"Image": models, code, and papers
Single Frame Deblurring with Laplacian Filters
Sep 17, 2020
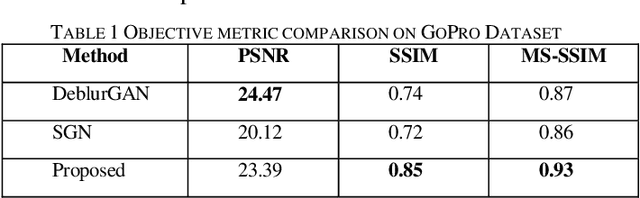
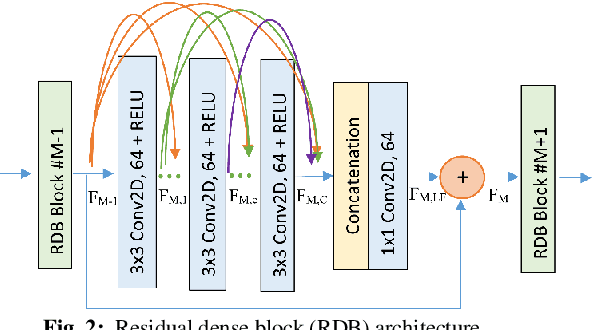

Blind single image deblurring has been a challenge over many decades due to the ill-posed nature of the problem. In this paper, we propose a single-frame blind deblurring solution with the aid of Laplacian filters. Utilized Residual Dense Network has proven its strengths in superresolution task, thus we selected it as a baseline architecture. We evaluated the proposed solution with state-of-art DNN methods on a benchmark dataset. The proposed method shows significant improvement in image quality measured objectively and subjectively.
Cycle-Dehaze: Enhanced CycleGAN for Single Image Dehazing
May 14, 2018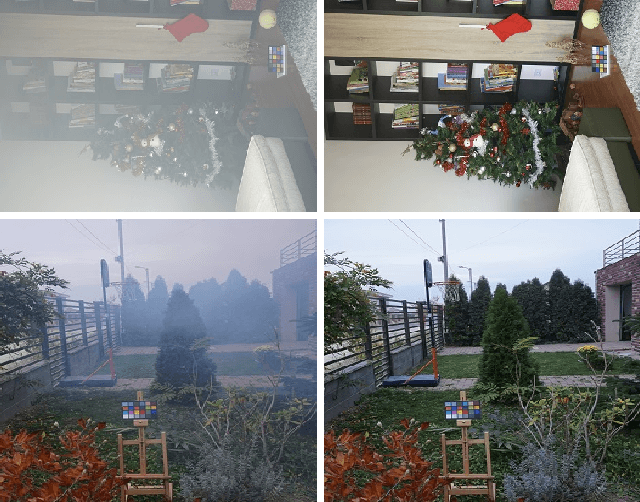

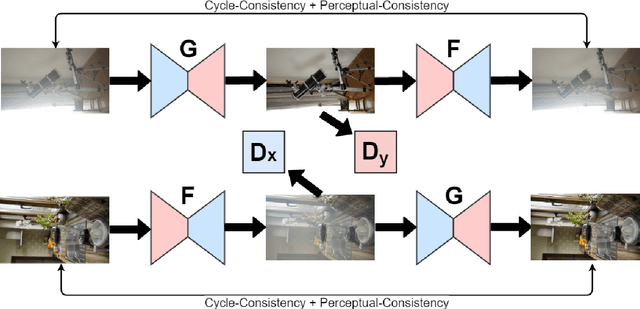
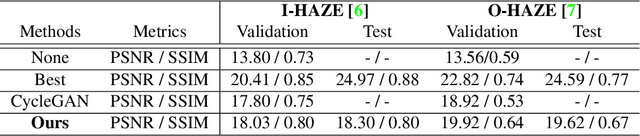
In this paper, we present an end-to-end network, called Cycle-Dehaze, for single image dehazing problem, which does not require pairs of hazy and corresponding ground truth images for training. That is, we train the network by feeding clean and hazy images in an unpaired manner. Moreover, the proposed approach does not rely on estimation of the atmospheric scattering model parameters. Our method enhances CycleGAN formulation by combining cycle-consistency and perceptual losses in order to improve the quality of textural information recovery and generate visually better haze-free images. Typically, deep learning models for dehazing take low resolution images as input and produce low resolution outputs. However, in the NTIRE 2018 challenge on single image dehazing, high resolution images were provided. Therefore, we apply bicubic downscaling. After obtaining low-resolution outputs from the network, we utilize the Laplacian pyramid to upscale the output images to the original resolution. We conduct experiments on NYU-Depth, I-HAZE, and O-HAZE datasets. Extensive experiments demonstrate that the proposed approach improves CycleGAN method both quantitatively and qualitatively.
Inspecting state of the art performance and NLP metrics in image-based medical report generation
Nov 21, 2020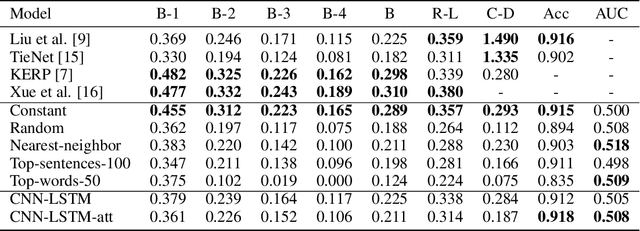
Several deep learning architectures have been proposed over the last years to deal with the problem of generating a written report given an imaging exam as input. Most works evaluate the generated reports using standard Natural Language Processing (NLP) metrics (e.g. BLEU, ROUGE), reporting significant progress. In this article, we contrast this progress by comparing state of the art (SOTA) models against weak baselines. We show that simple and even naive approaches yield near SOTA performance on most traditional NLP metrics. We conclude that evaluation methods in this task should be further studied towards correctly measuring clinical accuracy, ideally involving physicians to contribute to this end.
Demystifying Batch Normalization in ReLU Networks: Equivalent Convex Optimization Models and Implicit Regularization
Mar 02, 2021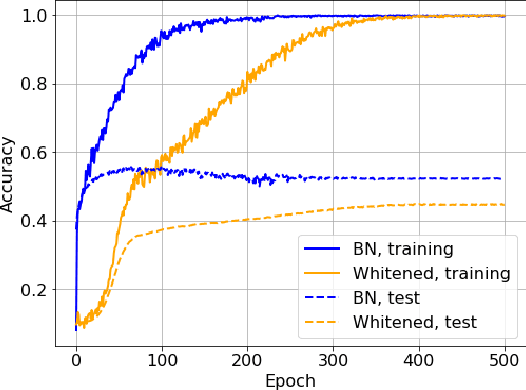
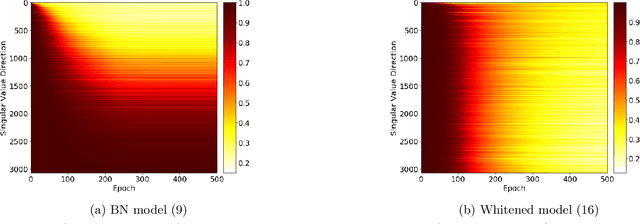
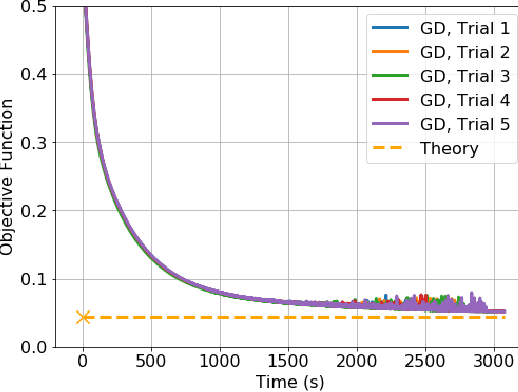
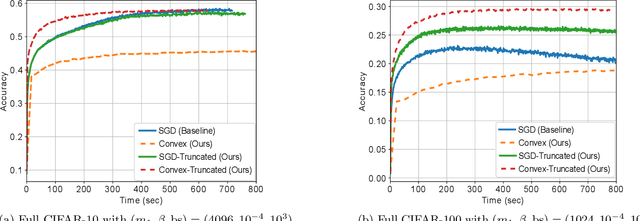
Batch Normalization (BN) is a commonly used technique to accelerate and stabilize training of deep neural networks. Despite its empirical success, a full theoretical understanding of BN is yet to be developed. In this work, we analyze BN through the lens of convex optimization. We introduce an analytic framework based on convex duality to obtain exact convex representations of weight-decay regularized ReLU networks with BN, which can be trained in polynomial-time. Our analyses also show that optimal layer weights can be obtained as simple closed-form formulas in the high-dimensional and/or overparameterized regimes. Furthermore, we find that Gradient Descent provides an algorithmic bias effect on the standard non-convex BN network, and we design an approach to explicitly encode this implicit regularization into the convex objective. Experiments with CIFAR image classification highlight the effectiveness of this explicit regularization for mimicking and substantially improving the performance of standard BN networks.
Human 3D keypoints via spatial uncertainty modeling
Dec 18, 2020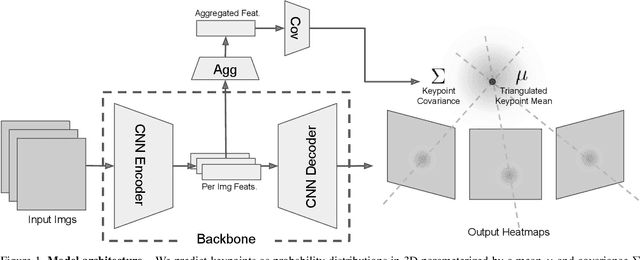
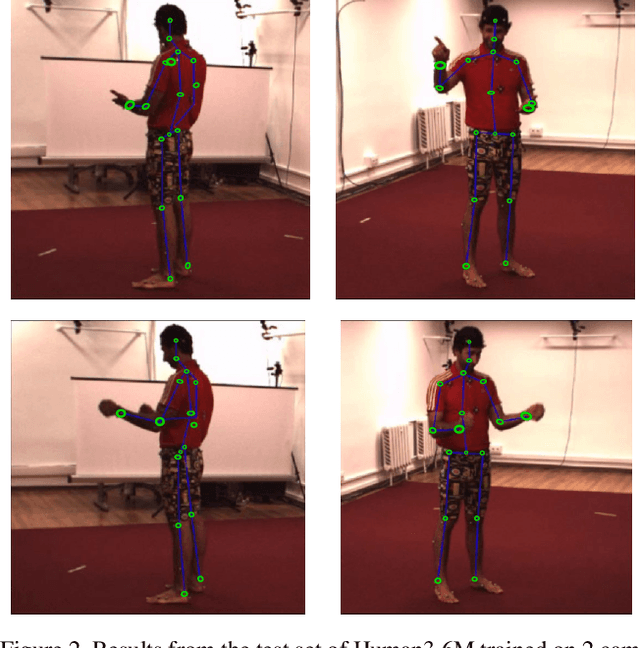
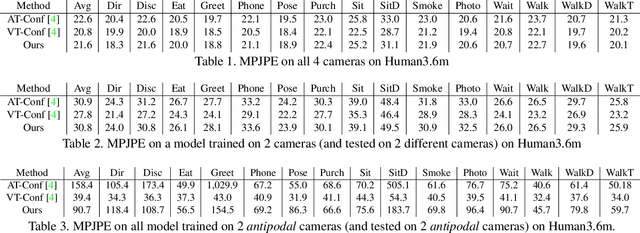

We introduce a technique for 3D human keypoint estimation that directly models the notion of spatial uncertainty of a keypoint. Our technique employs a principled approach to modelling spatial uncertainty inspired from techniques in robust statistics. Furthermore, our pipeline requires no 3D ground truth labels, relying instead on (possibly noisy) 2D image-level keypoints. Our method achieves near state-of-the-art performance on Human3.6m while being efficient to evaluate and straightforward to
Fast Adaptation with Linearized Neural Networks
Mar 02, 2021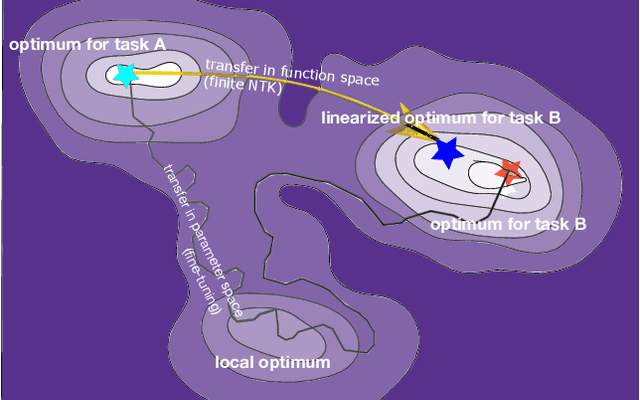

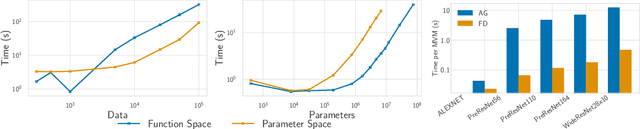
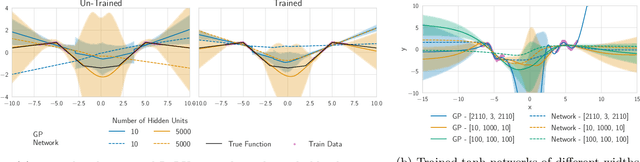
The inductive biases of trained neural networks are difficult to understand and, consequently, to adapt to new settings. We study the inductive biases of linearizations of neural networks, which we show to be surprisingly good summaries of the full network functions. Inspired by this finding, we propose a technique for embedding these inductive biases into Gaussian processes through a kernel designed from the Jacobian of the network. In this setting, domain adaptation takes the form of interpretable posterior inference, with accompanying uncertainty estimation. This inference is analytic and free of local optima issues found in standard techniques such as fine-tuning neural network weights to a new task. We develop significant computational speed-ups based on matrix multiplies, including a novel implementation for scalable Fisher vector products. Our experiments on both image classification and regression demonstrate the promise and convenience of this framework for transfer learning, compared to neural network fine-tuning. Code is available at https://github.com/amzn/xfer/tree/master/finite_ntk.
Benchmarking Deep Learning Classifiers: Beyond Accuracy
Mar 02, 2021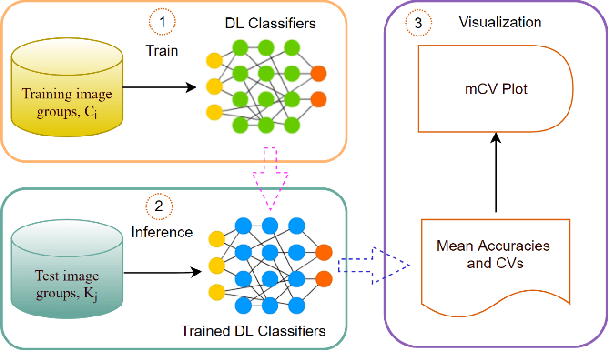
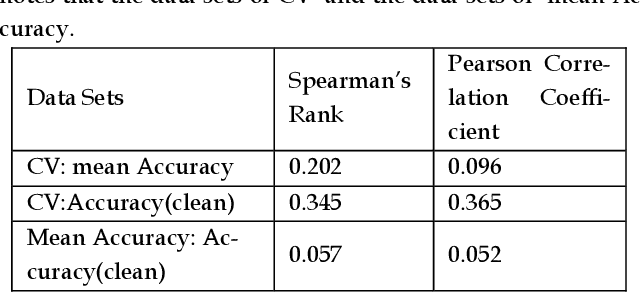
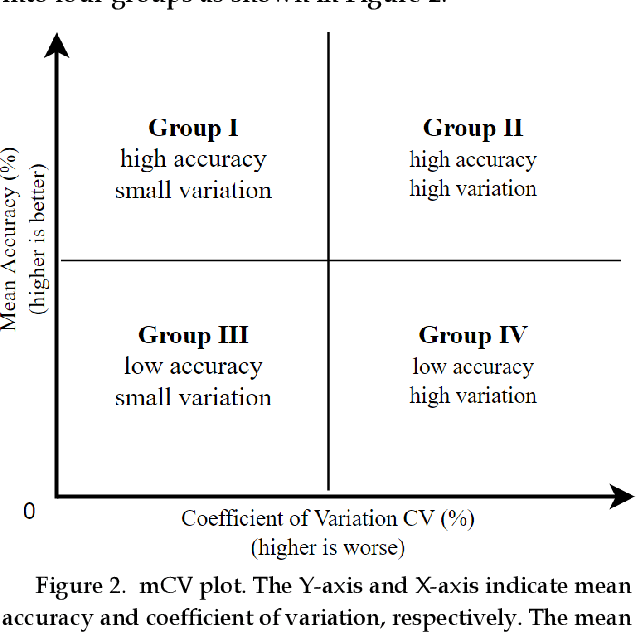
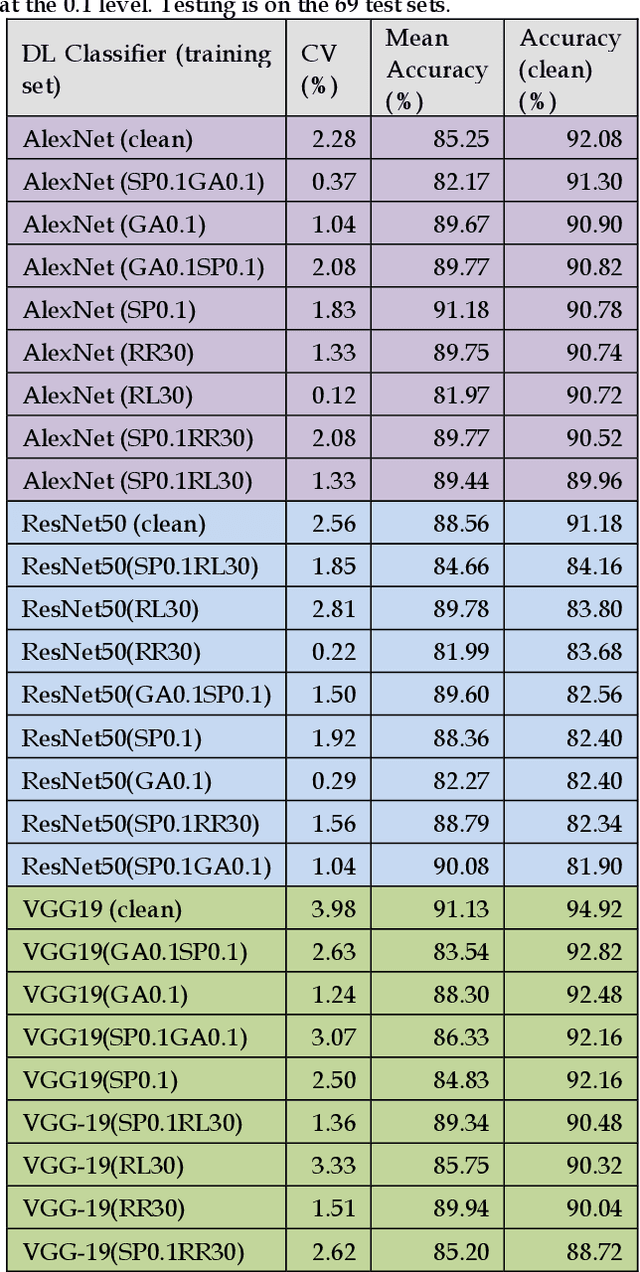
Previous research evaluating deep learning (DL) classifiers has often used top-1/top-5 accuracy. However, the accuracy of DL classifiers is unstable in that it often changes significantly when retested on imperfect or adversarial images. This paper adds to the small but fundamental body of work on benchmarking the robustness of DL classifiers on imperfect images by proposing a two-dimensional metric, consisting of mean accuracy and coefficient of variation, to measure the robustness of DL classifiers. Spearman's rank correlation coefficient and Pearson's correlation coefficient are used and their independence evaluated. A statistical plot we call mCV is presented which aims to help visualize the robustness of the performance of DL classifiers across varying amounts of imperfection in tested images. Finally, we demonstrate that defective images corrupted by two-factor corruption could be used to improve the robustness of DL classifiers. All source codes and related image sets are shared on a website (http://www.animpala.com) to support future research projects.
Fixing the Teacher-Student Knowledge Discrepancy in Distillation
Mar 31, 2021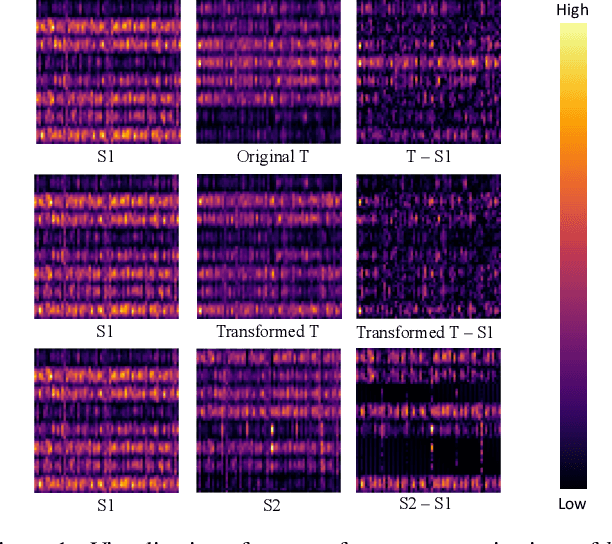

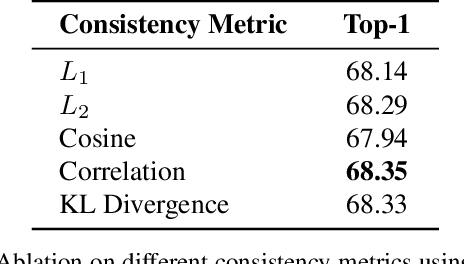
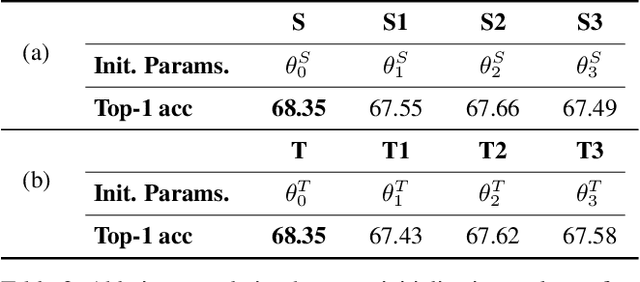
Training a small student network with the guidance of a larger teacher network is an effective way to promote the performance of the student. Despite the different types, the guided knowledge used to distill is always kept unchanged for different teacher and student pairs in previous knowledge distillation methods. However, we find that teacher and student models with different networks or trained from different initialization could have distinct feature representations among different channels. (e.g. the high activated channel for different categories). We name this incongruous representation of channels as teacher-student knowledge discrepancy in the distillation process. Ignoring the knowledge discrepancy problem of teacher and student models will make the learning of student from teacher more difficult. To solve this problem, in this paper, we propose a novel student-dependent distillation method, knowledge consistent distillation, which makes teacher's knowledge more consistent with the student and provides the best suitable knowledge to different student networks for distillation. Extensive experiments on different datasets (CIFAR100, ImageNet, COCO) and tasks (image classification, object detection) reveal the widely existing knowledge discrepancy problem between teachers and students and demonstrate the effectiveness of our proposed method. Our method is very flexible that can be easily combined with other state-of-the-art approaches.
Modelling brain lesion volume in patches with CNN-based Poisson Regression
Nov 26, 2020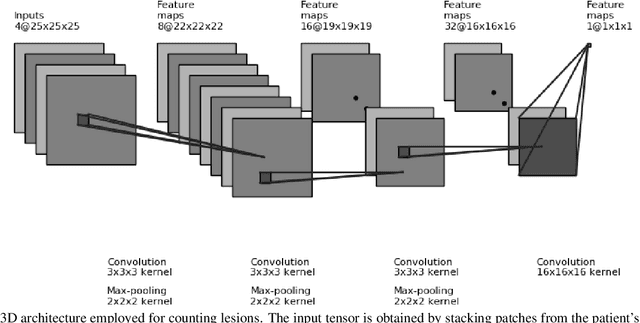
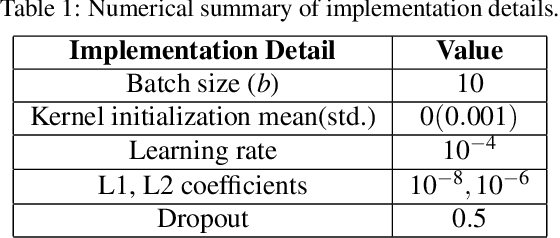
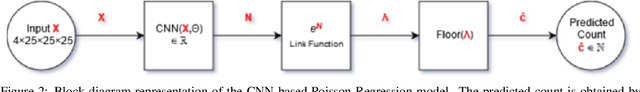
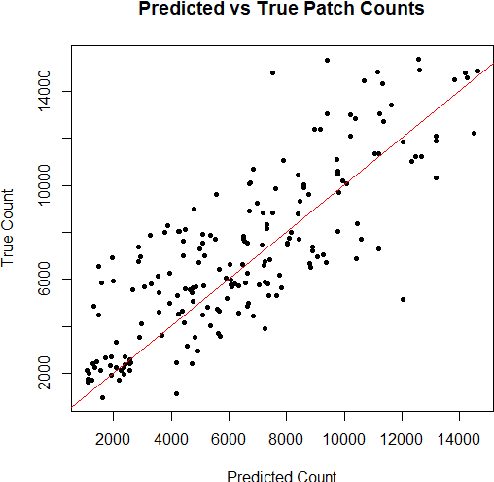
Monitoring the progression of lesions is important for clinical response. Summary statistics such as lesion volume are objective and easy to interpret, which can help clinicians assess lesion growth or decay. CNNs are commonly used in medical image segmentation for their ability to produce useful features within large contexts and their associated efficient iterative patch-based training. Many CNN architectures require hundreds of thousands parameters to yield a good segmentation. In this work, an efficient, computationally inexpensive CNN is implemented to estimate the number of lesion voxels in a predefined patch size from magnetic resonance (MR) images. The output of the CNN is interpreted as the conditional Poisson parameter over the patch, allowing standard mini-batch gradient descent to be employed. The ISLES2015 (SISS) data is used to train and evaluate the model, which by estimating lesion volume from raw features, accurately identified the lesion image with the larger lesion volume for 86% of paired sample patches. An argument for the development and use of estimating lesion volumes to also aid in model selection for segmentation is made.
Convolutional Neural Nets: Foundations, Computations, and New Applications
Jan 13, 2021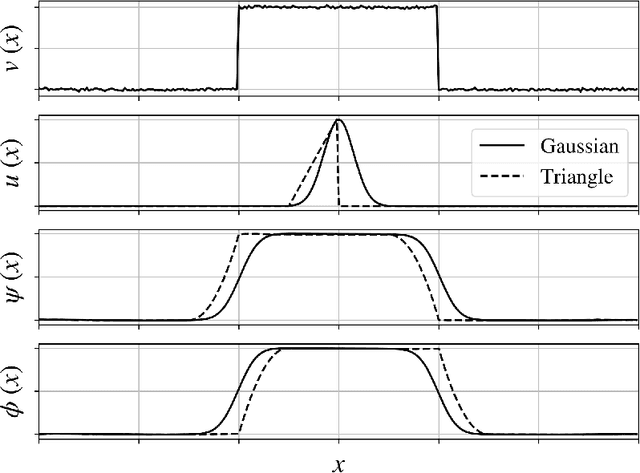
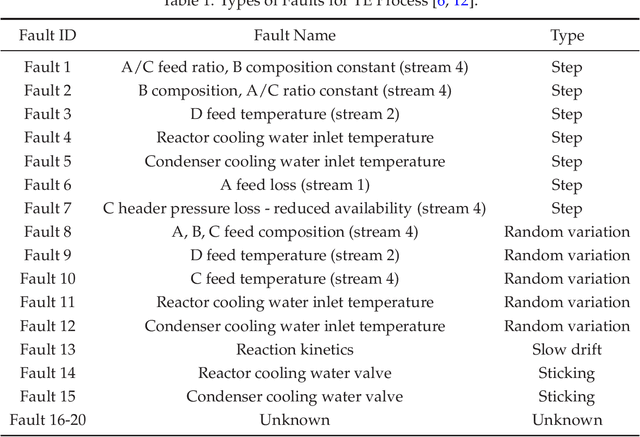

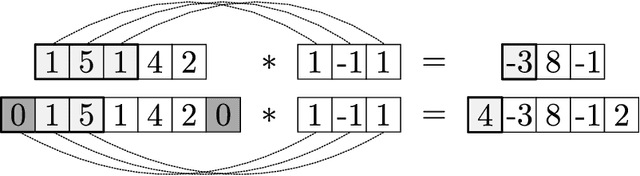
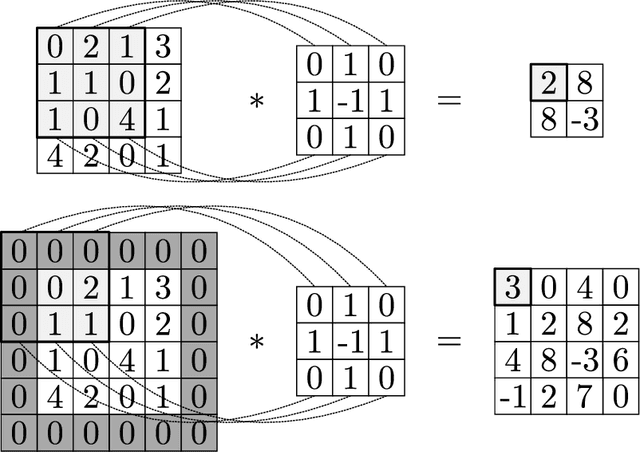
We review mathematical foundations of convolutional neural nets (CNNs) with the goals of: i) highlighting connections with techniques from statistics, signal processing, linear algebra, differential equations, and optimization, ii) demystifying underlying computations, and iii) identifying new types of applications. CNNs are powerful machine learning models that highlight features from grid data to make predictions (regression and classification). The grid data object can be represented as vectors (in 1D), matrices (in 2D), or tensors (in 3D or higher dimensions) and can incorporate multiple channels (thus providing high flexibility in the input data representation). For example, an image can be represented as a 2D grid data object that contains red, green, and blue (RBG) channels (each channel is a 2D matrix). Similarly, a video can be represented as a 3D grid data object (two spatial dimensions plus time) with RGB channels (each channel is a 3D tensor). CNNs highlight features from the grid data by performing convolution operations with different types of operators. The operators highlight different types of features (e.g., patterns, gradients, geometrical features) and are learned by using optimization techniques. In other words, CNNs seek to identify optimal operators that best map the input data to the output data. A common misconception is that CNNs are only capable of processing image or video data but their application scope is much wider; specifically, datasets encountered in diverse applications can be expressed as grid data. Here, we show how to apply CNNs to new types of applications such as optimal control, flow cytometry, multivariate process monitoring, and molecular simulations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge