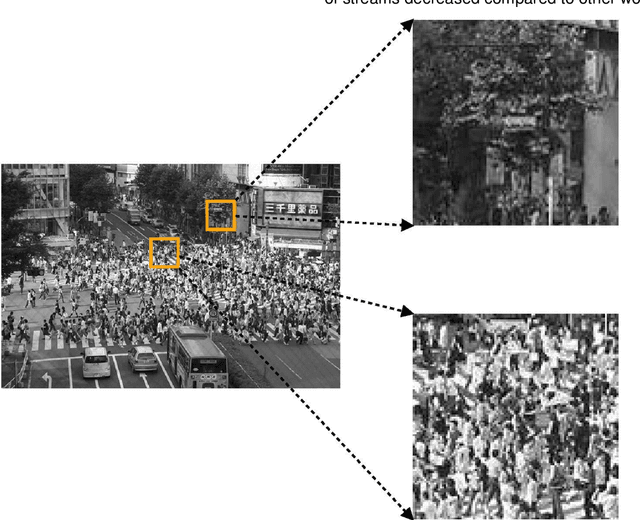
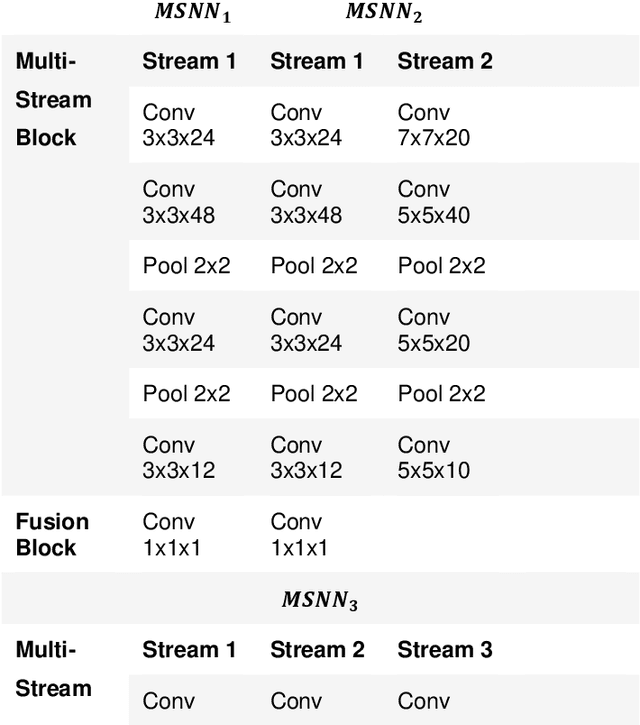
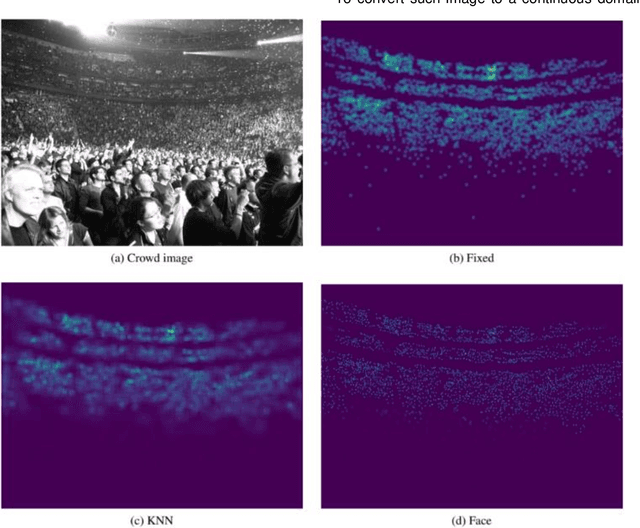
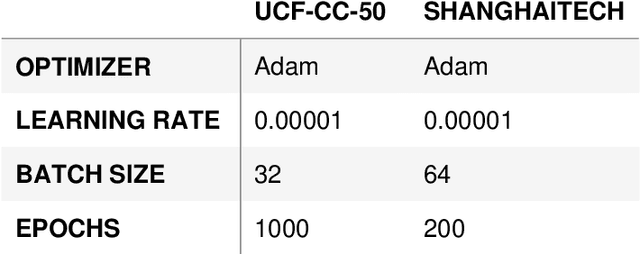
Crowd scene analysis has received a lot of attention recently due to the wide variety of applications, for instance, forensic science, urban planning, surveillance and security. In this context, a challenging task is known as crowd counting, whose main purpose is to estimate the number of people present in a single image. A Multi-Stream Convolutional Neural Network is developed and evaluated in this work, which receives an image as input and produces a density map that represents the spatial distribution of people in an end-to-end fashion. In order to address complex crowd counting issues, such as extremely unconstrained scale and perspective changes, the network architecture utilizes receptive fields with different size filters for each stream. In addition, we investigate the influence of the two most common fashions on the generation of ground truths and propose a hybrid method based on tiny face detection and scale interpolation. Experiments conducted on two challenging datasets, UCF-CC-50 and ShanghaiTech, demonstrate that using our ground truth generation methods achieves superior results.
Anatomy-Aware Cardiac Motion Estimation
Aug 17, 2020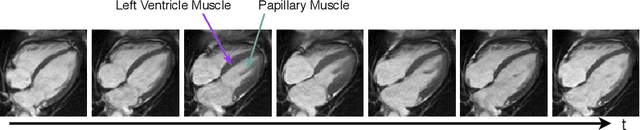
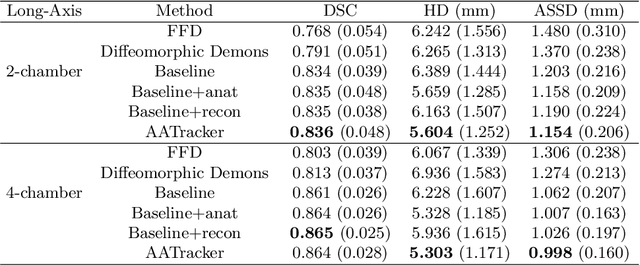
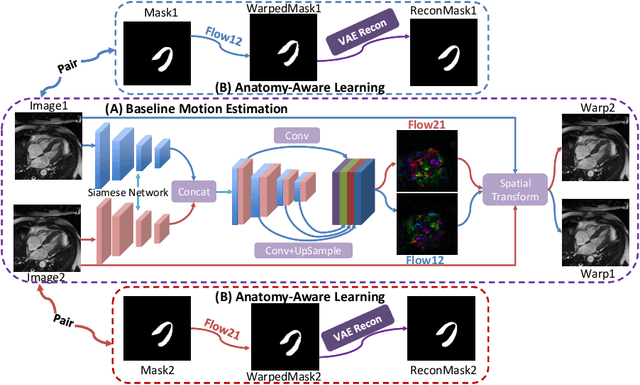
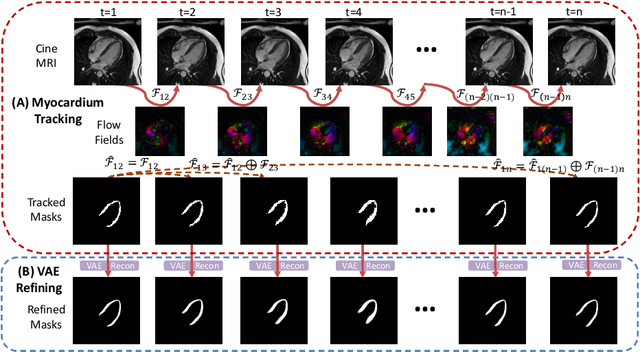
Cardiac motion estimation is critical to the assessment of cardiac function. Myocardium feature tracking (FT) can directly estimate cardiac motion from cine MRI, which requires no special scanning procedure. However, current deep learning-based FT methods may result in unrealistic myocardium shapes since the learning is solely guided by image intensities without considering anatomy. On the other hand, motion estimation through learning is challenging because ground-truth motion fields are almost impossible to obtain. In this study, we propose a novel Anatomy-Aware Tracker (AATracker) for cardiac motion estimation that preserves anatomy by weak supervision. A convolutional variational autoencoder (VAE) is trained to encapsulate realistic myocardium shapes. A baseline dense motion tracker is trained to approximate the motion fields and then refined to estimate anatomy-aware motion fields under the weak supervision from the VAE. We evaluate the proposed method on long-axis cardiac cine MRI, which has more complex myocardium appearances and motions than short-axis. Compared with other methods, AATracker significantly improves the tracking performance and provides visually more realistic tracking results, demonstrating the effectiveness of the proposed weakly-supervision scheme in cardiac motion estimation.
Improving Semantic Segmentation through Spatio-Temporal Consistency Learned from Videos
Apr 11, 2020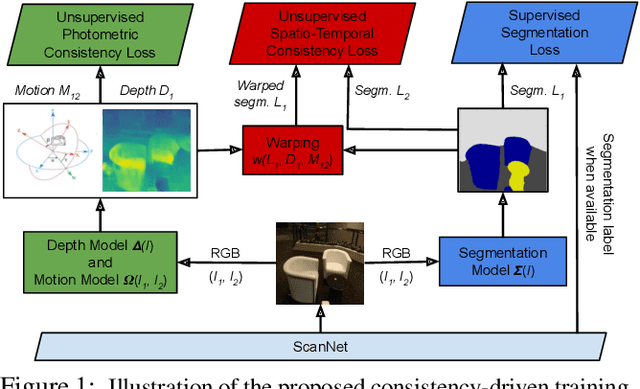
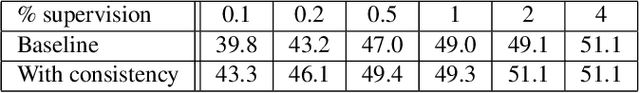
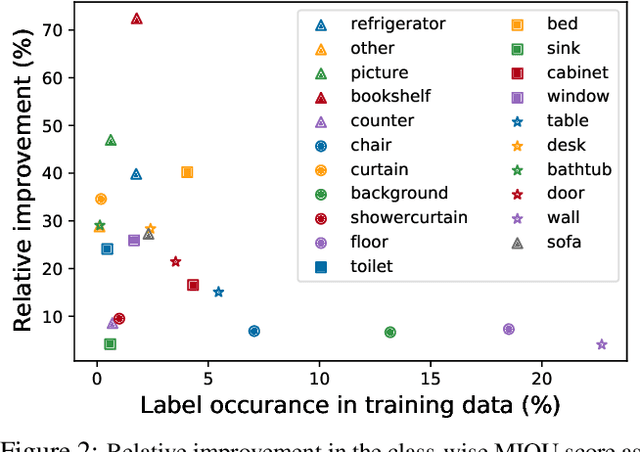
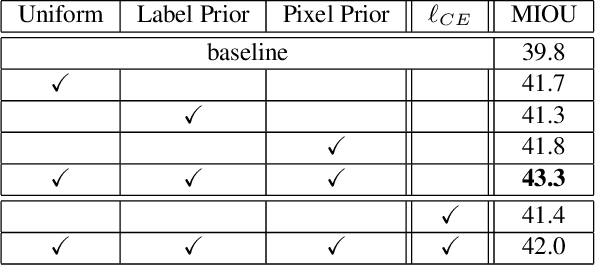
We leverage unsupervised learning of depth, egomotion, and camera intrinsics to improve the performance of single-image semantic segmentation, by enforcing 3D-geometric and temporal consistency of segmentation masks across video frames. The predicted depth, egomotion, and camera intrinsics are used to provide an additional supervision signal to the segmentation model, significantly enhancing its quality, or, alternatively, reducing the number of labels the segmentation model needs. Our experiments were performed on the ScanNet dataset.
Learning Stereo from Single Images
Aug 04, 2020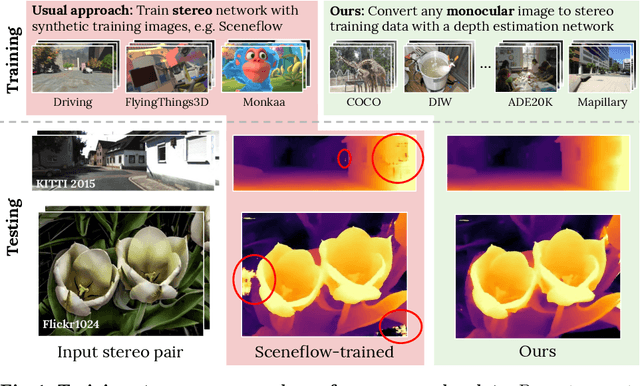
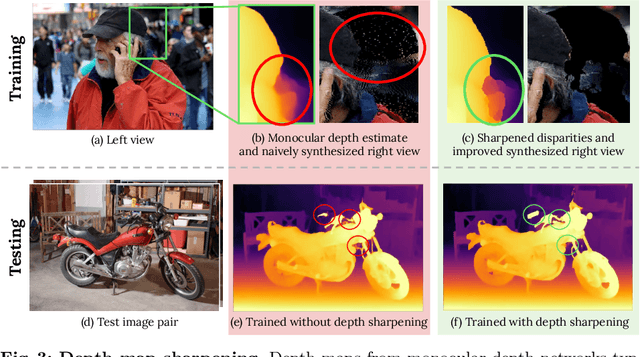
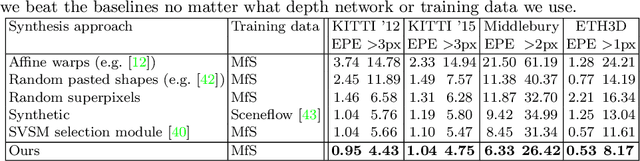
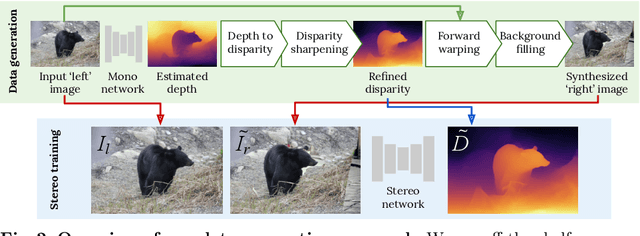

Supervised deep networks are among the best methods for finding correspondences in stereo image pairs. Like all supervised approaches, these networks require ground truth data during training. However, collecting large quantities of accurate dense correspondence data is very challenging. We propose that it is unnecessary to have such a high reliance on ground truth depths or even corresponding stereo pairs. Inspired by recent progress in monocular depth estimation, we generate plausible disparity maps from single images. In turn, we use those flawed disparity maps in a carefully designed pipeline to generate stereo training pairs. Training in this manner makes it possible to convert any collection of single RGB images into stereo training data. This results in a significant reduction in human effort, with no need to collect real depths or to hand-design synthetic data. We can consequently train a stereo matching network from scratch on datasets like COCO, which were previously hard to exploit for stereo. Through extensive experiments we show that our approach outperforms stereo networks trained with standard synthetic datasets, when evaluated on KITTI, ETH3D, and Middlebury.
Tracking Skin Colour and Wrinkle Changes During Cosmetic Product Trials Using Smartphone Images
Aug 04, 2020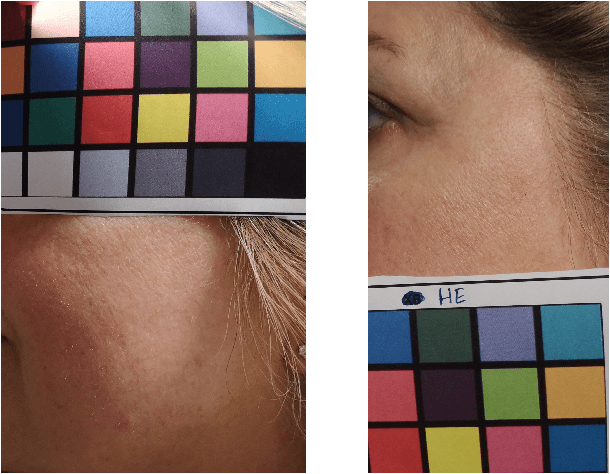
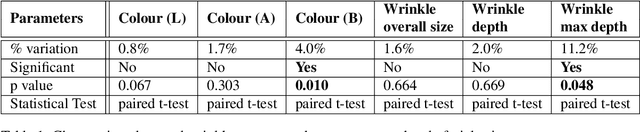
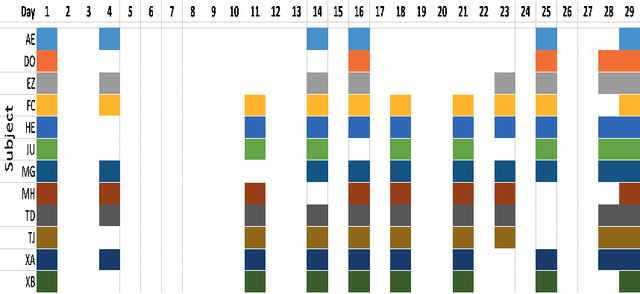
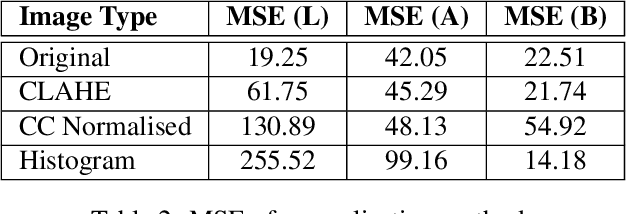
Background: To explore how the efficacy of product trials for skin cosmetics can be improved through the use of consumer-level images taken by volunteers using a conventional smartphone. Materials and Methods: 12 women aged 30 to 60 years participated in a product trial and had close-up images of the cheek and temple regions of their faces taken with a high-resolution Antera 3D CS camera at the start and end of a 4-week period. Additionally, they each had ``selfies'' of the same regions of their faces taken regularly throughout the trial period. Automatic image analysis to identify changes in skin colour used three kinds of colour normalisation and analysis for wrinkle composition identified edges and calculated their magnitude. Results: Images taken at the start and end of the trial acted as baseline ground truth for normalisation of smartphone images and showed large changes in both colour and wrinkle magnitude during the trial for many volunteers. Conclusions: Results demonstrate that regular use of selfie smartphone images within trial periods can add value to interpretation of the efficacy of the trial.
First U-Net Layers Contain More Domain Specific Information Than The Last Ones
Aug 17, 2020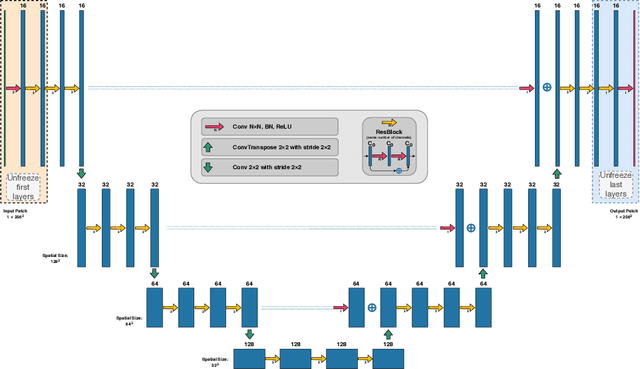
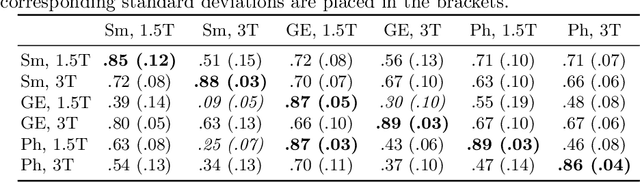
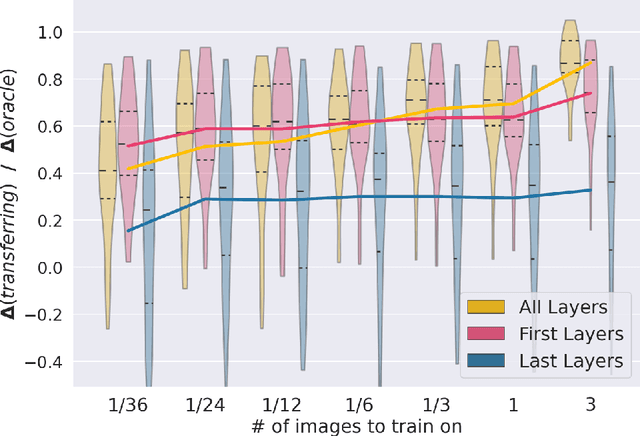
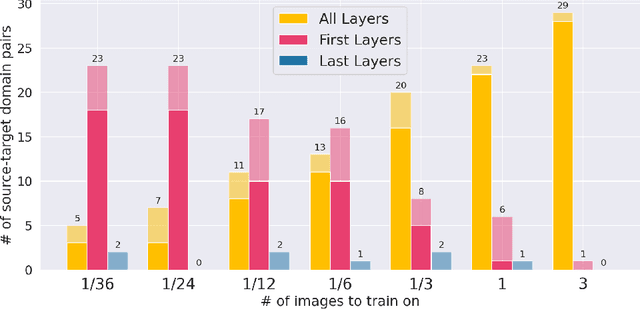
MRI scans appearance significantly depends on scanning protocols and, consequently, the data-collection institution. These variations between clinical sites result in dramatic drops of CNN segmentation quality on unseen domains. Many of the recently proposed MRI domain adaptation methods operate with the last CNN layers to suppress domain shift. At the same time, the core manifestation of MRI variability is a considerable diversity of image intensities. We hypothesize that these differences can be eliminated by modifying the first layers rather than the last ones. To validate this simple idea, we conducted a set of experiments with brain MRI scans from six domains. Our results demonstrate that 1) domain-shift may deteriorate the quality even for a simple brain extraction segmentation task (surface Dice Score drops from 0.85-0.89 even to 0.09); 2) fine-tuning of the first layers significantly outperforms fine-tuning of the last layers in almost all supervised domain adaptation setups. Moreover, fine-tuning of the first layers is a better strategy than fine-tuning of the whole network, if the amount of annotated data from the new domain is strictly limited.
Mining self-similarity: Label super-resolution with epitomic representations
Apr 24, 2020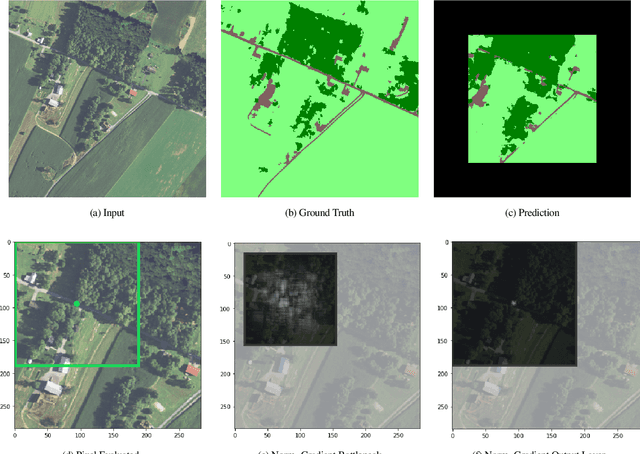
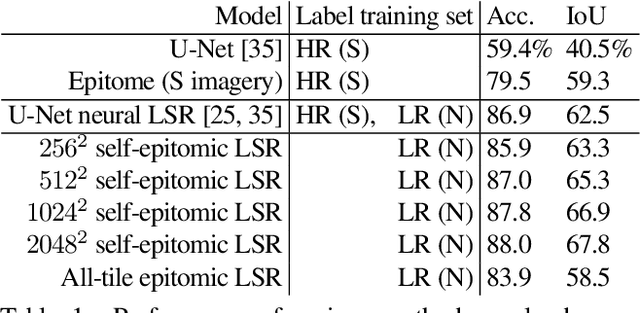

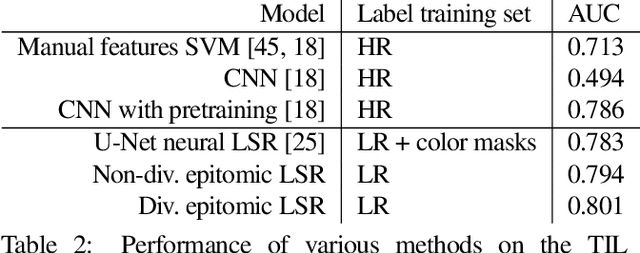
We show that simple patch-based models, such as epitomes, can have superior performance to the current state of the art in semantic segmentation and label super-resolution, which uses deep convolutional neural networks. We derive a new training algorithm for epitomes which allows, for the first time, learning from very large data sets and derive a label super-resolution algorithm as a statistical inference algorithm over epitomic representations. We illustrate our methods on land cover mapping and medical image analysis tasks.
DOC: Deep OCclusion Estimation From a Single Image
Jul 24, 2016
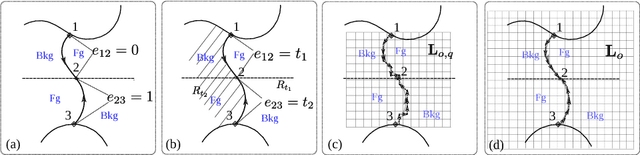
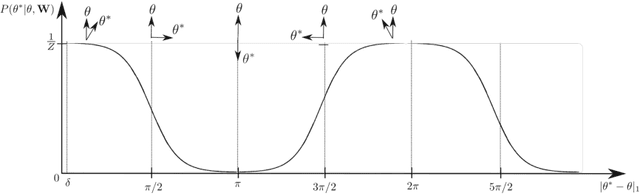
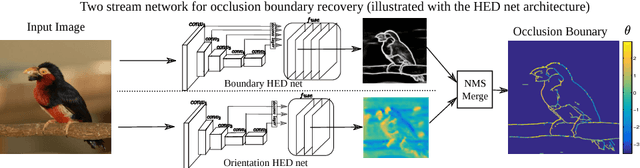
Recovering the occlusion relationships between objects is a fundamental human visual ability which yields important information about the 3D world. In this paper we propose a deep network architecture, called DOC, which acts on a single image, detects object boundaries and estimates the border ownership (i.e. which side of the boundary is foreground and which is background). We represent occlusion relations by a binary edge map, to indicate the object boundary, and an occlusion orientation variable which is tangential to the boundary and whose direction specifies border ownership by a left-hand rule. We train two related deep convolutional neural networks, called DOC, which exploit local and non-local image cues to estimate this representation and hence recover occlusion relations. In order to train and test DOC we construct a large-scale instance occlusion boundary dataset using PASCAL VOC images, which we call the PASCAL instance occlusion dataset (PIOD). This contains 10,000 images and hence is two orders of magnitude larger than existing occlusion datasets for outdoor images. We test two variants of DOC on PIOD and on the BSDS occlusion dataset and show they outperform state-of-the-art methods. Finally, we perform numerous experiments investigating multiple settings of DOC and transfer between BSDS and PIOD, which provides more insights for further study of occlusion estimation.
Learning Invariant Feature Representation to Improve Generalization across Chest X-ray Datasets
Aug 04, 2020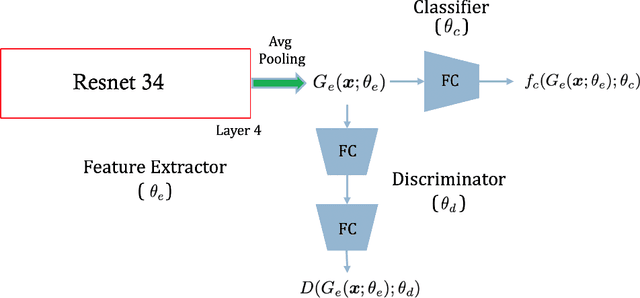
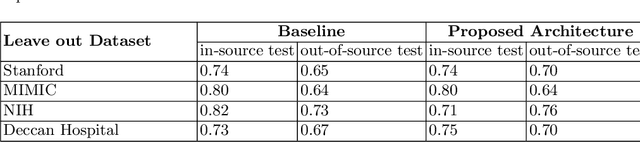
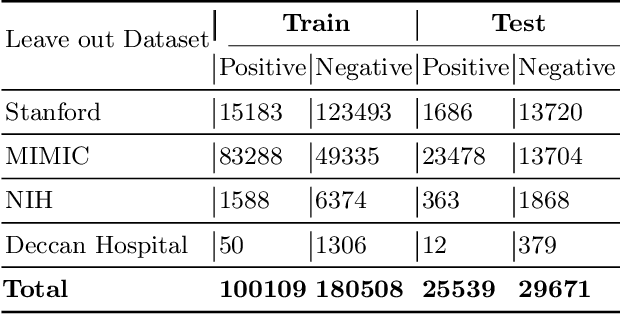

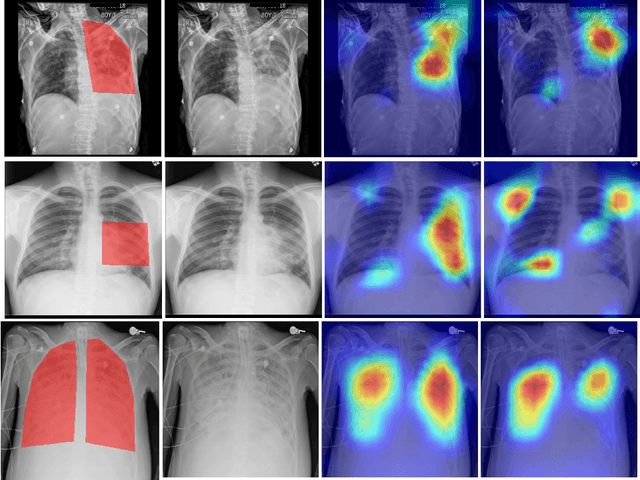
Chest radiography is the most common medical image examination for screening and diagnosis in hospitals. Automatic interpretation of chest X-rays at the level of an entry-level radiologist can greatly benefit work prioritization and assist in analyzing a larger population. Subsequently, several datasets and deep learning-based solutions have been proposed to identify diseases based on chest X-ray images. However, these methods are shown to be vulnerable to shift in the source of data: a deep learning model performing well when tested on the same dataset as training data, starts to perform poorly when it is tested on a dataset from a different source. In this work, we address this challenge of generalization to a new source by forcing the network to learn a source-invariant representation. By employing an adversarial training strategy, we show that a network can be forced to learn a source-invariant representation. Through pneumonia-classification experiments on multi-source chest X-ray datasets, we show that this algorithm helps in improving classification accuracy on a new source of X-ray dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge