Joy T. Wu
Evaluating Automated Radiology Report Quality through Fine-Grained Phrasal Grounding of Clinical Findings
Dec 02, 2024



Abstract:Several evaluation metrics have been developed recently to automatically assess the quality of generative AI reports for chest radiographs based only on textual information using lexical, semantic, or clinical named entity recognition methods. In this paper, we develop a new method of report quality evaluation by first extracting fine-grained finding patterns capturing the location, laterality, and severity of a large number of clinical findings. We then performed phrasal grounding to localize their associated anatomical regions on chest radiograph images. The textual and visual measures are then combined to rate the quality of the generated reports. We present results that compare this evaluation metric with other textual metrics on a gold standard dataset derived from the MIMIC collection and show its robustness and sensitivity to factual errors.
CheXRelNet: An Anatomy-Aware Model for Tracking Longitudinal Relationships between Chest X-Rays
Aug 08, 2022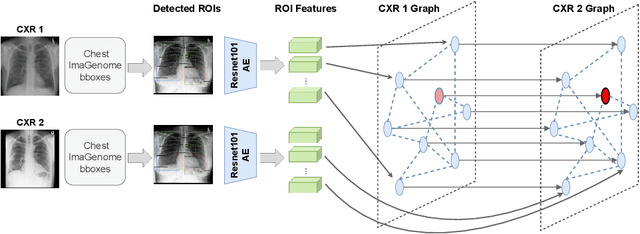
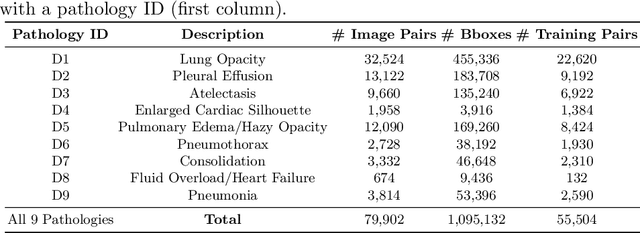
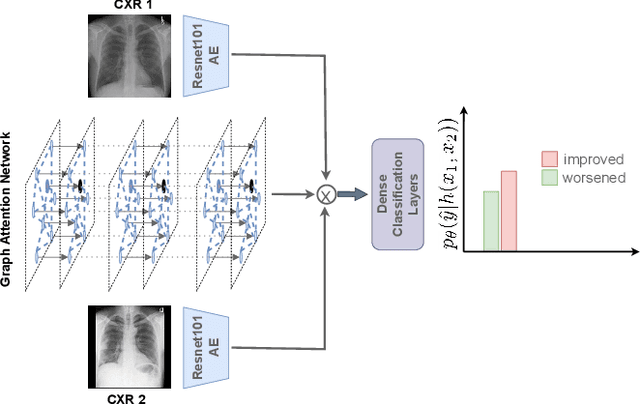
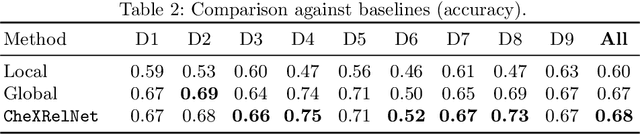
Abstract:Despite the progress in utilizing deep learning to automate chest radiograph interpretation and disease diagnosis tasks, change between sequential Chest X-rays (CXRs) has received limited attention. Monitoring the progression of pathologies that are visualized through chest imaging poses several challenges in anatomical motion estimation and image registration, i.e., spatially aligning the two images and modeling temporal dynamics in change detection. In this work, we propose CheXRelNet, a neural model that can track longitudinal pathology change relations between two CXRs. CheXRelNet incorporates local and global visual features, utilizes inter-image and intra-image anatomical information, and learns dependencies between anatomical region attributes, to accurately predict disease change for a pair of CXRs. Experimental results on the Chest ImaGenome dataset show increased downstream performance compared to baselines. Code is available at https://github.com/PLAN-Lab/ChexRelNet
Chest ImaGenome Dataset for Clinical Reasoning
Jul 31, 2021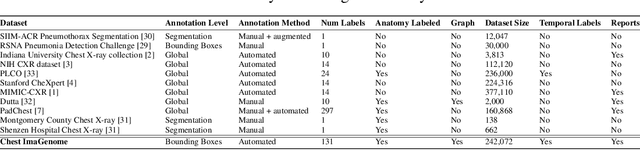
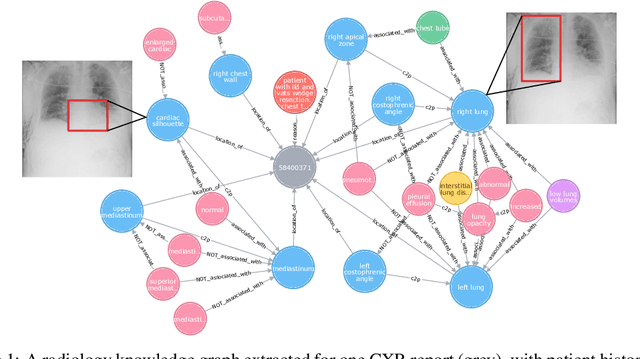
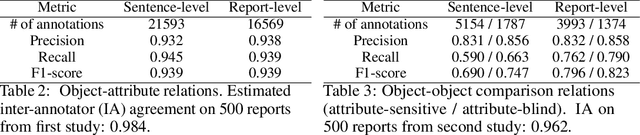
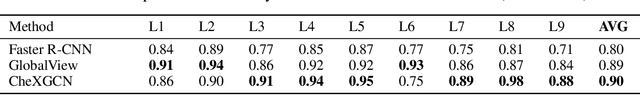
Abstract:Despite the progress in automatic detection of radiologic findings from chest X-ray (CXR) images in recent years, a quantitative evaluation of the explainability of these models is hampered by the lack of locally labeled datasets for different findings. With the exception of a few expert-labeled small-scale datasets for specific findings, such as pneumonia and pneumothorax, most of the CXR deep learning models to date are trained on global "weak" labels extracted from text reports, or trained via a joint image and unstructured text learning strategy. Inspired by the Visual Genome effort in the computer vision community, we constructed the first Chest ImaGenome dataset with a scene graph data structure to describe $242,072$ images. Local annotations are automatically produced using a joint rule-based natural language processing (NLP) and atlas-based bounding box detection pipeline. Through a radiologist constructed CXR ontology, the annotations for each CXR are connected as an anatomy-centered scene graph, useful for image-level reasoning and multimodal fusion applications. Overall, we provide: i) $1,256$ combinations of relation annotations between $29$ CXR anatomical locations (objects with bounding box coordinates) and their attributes, structured as a scene graph per image, ii) over $670,000$ localized comparison relations (for improved, worsened, or no change) between the anatomical locations across sequential exams, as well as ii) a manually annotated gold standard scene graph dataset from $500$ unique patients.
AnaXNet: Anatomy Aware Multi-label Finding Classification in Chest X-ray
May 20, 2021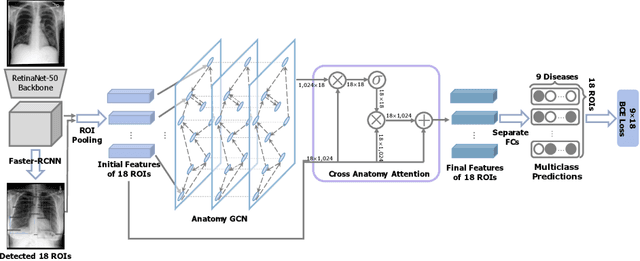
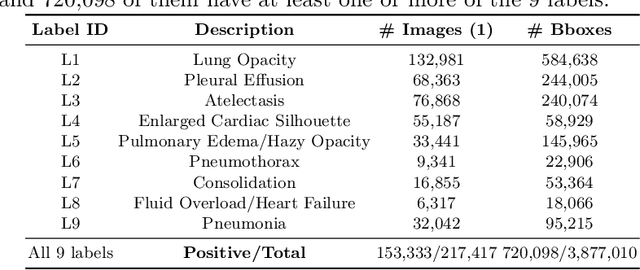
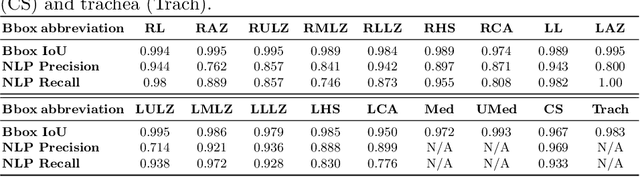
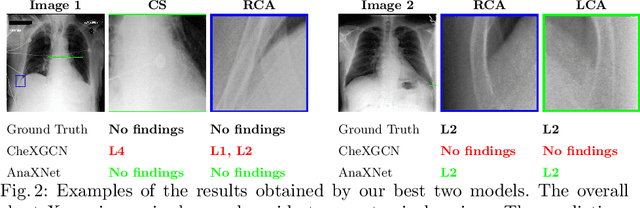
Abstract:Radiologists usually observe anatomical regions of chest X-ray images as well as the overall image before making a decision. However, most existing deep learning models only look at the entire X-ray image for classification, failing to utilize important anatomical information. In this paper, we propose a novel multi-label chest X-ray classification model that accurately classifies the image finding and also localizes the findings to their correct anatomical regions. Specifically, our model consists of two modules, the detection module and the anatomical dependency module. The latter utilizes graph convolutional networks, which enable our model to learn not only the label dependency but also the relationship between the anatomical regions in the chest X-ray. We further utilize a method to efficiently create an adjacency matrix for the anatomical regions using the correlation of the label across the different regions. Detailed experiments and analysis of our results show the effectiveness of our method when compared to the current state-of-the-art multi-label chest X-ray image classification methods while also providing accurate location information.
Extracting and Learning Fine-Grained Labels from Chest Radiographs
Nov 18, 2020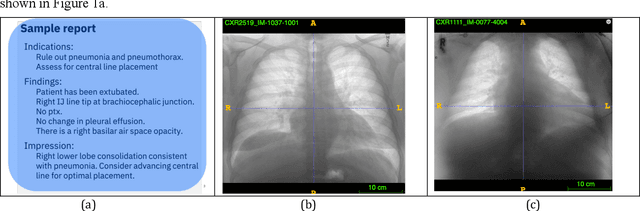
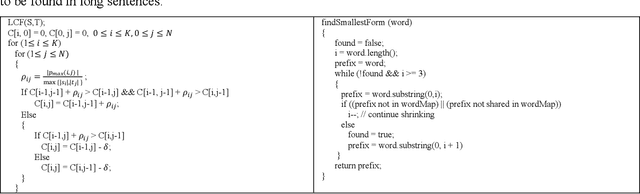
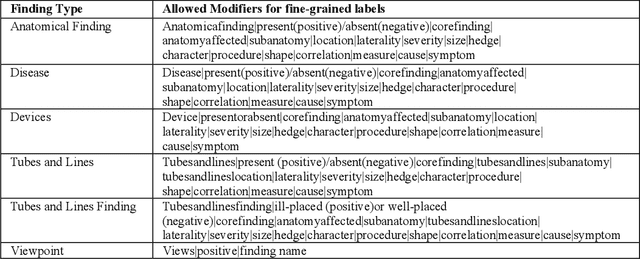
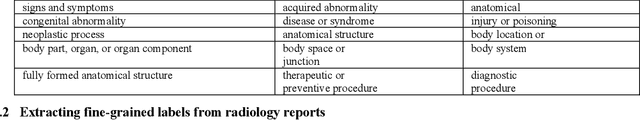

Abstract:Chest radiographs are the most common diagnostic exam in emergency rooms and intensive care units today. Recently, a number of researchers have begun working on large chest X-ray datasets to develop deep learning models for recognition of a handful of coarse finding classes such as opacities, masses and nodules. In this paper, we focus on extracting and learning fine-grained labels for chest X-ray images. Specifically we develop a new method of extracting fine-grained labels from radiology reports by combining vocabulary-driven concept extraction with phrasal grouping in dependency parse trees for association of modifiers with findings. A total of 457 fine-grained labels depicting the largest spectrum of findings to date were selected and sufficiently large datasets acquired to train a new deep learning model designed for fine-grained classification. We show results that indicate a highly accurate label extraction process and a reliable learning of fine-grained labels. The resulting network, to our knowledge, is the first to recognize fine-grained descriptions of findings in images covering over nine modifiers including laterality, location, severity, size and appearance.
Learning Invariant Feature Representation to Improve Generalization across Chest X-ray Datasets
Aug 04, 2020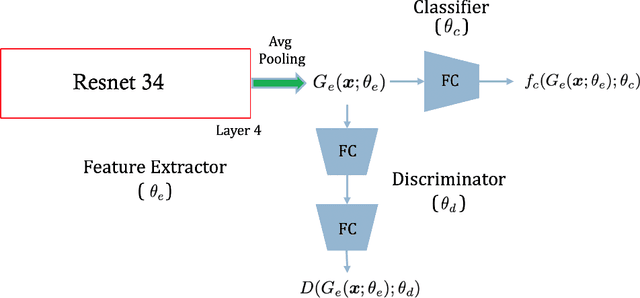
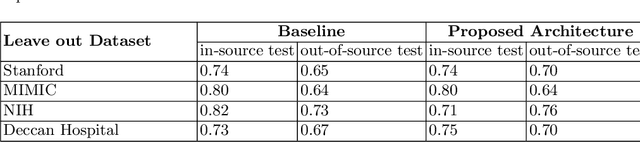
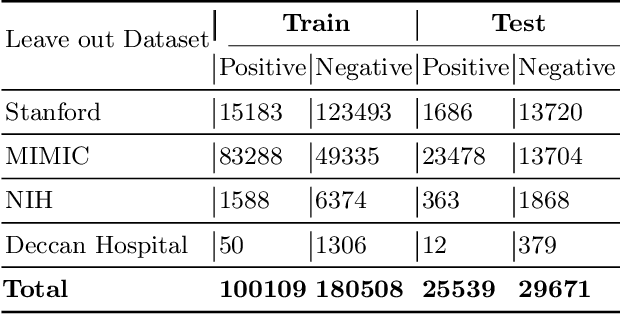

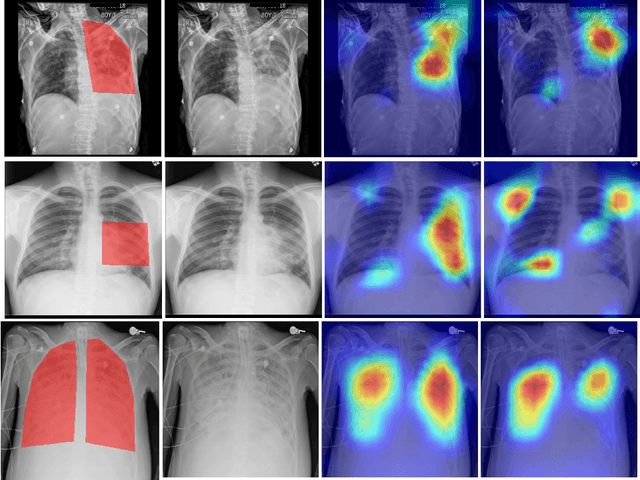
Abstract:Chest radiography is the most common medical image examination for screening and diagnosis in hospitals. Automatic interpretation of chest X-rays at the level of an entry-level radiologist can greatly benefit work prioritization and assist in analyzing a larger population. Subsequently, several datasets and deep learning-based solutions have been proposed to identify diseases based on chest X-ray images. However, these methods are shown to be vulnerable to shift in the source of data: a deep learning model performing well when tested on the same dataset as training data, starts to perform poorly when it is tested on a dataset from a different source. In this work, we address this challenge of generalization to a new source by forcing the network to learn a source-invariant representation. By employing an adversarial training strategy, we show that a network can be forced to learn a source-invariant representation. Through pneumonia-classification experiments on multi-source chest X-ray datasets, we show that this algorithm helps in improving classification accuracy on a new source of X-ray dataset.
Chest X-ray Report Generation through Fine-Grained Label Learning
Jul 27, 2020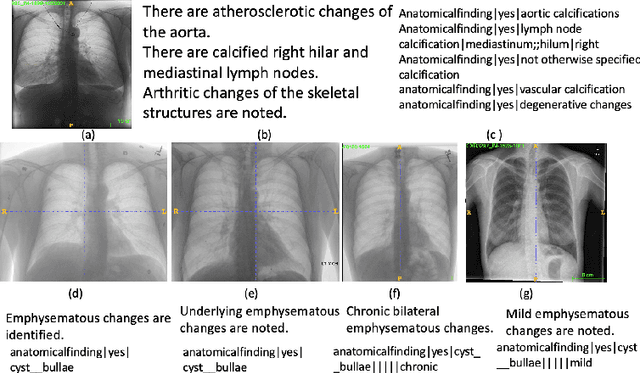
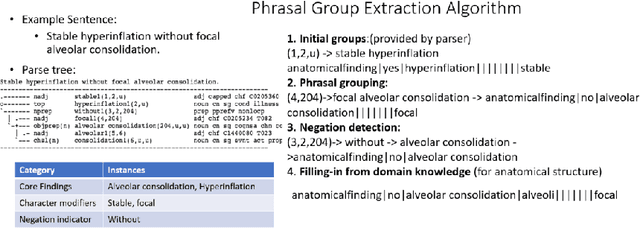
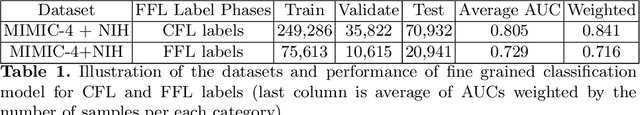

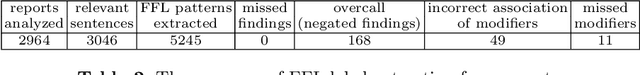
Abstract:Obtaining automated preliminary read reports for common exams such as chest X-rays will expedite clinical workflows and improve operational efficiencies in hospitals. However, the quality of reports generated by current automated approaches is not yet clinically acceptable as they cannot ensure the correct detection of a broad spectrum of radiographic findings nor describe them accurately in terms of laterality, anatomical location, severity, etc. In this work, we present a domain-aware automatic chest X-ray radiology report generation algorithm that learns fine-grained description of findings from images and uses their pattern of occurrences to retrieve and customize similar reports from a large report database. We also develop an automatic labeling algorithm for assigning such descriptors to images and build a novel deep learning network that recognizes both coarse and fine-grained descriptions of findings. The resulting report generation algorithm significantly outperforms the state of the art using established score metrics.
A Corpus for Detecting High-Context Medical Conditions in Intensive Care Patient Notes Focusing on Frequently Readmitted Patients
Mar 06, 2020
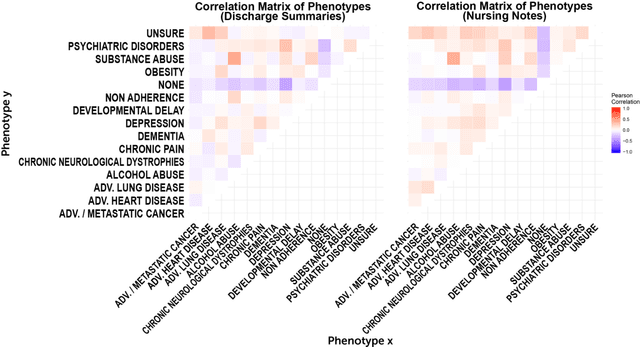
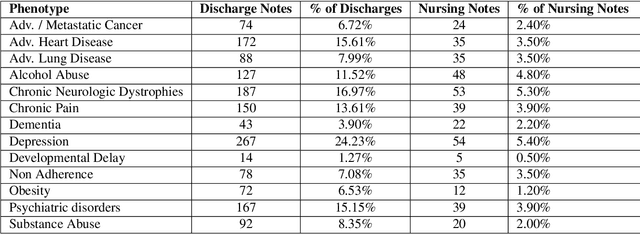
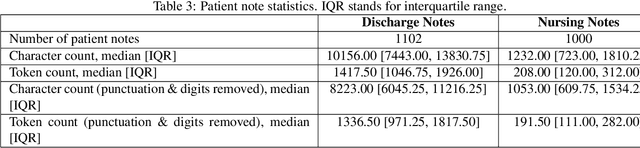
Abstract:A crucial step within secondary analysis of electronic health records (EHRs) is to identify the patient cohort under investigation. While EHRs contain medical billing codes that aim to represent the conditions and treatments patients may have, much of the information is only present in the patient notes. Therefore, it is critical to develop robust algorithms to infer patients' conditions and treatments from their written notes. In this paper, we introduce a dataset for patient phenotyping, a task that is defined as the identification of whether a patient has a given medical condition (also referred to as clinical indication or phenotype) based on their patient note. Nursing Progress Notes and Discharge Summaries from the Intensive Care Unit of a large tertiary care hospital were manually annotated for the presence of several high-context phenotypes relevant to treatment and risk of re-hospitalization. This dataset contains 1102 Discharge Summaries and 1000 Nursing Progress Notes. Each Discharge Summary and Progress Note has been annotated by at least two expert human annotators (one clinical researcher and one resident physician). Annotated phenotypes include treatment non-adherence, chronic pain, advanced/metastatic cancer, as well as 10 other phenotypes. This dataset can be utilized for academic and industrial research in medicine and computer science, particularly within the field of medical natural language processing.
Automated Detection and Type Classification of Central Venous Catheters in Chest X-Rays
Jul 25, 2019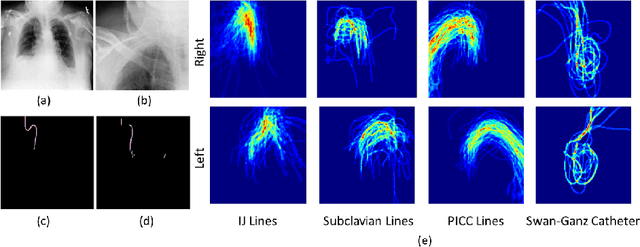
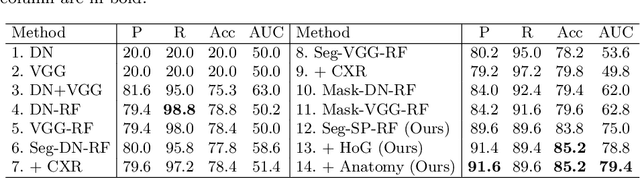
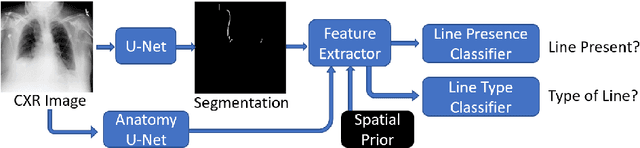
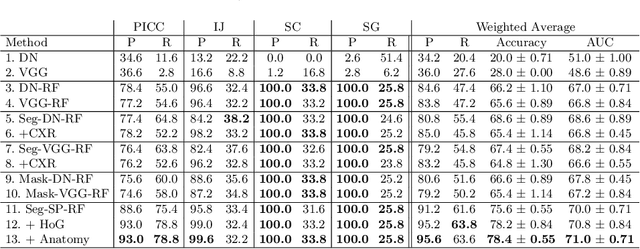
Abstract:Central venous catheters (CVCs) are commonly used in critical care settings for monitoring body functions and administering medications. They are often described in radiology reports by referring to their presence, identity and placement. In this paper, we address the problem of automatic detection of their presence and identity through automated segmentation using deep learning networks and classification based on their intersection with previously learned shape priors from clinician annotations of CVCs. The results not only outperform existing methods of catheter detection achieving 85.2% accuracy at 91.6% precision, but also enable high precision (95.2%) classification of catheter types on a large dataset of over 10,000 chest X-rays, presenting a robust and practical solution to this problem.
Boosting the rule-out accuracy of deep disease detection using class weight modifiers
Jun 21, 2019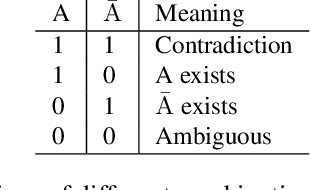
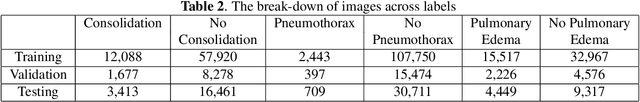
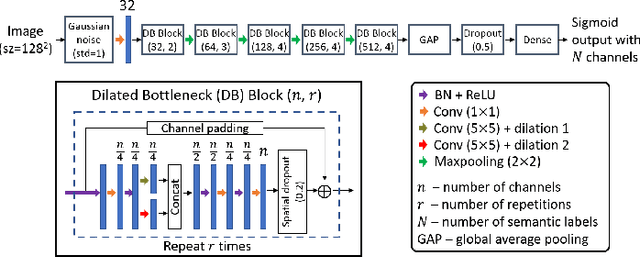

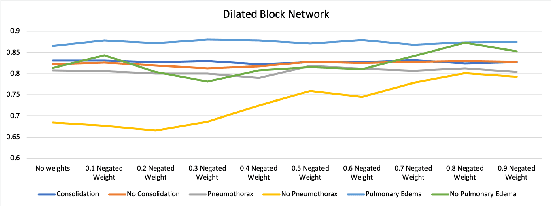
Abstract:In many screening applications, the primary goal of a radiologist or assisting artificial intelligence is to rule out certain findings. The classifiers built for such applications are often trained on large datasets that derive labels from clinical notes written for patients. While the quality of the positive findings described in these notes is often reliable, lack of the mention of a finding does not always rule out the presence of it. This happens because radiologists comment on the patient in the context of the exam, for example focusing on trauma as opposed to chronic disease at emergency rooms. However, this disease finding ambiguity can affect the performance of algorithms. Hence it is critical to model the ambiguity during training. We propose a scheme to apply reasonable class weight modifiers to our loss function for the no mention cases during training. We experiment with two different deep neural network architectures and show that the proposed method results in a large improvement in the performance of the classifiers, specially on negated findings. The baseline performance of a custom-made dilated block network proposed in this paper shows an improvement in comparison with baseline DenseNet-201, while both architectures benefit from the new proposed loss function weighting scheme. Over 200,000 chest X-ray images and three highly common diseases, along with their negated counterparts, are included in this study.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge