"Image": models, code, and papers
Spatial Moment Pooling Improves Neural Image Assessment
Sep 29, 2022
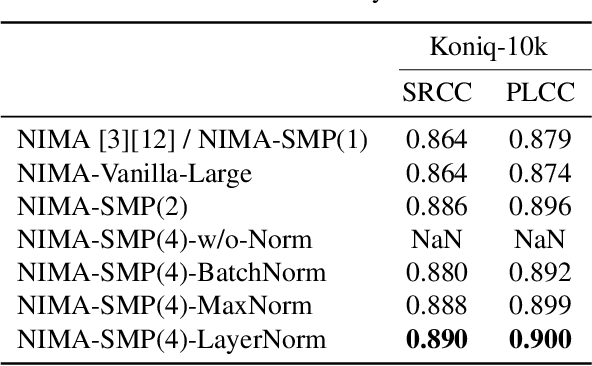
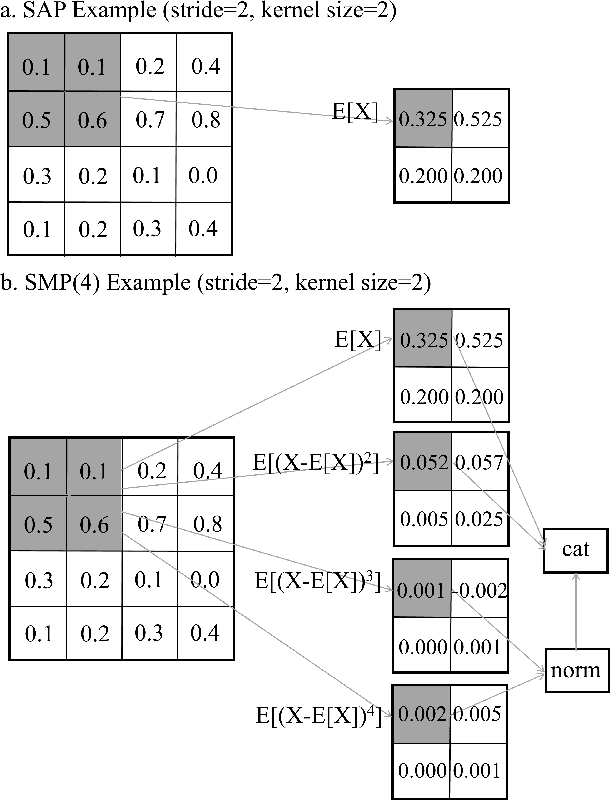
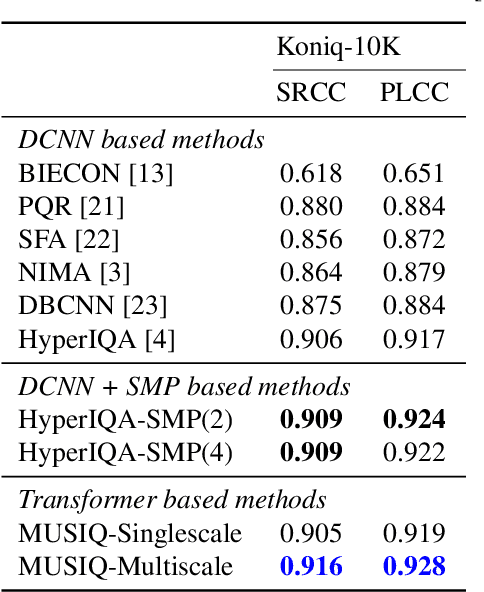
In recent years, there has been widespread attention drawn to convolutional neural network (CNN) based blind image quality assessment (IQA). A large number of works start by extracting deep features from CNN. Then, those features are processed through spatial average pooling (SAP) and fully connected layers to predict quality. Inspired by full reference IQA and texture features, in this paper, we extend SAP ($1^{st}$ moment) into spatial moment pooling (SMP) by incorporating higher order moments (such as variance, skewness). Moreover, we provide learning friendly normalization to circumvent numerical issue when computing gradients of higher moments. Experimental results suggest that simply upgrading SAP to SMP significantly enhances CNN-based blind IQA methods and achieves state of the art performance.
Directed Diffusion: Direct Control of Object Placement through Attention Guidance
Feb 25, 2023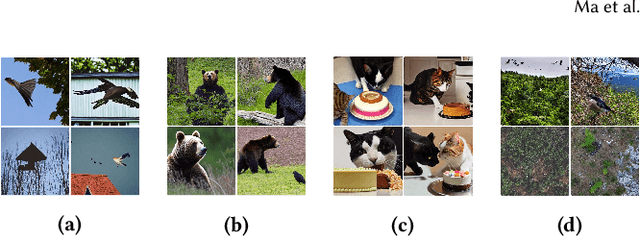
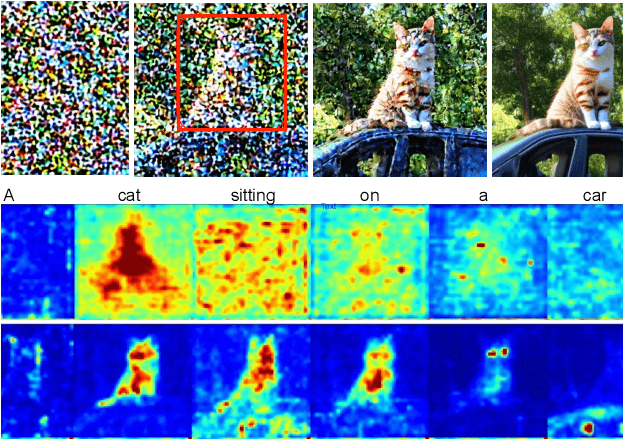

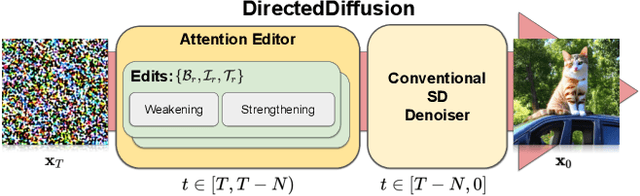
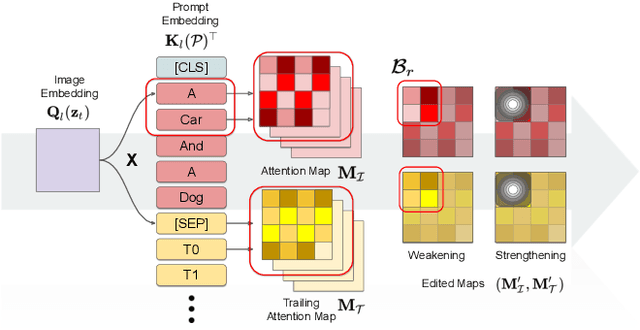
Text-guided diffusion models such as DALLE-2, IMAGEN, and Stable Diffusion are able to generate an effectively endless variety of images given only a short text prompt describing the desired image content. In many cases the images are very high quality as well. However, these models often struggle to compose scenes containing several key objects such as characters in specified positional relationships. Unfortunately, this capability to ``direct'' the placement of characters and objects both within and across images is crucial in storytelling, as recognized in the literature on film and animation theory. In this work we take a particularly straightforward approach to providing the needed direction, by injecting ``activation'' at desired positions in the cross-attention maps corresponding to the objects under control, while attenuating the remainder of the map. The resulting approach is a step toward generalizing the applicability of text-guided diffusion models beyond single images to collections of related images, as in storybooks. To the best of our knowledge, our Directed Diffusion method is the first diffusion technique that provides positional control over multiple objects, while making use of an existing pre-trained model and maintaining a coherent blend between the positioned objects and the background. Moreover, it requires only a few lines to implement.
A Surrogate-Assisted Highly Cooperative Coevolutionary Algorithm for Hyperparameter Optimization in Deep Convolutional Neural Network
Feb 25, 2023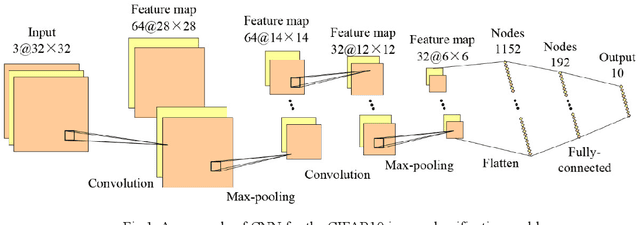
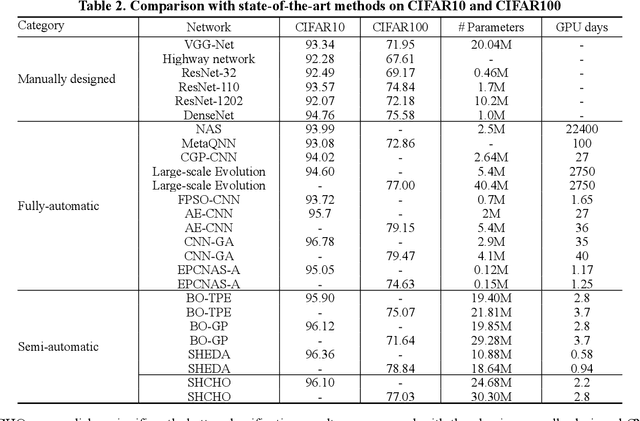
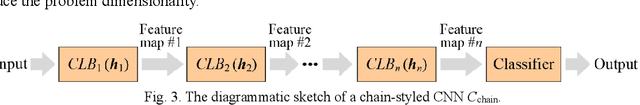
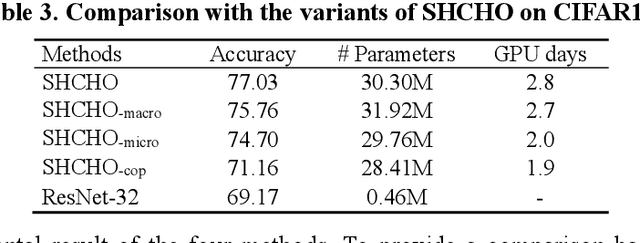
Convolutional neural networks (CNNs) have gained remarkable success in recent years. However, their performance highly relies on the architecture hyperparameters, and finding proper hyperparameters for a deep CNN is a challenging optimization problem owing to its high-dimensional and computationally expensive characteristics. Given these difficulties, this study proposes a surrogate-assisted highly cooperative hyperparameter optimization (SHCHO) algorithm for chain-styled CNNs. To narrow the large search space, SHCHO first decomposes the whole CNN into several overlapping sub-CNNs in accordance with the overlapping hyperparameter interaction structure and then cooperatively optimizes these hyperparameter subsets. Two cooperation mechanisms are designed during this process. One coordinates all the sub-CNNs to reproduce the information flow in the whole CNN and achieve macro cooperation among them, and the other tackles the overlapping components by simultaneously considering the involved two sub-CNNs and facilitates micro cooperation between them. As a result, a proper hyperparameter configuration can be effectively located for the whole CNN. Besides, SHCHO also employs the well-performing surrogate technique to assist in the hyperparameter optimization of each sub-CNN, thereby greatly reducing the expensive computational cost. Extensive experimental results on two widely-used image classification datasets indicate that SHCHO can significantly improve the performance of CNNs.
Deep Anatomical Federated Network (Dafne): an open client/server framework for the continuous collaborative improvement of deep-learning-based medical image segmentation
Feb 14, 2023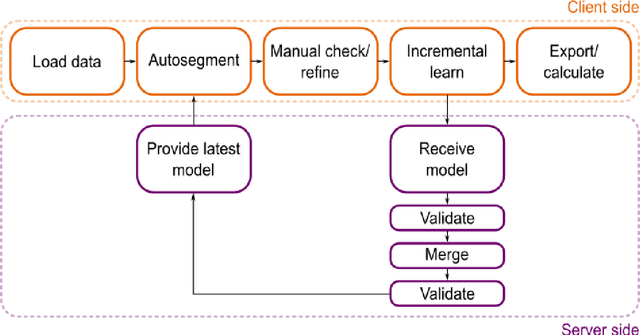
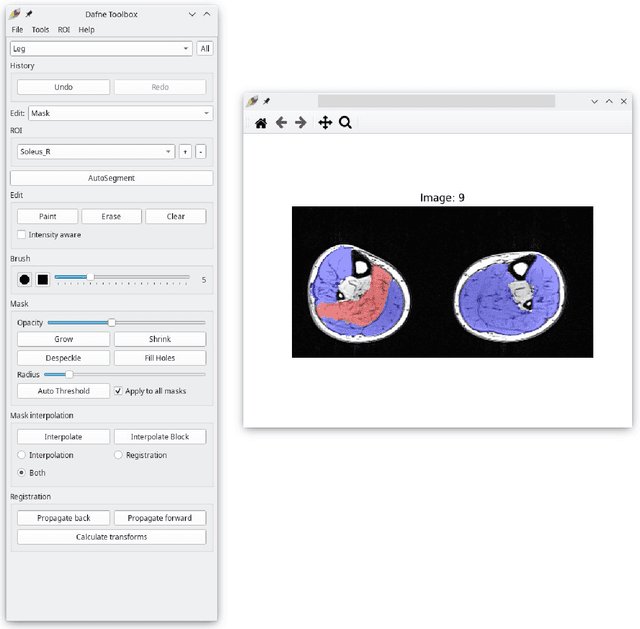
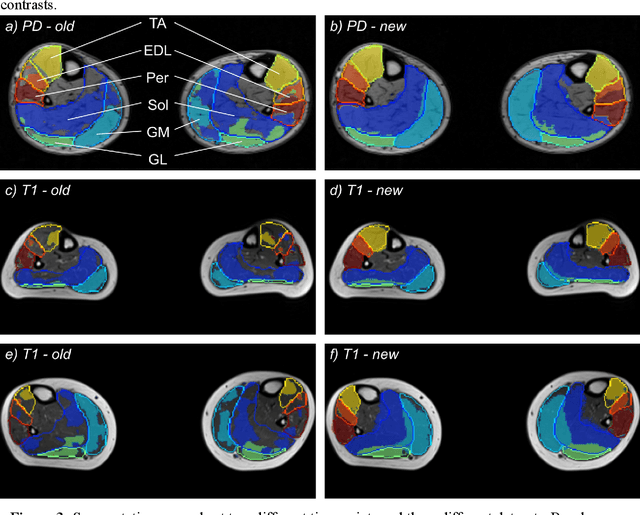
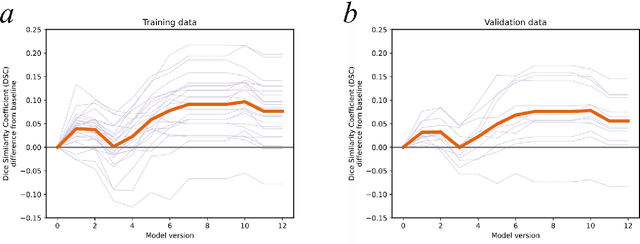
Semantic segmentation is a crucial step to extract quantitative information from medical (and, specifically, radiological) images to aid the diagnostic process, clinical follow-up. and to generate biomarkers for clinical research. In recent years, machine learning algorithms have become the primary tool for this task. However, its real-world performance is heavily reliant on the comprehensiveness of training data. Dafne is the first decentralized, collaborative solution that implements continuously evolving deep learning models exploiting the collective knowledge of the users of the system. In the Dafne workflow, the result of each automated segmentation is refined by the user through an integrated interface, so that the new information is used to continuously expand the training pool via federated incremental learning. The models deployed through Dafne are able to improve their performance over time and to generalize to data types not seen in the training sets, thus becoming a viable and practical solution for real-life medical segmentation tasks.
Fact or Artifact? Revise Layer-wise Relevance Propagation on various ANN Architectures
Feb 23, 2023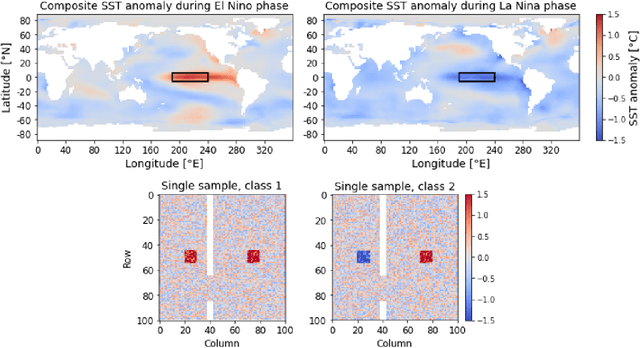
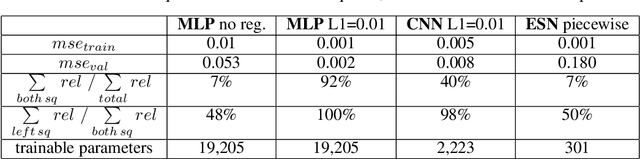
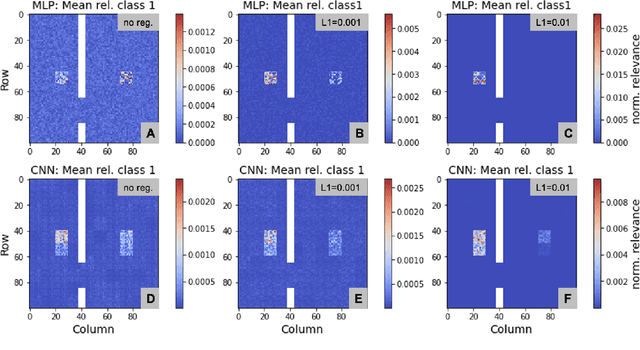
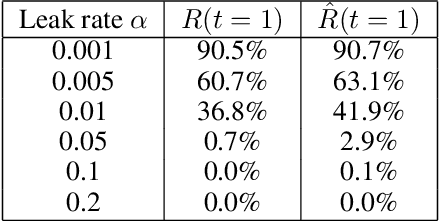
Layer-wise relevance propagation (LRP) is a widely used and powerful technique to reveal insights into various artificial neural network (ANN) architectures. LRP is often used in the context of image classification. The aim is to understand, which parts of the input sample have highest relevance and hence most influence on the model prediction. Relevance can be traced back through the network to attribute a certain score to each input pixel. Relevance scores are then combined and displayed as heat maps and give humans an intuitive visual understanding of classification models. Opening the black box to understand the classification engine in great detail is essential for domain experts to gain trust in ANN models. However, there are pitfalls in terms of model-inherent artifacts included in the obtained relevance maps, that can easily be missed. But for a valid interpretation, these artifacts must not be ignored. Here, we apply and revise LRP on various ANN architectures trained as classifiers on geospatial and synthetic data. Depending on the network architecture, we show techniques to control model focus and give guidance to improve the quality of obtained relevance maps to separate facts from artifacts.
Out-of-Domain Robustness via Targeted Augmentations
Feb 23, 2023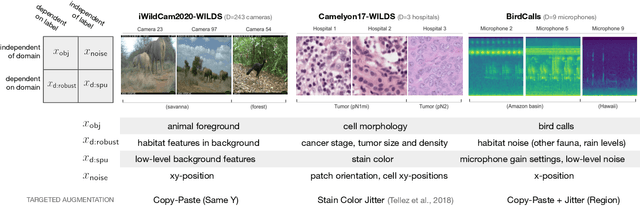
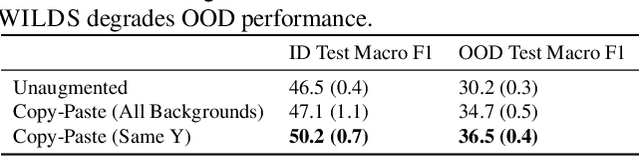

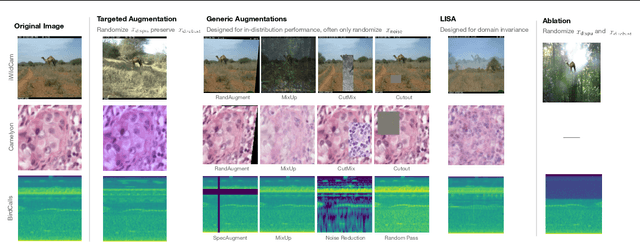
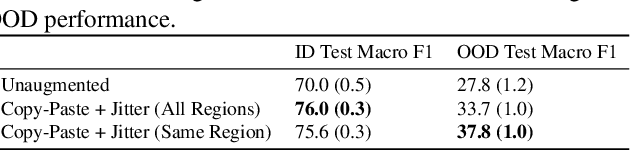
Models trained on one set of domains often suffer performance drops on unseen domains, e.g., when wildlife monitoring models are deployed in new camera locations. In this work, we study principles for designing data augmentations for out-of-domain (OOD) generalization. In particular, we focus on real-world scenarios in which some domain-dependent features are robust, i.e., some features that vary across domains are predictive OOD. For example, in the wildlife monitoring application above, image backgrounds vary across camera locations but indicate habitat type, which helps predict the species of photographed animals. Motivated by theoretical analysis on a linear setting, we propose targeted augmentations, which selectively randomize spurious domain-dependent features while preserving robust ones. We prove that targeted augmentations improve OOD performance, allowing models to generalize better with fewer domains. In contrast, existing approaches such as generic augmentations, which fail to randomize domain-dependent features, and domain-invariant augmentations, which randomize all domain-dependent features, both perform poorly OOD. In experiments on three real-world datasets, we show that targeted augmentations set new states-of-the-art for OOD performance by 3.2-15.2%.
EquiPocket: an E(3)-Equivariant Geometric Graph Neural Network for Ligand Binding Site Prediction
Feb 23, 2023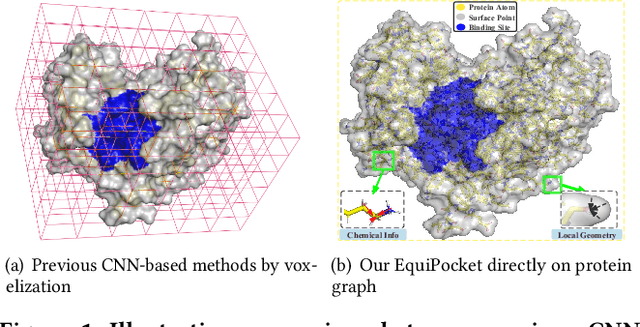
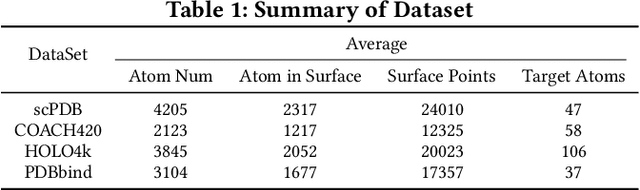
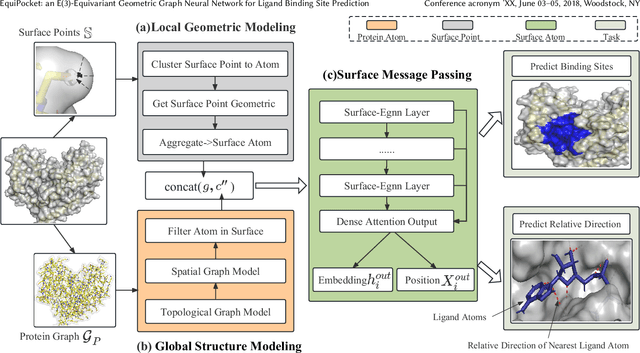
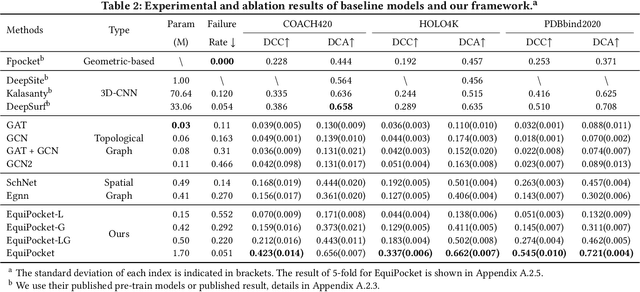
Predicting the binding sites of the target proteins plays a fundamental role in drug discovery. Most existing deep-learning methods consider a protein as a 3D image by spatially clustering its atoms into voxels and then feed the voxelized protein into a 3D CNN for prediction. However, the CNN-based methods encounter several critical issues: 1) defective in representing irregular protein structures; 2) sensitive to rotations; 3) insufficient to characterize the protein surface; 4) unaware of data distribution shift. To address the above issues, this work proposes EquiPocket, an E(3)-equivariant Graph Neural Network (GNN) for binding site prediction. In particular, EquiPocket consists of three modules: the first one to extract local geometric information for each surface atom, the second one to model both the chemical and spatial structure of the protein, and the last one to capture the geometry of the surface via equivariant message passing over the surface atoms. We further propose a dense attention output layer to better alleviate the data distribution shift effect incurred by the variable protein size. Extensive experiments on several representative benchmarks demonstrate the superiority of our framework to the state-of-the-art methods.
Language Model Crossover: Variation through Few-Shot Prompting
Feb 23, 2023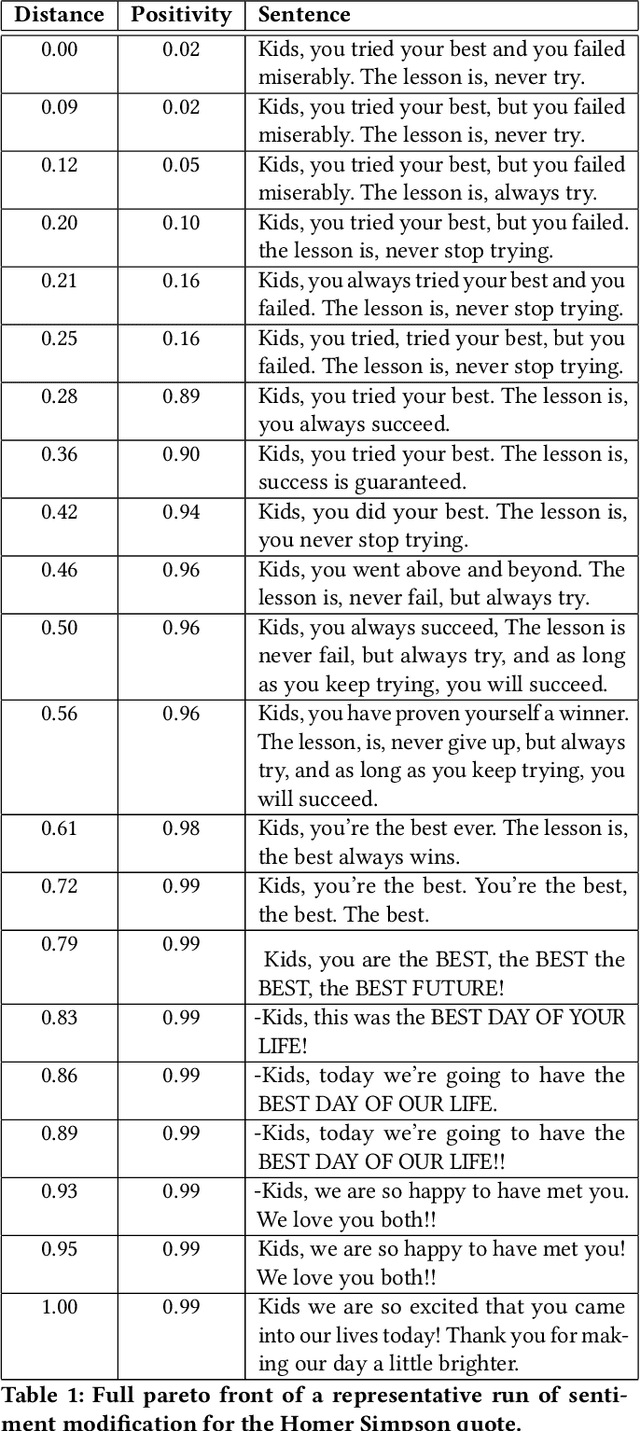
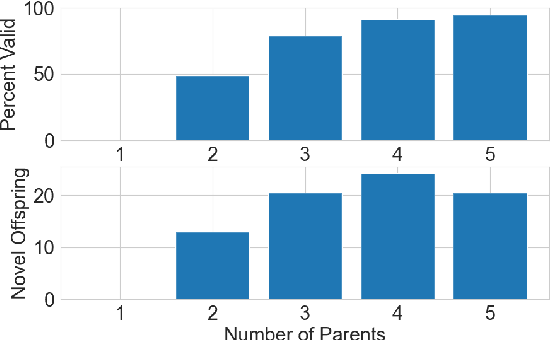
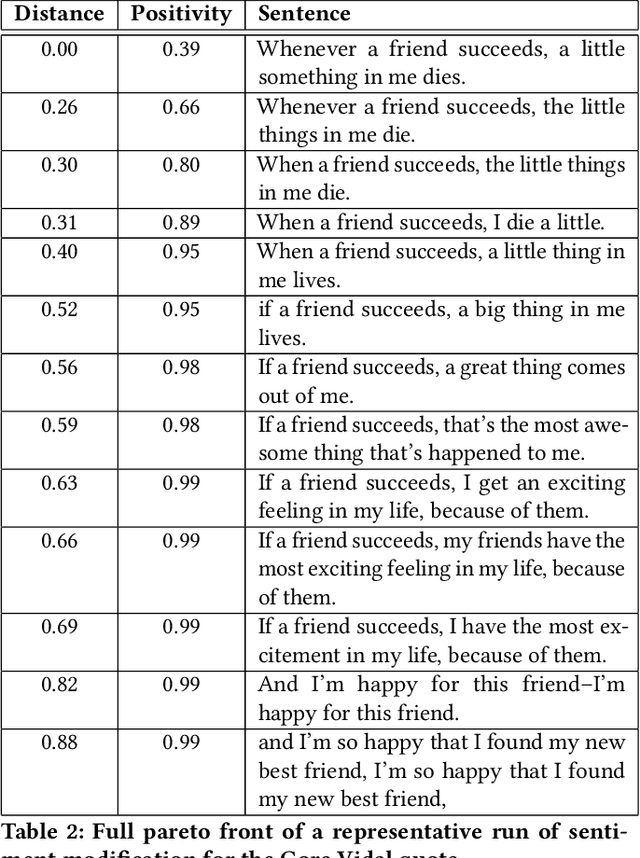
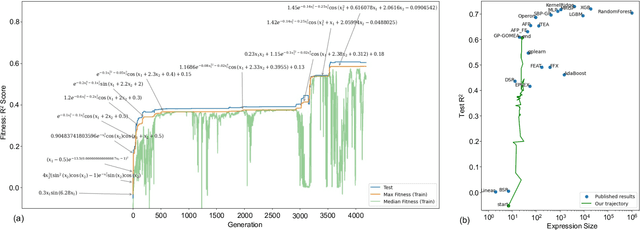
This paper pursues the insight that language models naturally enable an intelligent variation operator similar in spirit to evolutionary crossover. In particular, language models of sufficient scale demonstrate in-context learning, i.e. they can learn from associations between a small number of input patterns to generate outputs incorporating such associations (also called few-shot prompting). This ability can be leveraged to form a simple but powerful variation operator, i.e. to prompt a language model with a few text-based genotypes (such as code, plain-text sentences, or equations), and to parse its corresponding output as those genotypes' offspring. The promise of such language model crossover (which is simple to implement and can leverage many different open-source language models) is that it enables a simple mechanism to evolve semantically-rich text representations (with few domain-specific tweaks), and naturally benefits from current progress in language models. Experiments in this paper highlight the versatility of language-model crossover, through evolving binary bit-strings, sentences, equations, text-to-image prompts, and Python code. The conclusion is that language model crossover is a promising method for evolving genomes representable as text.
CHeart: A Conditional Spatio-Temporal Generative Model for Cardiac Anatomy
Jan 30, 2023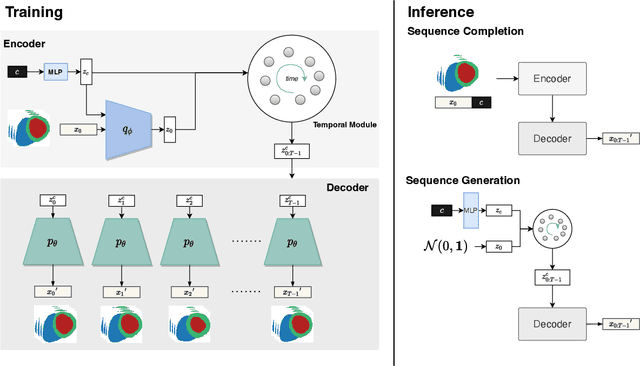
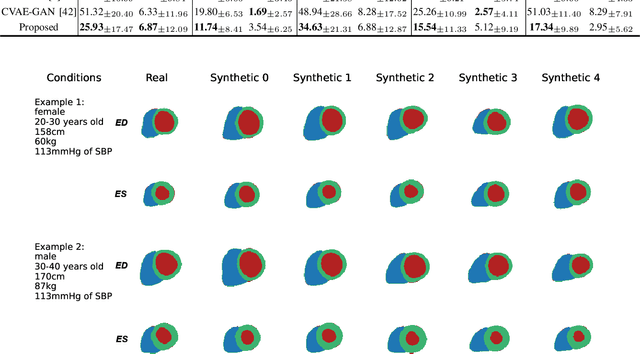
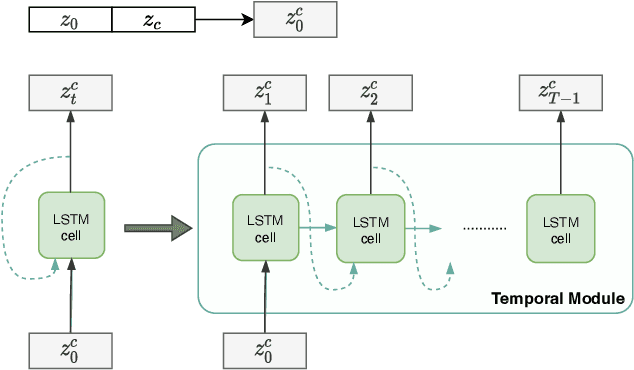
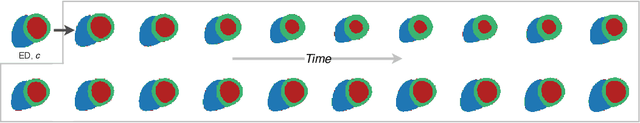

Two key questions in cardiac image analysis are to assess the anatomy and motion of the heart from images; and to understand how they are associated with non-imaging clinical factors such as gender, age and diseases. While the first question can often be addressed by image segmentation and motion tracking algorithms, our capability to model and to answer the second question is still limited. In this work, we propose a novel conditional generative model to describe the 4D spatio-temporal anatomy of the heart and its interaction with non-imaging clinical factors. The clinical factors are integrated as the conditions of the generative modelling, which allows us to investigate how these factors influence the cardiac anatomy. We evaluate the model performance in mainly two tasks, anatomical sequence completion and sequence generation. The model achieves a high performance in anatomical sequence completion, comparable to or outperforming other state-of-the-art generative models. In terms of sequence generation, given clinical conditions, the model can generate realistic synthetic 4D sequential anatomies that share similar distributions with the real data.
Massively Scaling Heteroscedastic Classifiers
Jan 30, 2023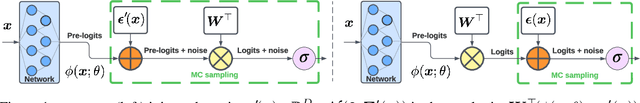

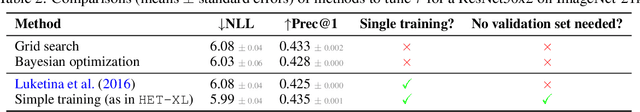
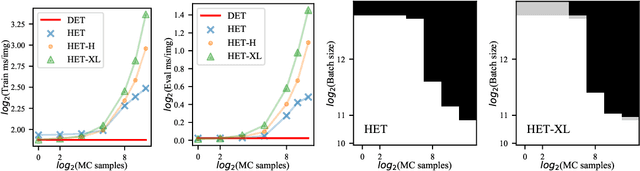
Heteroscedastic classifiers, which learn a multivariate Gaussian distribution over prediction logits, have been shown to perform well on image classification problems with hundreds to thousands of classes. However, compared to standard classifiers, they introduce extra parameters that scale linearly with the number of classes. This makes them infeasible to apply to larger-scale problems. In addition heteroscedastic classifiers introduce a critical temperature hyperparameter which must be tuned. We propose HET-XL, a heteroscedastic classifier whose parameter count when compared to a standard classifier scales independently of the number of classes. In our large-scale settings, we show that we can remove the need to tune the temperature hyperparameter, by directly learning it on the training data. On large image classification datasets with up to 4B images and 30k classes our method requires 14X fewer additional parameters, does not require tuning the temperature on a held-out set and performs consistently better than the baseline heteroscedastic classifier. HET-XL improves ImageNet 0-shot classification in a multimodal contrastive learning setup which can be viewed as a 3.5 billion class classification problem.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge