"Image": models, code, and papers
Automatically Designing CNN Architectures Using Genetic Algorithm for Image Classification
Aug 11, 2018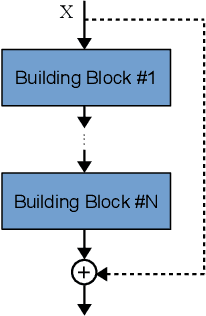
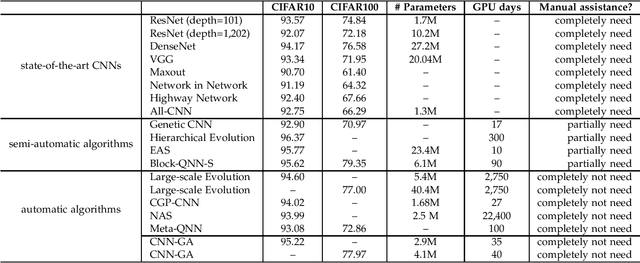

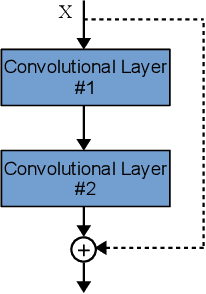
Convolutional Neural Networks (CNNs) have gained a remarkable success on many real-world problems in recent years. However, the performance of CNNs is highly relied on their architectures. For some state-of-the-art CNNs, their architectures are hand-crafted with expertise in both CNNs and the investigated problems. To this end, it is difficult for researchers, who have no extended expertise in CNNs, to explore CNNs for their own problems of interest. In this paper, we propose an automatic architecture design method for CNNs by using genetic algorithms, which is capable of discovering a promising architecture of a CNN on handling image classification tasks. The proposed algorithm does not need any pre-processing before it works, nor any post-processing on the discovered CNN, which means it is completely automatic. The proposed algorithm is validated on widely used benchmark datasets, by comparing to the state-of-the-art peer competitors covering eight manually designed CNNs, four semi-automatically designed CNNs and additional four automatically designed CNNs. The experimental results indicate that the proposed algorithm achieves the best classification accuracy consistently among manually and automatically designed CNNs. Furthermore, the proposed algorithm also shows the competitive classification accuracy to the semi-automatic peer competitors, while reducing 10 times of the parameters. In addition, on the average the proposed algorithm takes only one percentage of computational resource compared to that of all the other architecture discovering algorithms.
Cross-modal Subspace Learning for Fine-grained Sketch-based Image Retrieval
May 28, 2017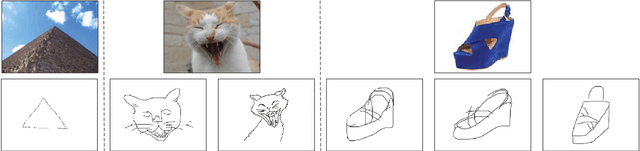
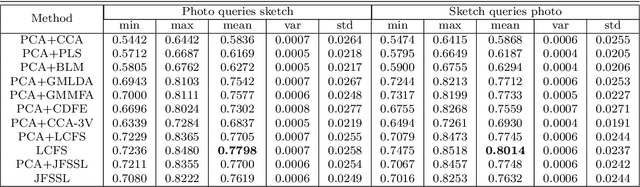
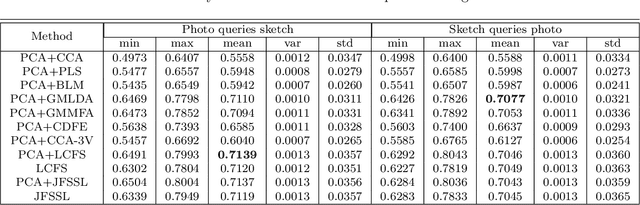
Sketch-based image retrieval (SBIR) is challenging due to the inherent domain-gap between sketch and photo. Compared with pixel-perfect depictions of photos, sketches are iconic renderings of the real world with highly abstract. Therefore, matching sketch and photo directly using low-level visual clues are unsufficient, since a common low-level subspace that traverses semantically across the two modalities is non-trivial to establish. Most existing SBIR studies do not directly tackle this cross-modal problem. This naturally motivates us to explore the effectiveness of cross-modal retrieval methods in SBIR, which have been applied in the image-text matching successfully. In this paper, we introduce and compare a series of state-of-the-art cross-modal subspace learning methods and benchmark them on two recently released fine-grained SBIR datasets. Through thorough examination of the experimental results, we have demonstrated that the subspace learning can effectively model the sketch-photo domain-gap. In addition we draw a few key insights to drive future research.
Polyhedron Volume-Ratio-based Classification for Image Recognition
Jan 26, 2016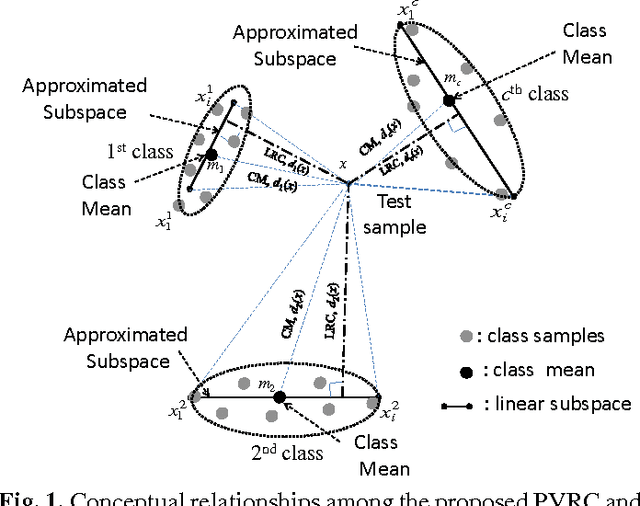

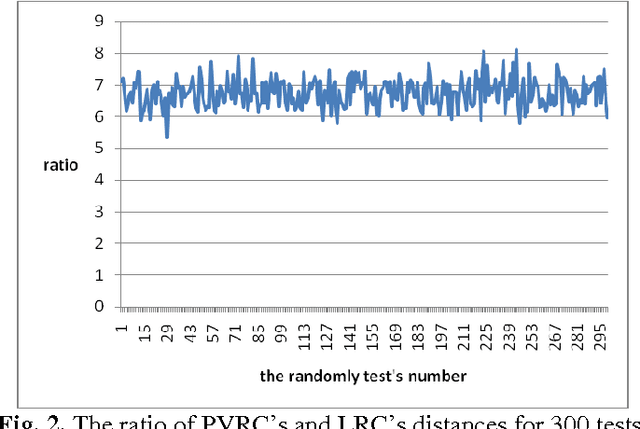

In this paper, a novel method, called polyhedron volume ratio classification (PVRC) is proposed for image recognition
Learning Colour Representations of Search Queries
Jun 17, 2020
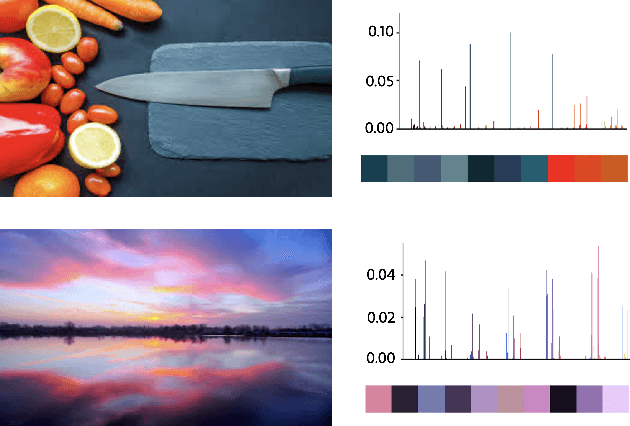

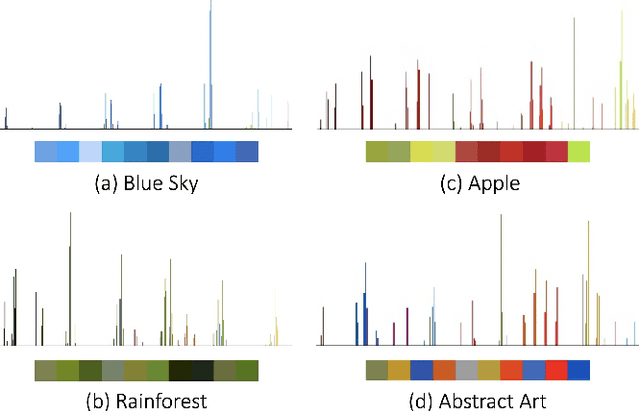
Image search engines rely on appropriately designed ranking features that capture various aspects of the content semantics as well as the historic popularity. In this work, we consider the role of colour in this relevance matching process. Our work is motivated by the observation that a significant fraction of user queries have an inherent colour associated with them. While some queries contain explicit colour mentions (such as 'black car' and 'yellow daisies'), other queries have implicit notions of colour (such as 'sky' and 'grass'). Furthermore, grounding queries in colour is not a mapping to a single colour, but a distribution in colour space. For instance, a search for 'trees' tends to have a bimodal distribution around the colours green and brown. We leverage historical clickthrough data to produce a colour representation for search queries and propose a recurrent neural network architecture to encode unseen queries into colour space. We also show how this embedding can be learnt alongside a cross-modal relevance ranker from impression logs where a subset of the result images were clicked. We demonstrate that the use of a query-image colour distance feature leads to an improvement in the ranker performance as measured by users' preferences of clicked versus skipped images.
Does Data Augmentation Benefit from Split BatchNorms
Oct 15, 2020
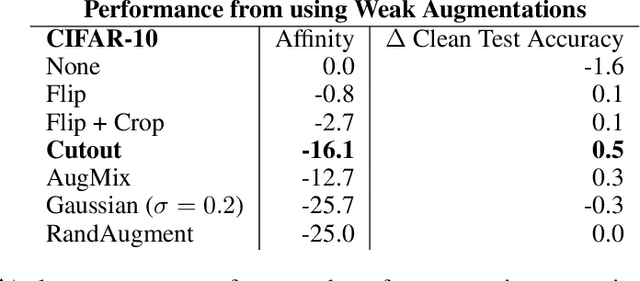
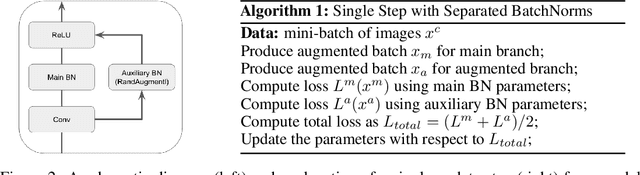


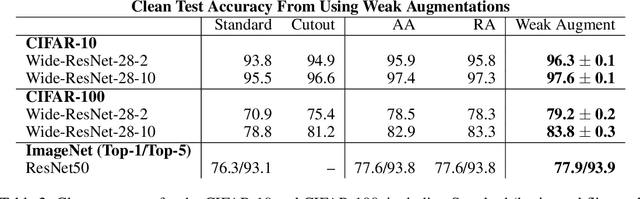
Data augmentation has emerged as a powerful technique for improving the performance of deep neural networks and led to state-of-the-art results in computer vision. However, state-of-the-art data augmentation strongly distorts training images, leading to a disparity between examples seen during training and inference. In this work, we explore a recently proposed training paradigm in order to correct for this disparity: using an auxiliary BatchNorm for the potentially out-of-distribution, strongly augmented images. Our experiments then focus on how to define the BatchNorm parameters that are used at evaluation. To eliminate the train-test disparity, we experiment with using the batch statistics defined by clean training images only, yet surprisingly find that this does not yield improvements in model performance. Instead, we investigate using BatchNorm parameters defined by weak augmentations and find that this method significantly improves the performance of common image classification benchmarks such as CIFAR-10, CIFAR-100, and ImageNet. We then explore a fundamental trade-off between accuracy and robustness coming from using different BatchNorm parameters, providing greater insight into the benefits of data augmentation on model performance.
A Comparison of Deep Learning Convolution Neural Networks for Liver Segmentation in Radial Turbo Spin Echo Images
Apr 13, 2020



Motion-robust 2D Radial Turbo Spin Echo (RADTSE) pulse sequence can provide a high-resolution composite image, T2-weighted images at multiple echo times (TEs), and a quantitative T2 map, all from a single k-space acquisition. In this work, we use a deep-learning convolutional neural network (CNN) for the segmentation of liver in abdominal RADTSE images. A modified UNET architecture with generalized dice loss objective function was implemented. Three 2D CNNs were trained, one for each image type obtained from the RADTSE sequence. On evaluating the performance of the CNNs on the validation set, we found that CNNs trained on TE images or the T2 maps had higher average dice scores than the composite images. This, in turn, implies that the information regarding T2 variation in tissues aids in improving the segmentation performance.
Longevity Associated Geometry Identified in Satellite Images: Sidewalks, Driveways and Hiking Trails
Mar 05, 2020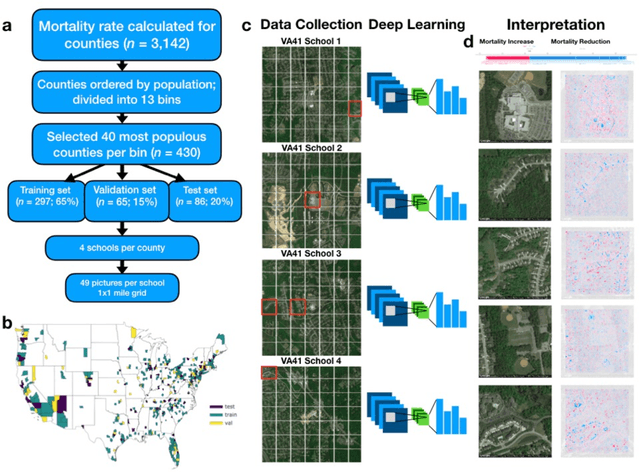
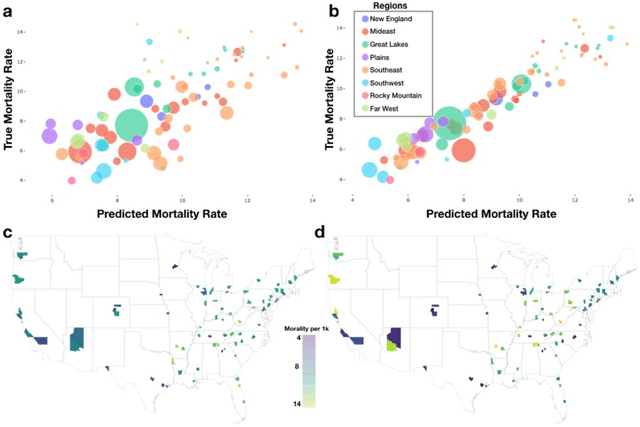
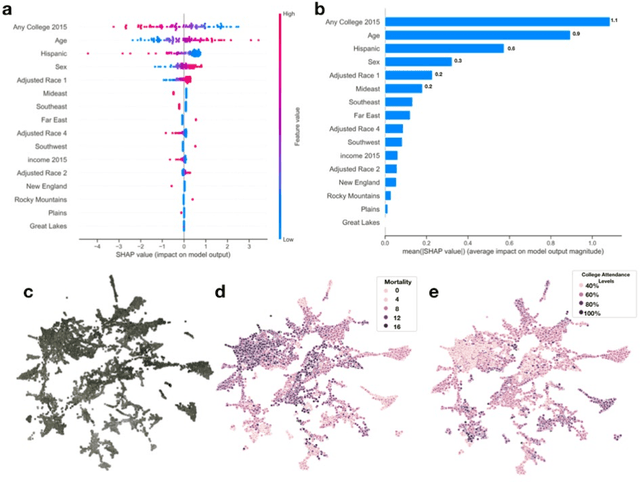
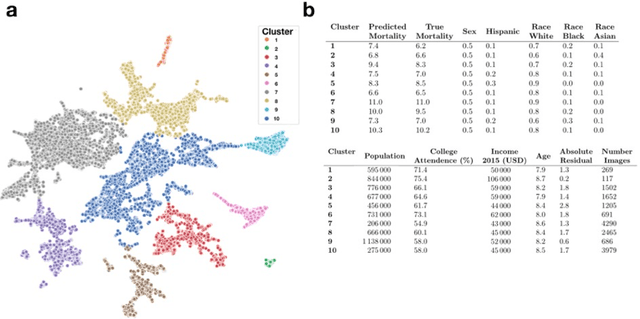
Importance: Following a century of increase, life expectancy in the United States has stagnated and begun to decline in recent decades. Using satellite images and street view images prior work has demonstrated associations of the built environment with income, education, access to care and health factors such as obesity. However, assessment of learned image feature relationships with variation in crude mortality rate across the United States has been lacking. Objective: Investigate prediction of county-level mortality rates in the U.S. using satellite images. Design: Satellite images were extracted with the Google Static Maps application programming interface for 430 counties representing approximately 68.9% of the US population. A convolutional neural network was trained using crude mortality rates for each county in 2015 to predict mortality. Learned image features were interpreted using Shapley Additive Feature Explanations, clustered, and compared to mortality and its associated covariate predictors. Main Outcomes and Measures: County mortality was predicted using satellite images. Results: Predicted mortality from satellite images in a held-out test set of counties was strongly correlated to the true crude mortality rate (Pearson r=0.72). Learned image features were clustered, and we identified 10 clusters that were associated with education, income, geographical region, race and age. Conclusion and Relevance: The application of deep learning techniques to remotely-sensed features of the built environment can serve as a useful predictor of mortality in the United States. Tools that are able to identify image features associated with health-related outcomes can inform targeted public health interventions.
Global hard thresholding algorithms for joint sparse image representation and denoising
May 27, 2017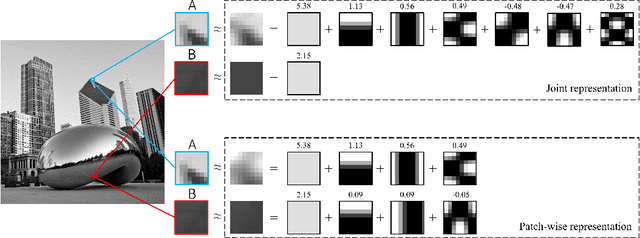
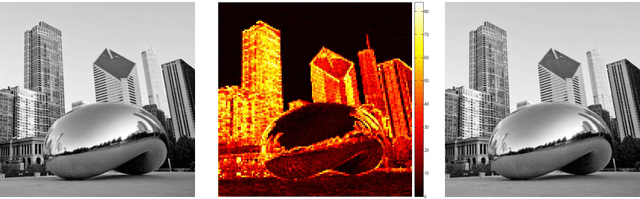
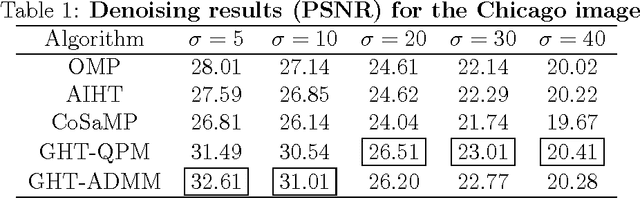

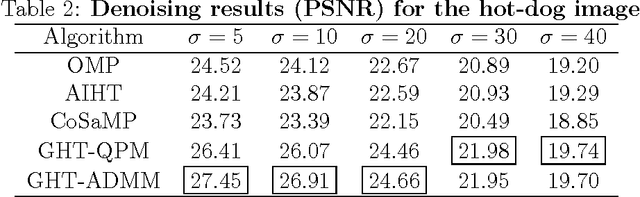
Sparse coding of images is traditionally done by cutting them into small patches and representing each patch individually over some dictionary given a pre-determined number of nonzero coefficients to use for each patch. In lack of a way to effectively distribute a total number (or global budget) of nonzero coefficients across all patches, current sparse recovery algorithms distribute the global budget equally across all patches despite the wide range of differences in structural complexity among them. In this work we propose a new framework for joint sparse representation and recovery of all image patches simultaneously. We also present two novel global hard thresholding algorithms, based on the notion of variable splitting, for solving the joint sparse model. Experimentation using both synthetic and real data shows effectiveness of the proposed framework for sparse image representation and denoising tasks. Additionally, time complexity analysis of the proposed algorithms indicate high scalability of both algorithms, making them favorable to use on large megapixel images.
Deep Depth Prior for Multi-View Stereo
Jan 21, 2020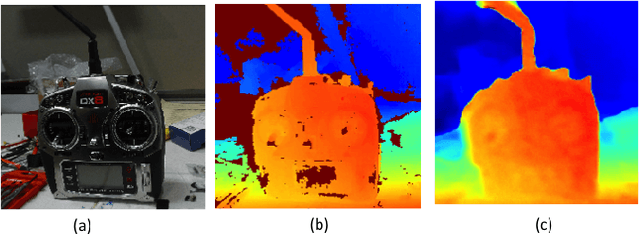
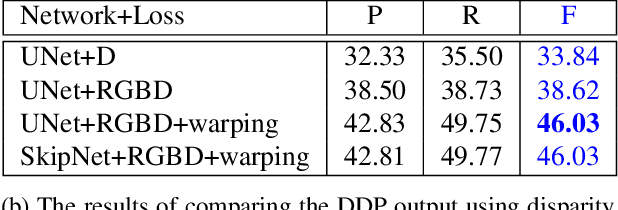
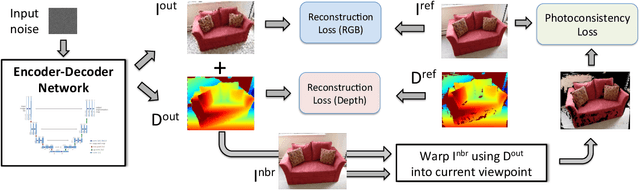
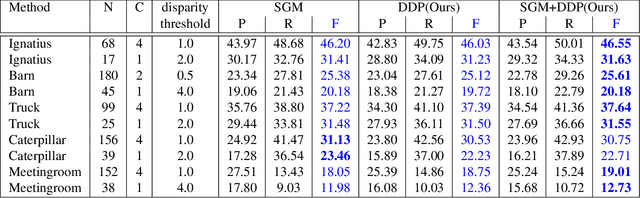
It was recently shown that the structure of convolutional neural networks induces a strong prior favoring natural color images, a phenomena referred to as a deep image prior (DIP), which can be an effective regularizer in inverse problems such as image denoising, inpainting etc. In this paper, we investigate a similar idea for depth images, which we call a deep depth prior. Specifically, given a color image and a noisy and incomplete target depth map from the same viewpoint, we optimize a randomly initialized CNN model to reconstruct an RGB-D image where the depth channel gets restored by virtue of using the network structure as a prior. We propose using deep depth priors for refining and inpainting noisy depth maps within a multi-view stereo pipeline. We optimize the network parameters to minimize two losses 1) a RGB-D reconstruction loss based on the noisy depth map and 2) a multi-view photoconsistency-based loss, which is computed using images from a geometrically calibrated camera from nearby viewpoints. Our quantitative and qualitative evaluation shows that our refined depth maps are more accurate and complete, and after fusion, produces dense 3D models of higher quality.
Weakly Supervised Deep Nuclei Segmentation Using Partial Points Annotation in Histopathology Images
Jul 10, 2020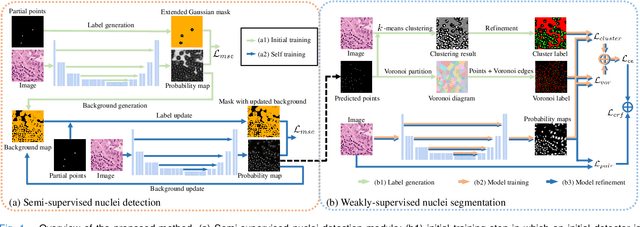
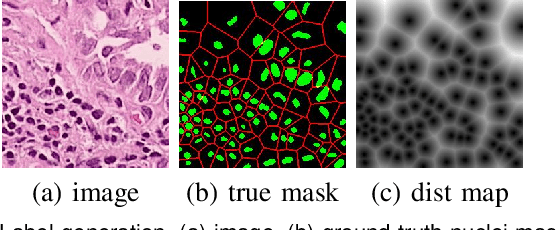
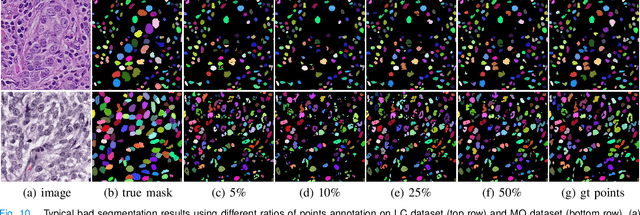
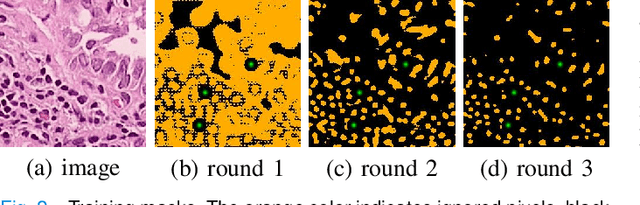

Nuclei segmentation is a fundamental task in histopathology image analysis. Typically, such segmentation tasks require significant effort to manually generate accurate pixel-wise annotations for fully supervised training. To alleviate such tedious and manual effort, in this paper we propose a novel weakly supervised segmentation framework based on partial points annotation, i.e., only a small portion of nuclei locations in each image are labeled. The framework consists of two learning stages. In the first stage, we design a semi-supervised strategy to learn a detection model from partially labeled nuclei locations. Specifically, an extended Gaussian mask is designed to train an initial model with partially labeled data. Then, selftraining with background propagation is proposed to make use of the unlabeled regions to boost nuclei detection and suppress false positives. In the second stage, a segmentation model is trained from the detected nuclei locations in a weakly-supervised fashion. Two types of coarse labels with complementary information are derived from the detected points and are then utilized to train a deep neural network. The fully-connected conditional random field loss is utilized in training to further refine the model without introducing extra computational complexity during inference. The proposed method is extensively evaluated on two nuclei segmentation datasets. The experimental results demonstrate that our method can achieve competitive performance compared to the fully supervised counterpart and the state-of-the-art methods while requiring significantly less annotation effort.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge