"Image To Image Translation": models, code, and papers
StarGAN: Unified Generative Adversarial Networks for Multi-Domain Image-to-Image Translation
Sep 21, 2018
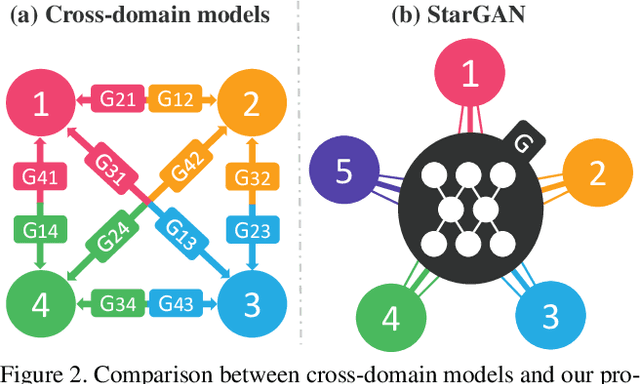
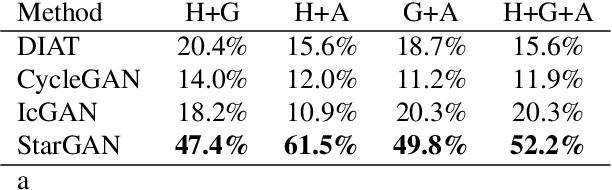
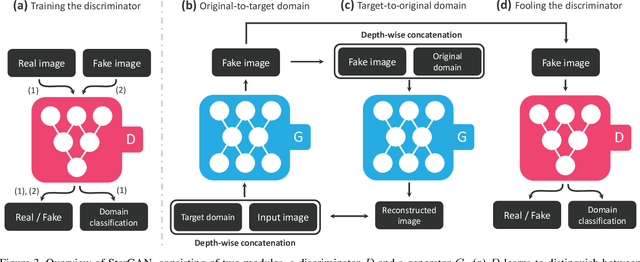
Recent studies have shown remarkable success in image-to-image translation for two domains. However, existing approaches have limited scalability and robustness in handling more than two domains, since different models should be built independently for every pair of image domains. To address this limitation, we propose StarGAN, a novel and scalable approach that can perform image-to-image translations for multiple domains using only a single model. Such a unified model architecture of StarGAN allows simultaneous training of multiple datasets with different domains within a single network. This leads to StarGAN's superior quality of translated images compared to existing models as well as the novel capability of flexibly translating an input image to any desired target domain. We empirically demonstrate the effectiveness of our approach on a facial attribute transfer and a facial expression synthesis tasks.
* Accepted to CVPR 2018 (Oral)
Brainomaly: Unsupervised Neurologic Disease Detection Utilizing Unannotated T1-weighted Brain MR Images
Feb 18, 2023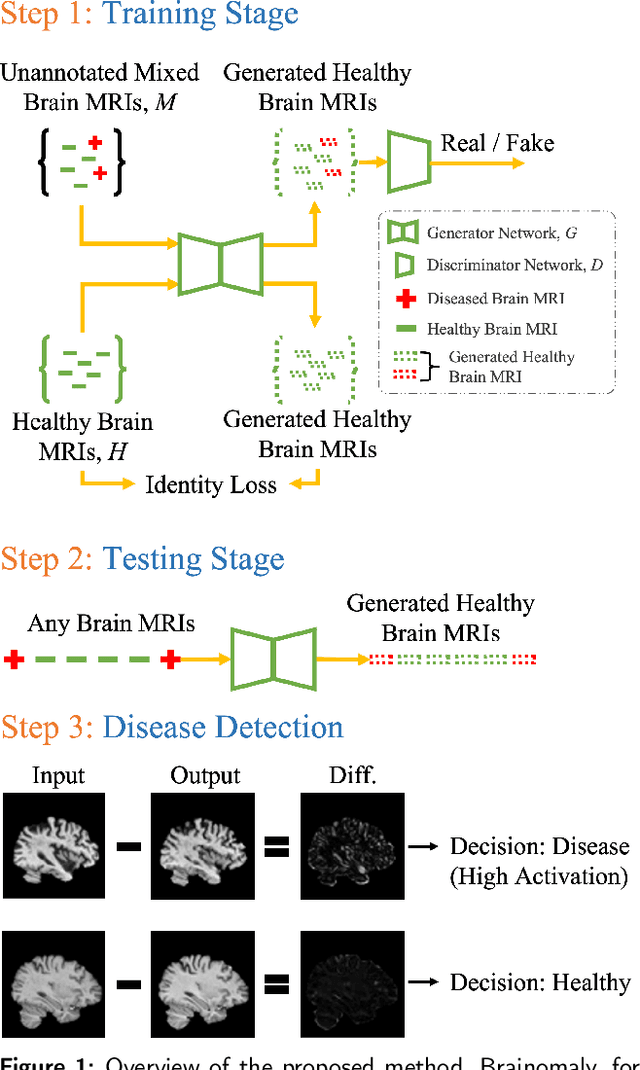
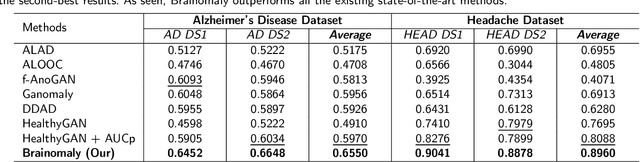
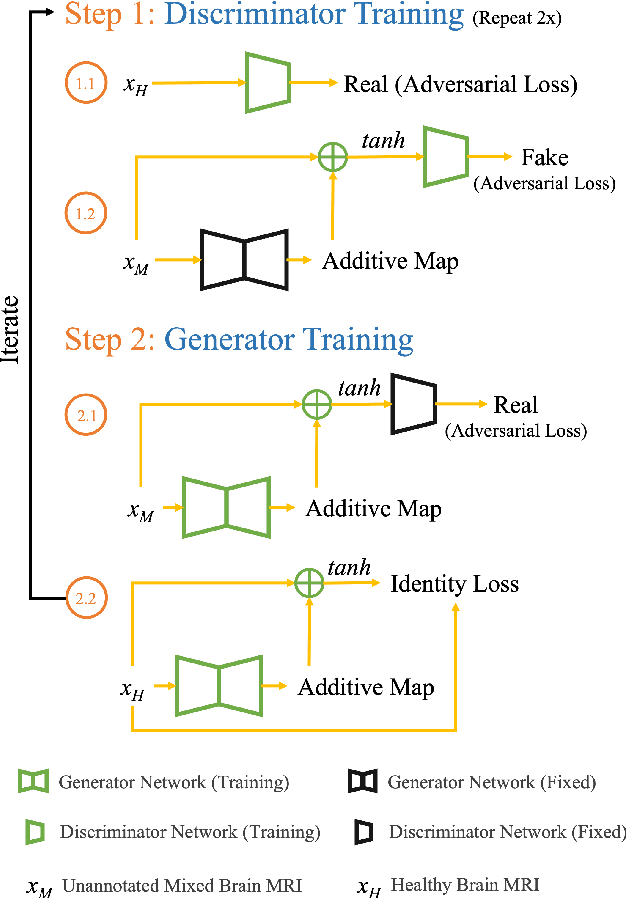
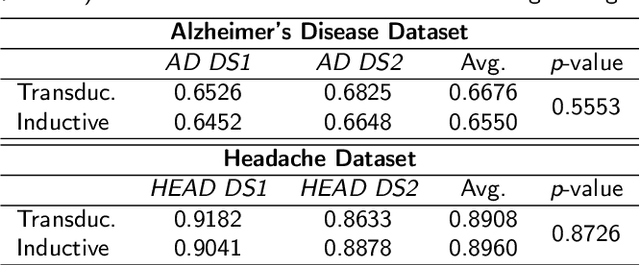
Deep neural networks have revolutionized the field of supervised learning by enabling accurate predictions through learning from large annotated datasets. However, acquiring large annotated medical imaging datasets is a challenging task, especially for rare diseases, due to the high cost, time, and effort required for annotation. In these scenarios, unsupervised disease detection methods, such as anomaly detection, can save significant human effort. A typically used approach for anomaly detection is to learn the images from healthy subjects only, assuming the model will detect the images from diseased subjects as outliers. However, in many real-world scenarios, unannotated datasets with a mix of healthy and diseased individuals are available. Recent studies have shown improvement in unsupervised disease/anomaly detection using such datasets of unannotated images from healthy and diseased individuals compared to datasets that only include images from healthy individuals. A major issue remains unaddressed in these studies, which is selecting the best model for inference from a set of trained models without annotated samples. To address this issue, we propose Brainomaly, a GAN-based image-to-image translation method for neurologic disease detection using unannotated T1-weighted brain MRIs of individuals with neurologic diseases and healthy subjects. Brainomaly is trained to remove the diseased regions from the input brain MRIs and generate MRIs of corresponding healthy brains. Instead of generating the healthy images directly, Brainomaly generates an additive map where each voxel indicates the amount of changes required to make the input image look healthy. In addition, Brainomaly uses a pseudo-AUC metric for inference model selection, which further improves the detection performance. Our Brainomaly outperforms existing state-of-the-art methods by large margins.
Continuous and Diverse Image-to-Image Translation via Signed Attribute Vectors
Nov 03, 2020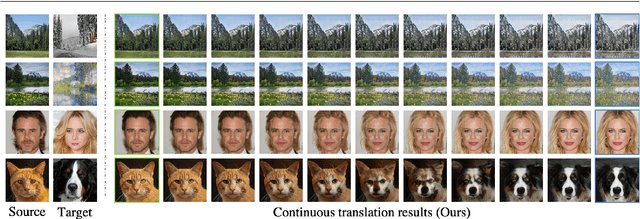

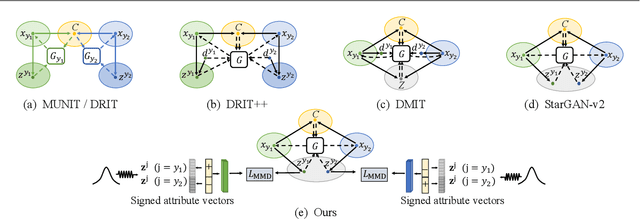
Recent image-to-image (I2I) translation algorithms focus on learning the mapping from a source to a target domain. However, the continuous translation problem that synthesizes intermediate results between the two domains has not been well-studied in the literature. Generating a smooth sequence of intermediate results bridges the gap of two different domains, facilitating the morphing effect across domains. Existing I2I approaches are limited to either intra-domain or deterministic inter-domain continuous translation. In this work, we present an effective signed attribute vector, which enables continuous translation on diverse mapping paths across various domains. In particular, utilizing the sign operation to encode the domain information, we introduce a unified attribute space shared by all domains, thereby allowing the interpolation on attribute vectors of different domains. To enhance the visual quality of continuous translation results, we generate a trajectory between two sign-symmetrical attribute vectors and leverage the domain information of the interpolated results along the trajectory for adversarial training. We evaluate the proposed method on a wide range of I2I translation tasks. Both qualitative and quantitative results demonstrate that the proposed framework generates more high-quality continuous translation results against the state-of-the-art methods.
MISO: Mutual Information Loss with Stochastic Style Representations for Multimodal Image-to-Image Translation
Feb 11, 2019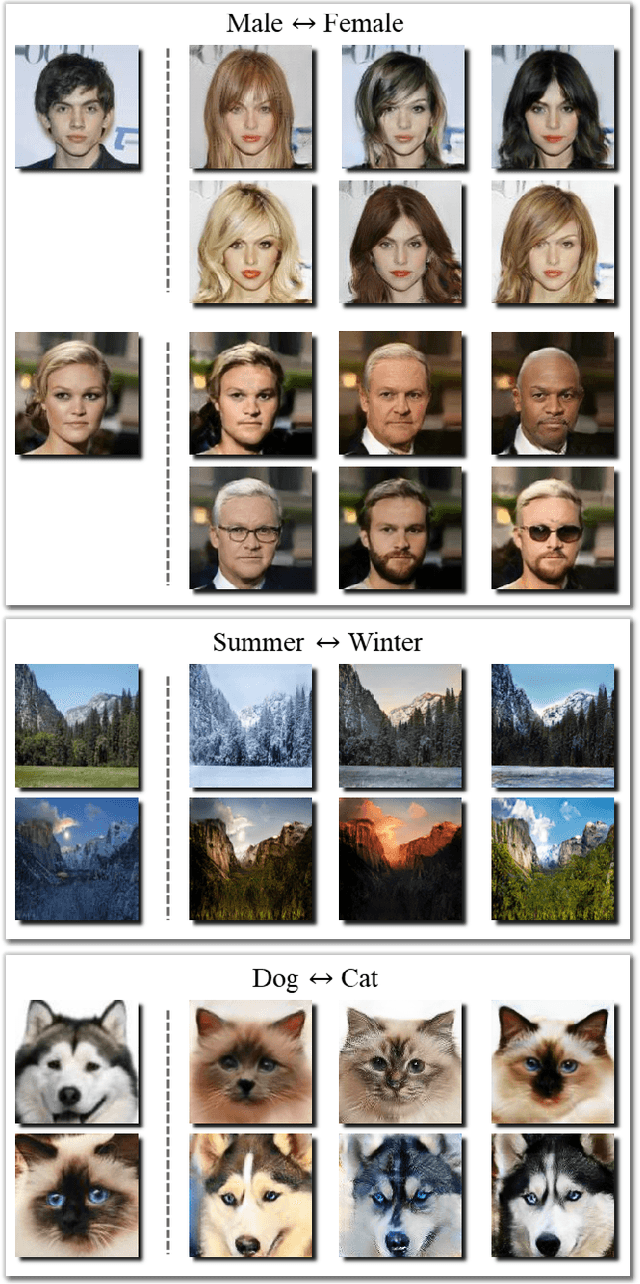
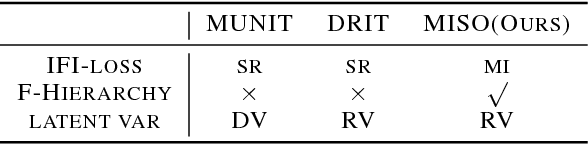
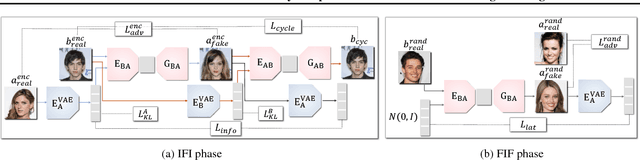
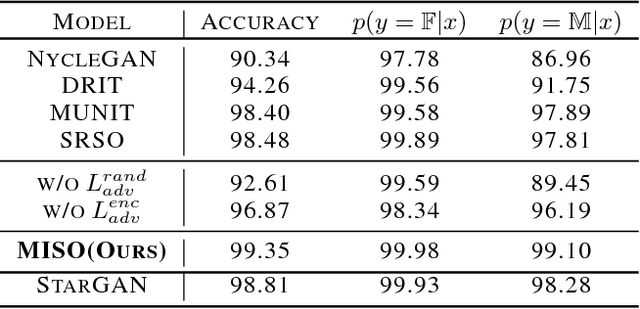
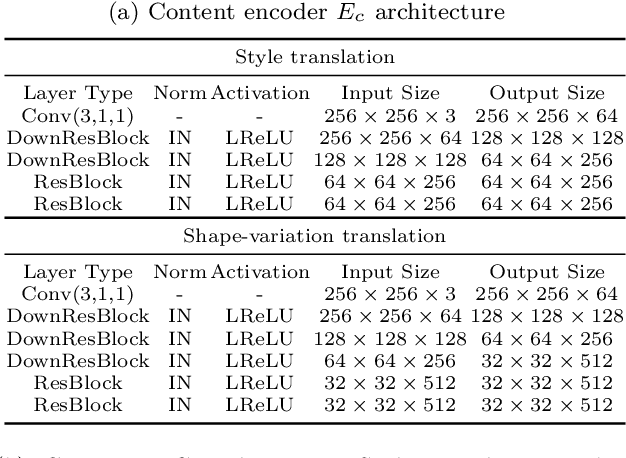
Unpaired multimodal image-to-image translation is a task of translating a given image in a source domain into diverse images in the target domain, overcoming the limitation of one-to-one mapping. Existing multimodal translation models are mainly based on the disentangled representations with an image reconstruction loss. We propose two approaches to improve multimodal translation quality. First, we use a content representation from the source domain conditioned on a style representation from the target domain. Second, rather than using a typical image reconstruction loss, we design MILO (Mutual Information LOss), a new stochastically-defined loss function based on information theory. This loss function directly reflects the interpretation of latent variables as a random variable. We show that our proposed model Mutual Information with StOchastic Style Representation(MISO) achieves state-of-the-art performance through extensive experiments on various real-world datasets.
Towards Learning a Self-inverse Network for Bidirectional Image-to-image Translation
Sep 16, 2019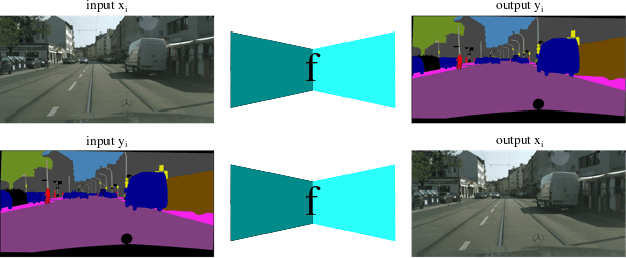
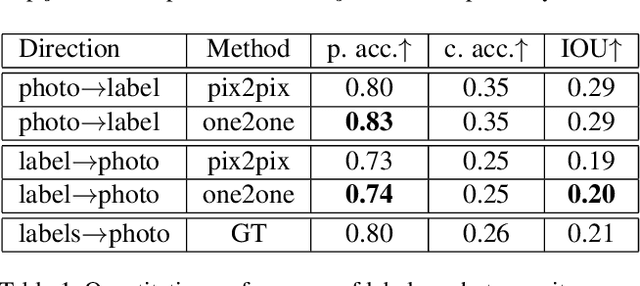
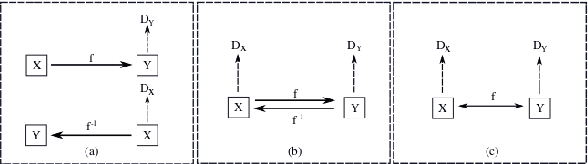
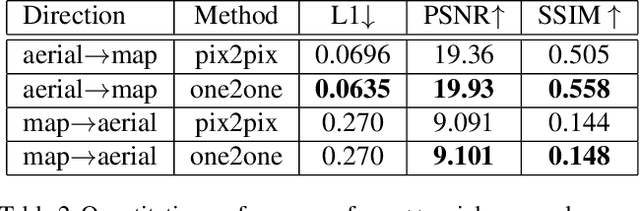
The one-to-one mapping is necessary for many bidirectional image-to-image translation applications, such as MRI image synthesis as MRI images are unique to the patient. State-of-the-art approaches for image synthesis from domain X to domain Y learn a convolutional neural network that meticulously maps between the domains. A different network is typically implemented to map along the opposite direction, from Y to X. In this paper, we explore the possibility of only wielding one network for bi-directional image synthesis. In other words, such an autonomous learning network implements a self-inverse function. A self-inverse network shares several distinct advantages: only one network instead of two, better generalization and more restricted parameter space. Most importantly, a self-inverse function guarantees a one-to-one mapping, a property that cannot be guaranteed by earlier approaches that are not self-inverse. The experiments on three datasets show that, compared with the baseline approaches that use two separate models for the image synthesis along two directions, our self-inverse network achieves better synthesis results in terms of standard metrics. Finally, our sensitivity analysis confirms the feasibility of learning a self-inverse function for the bidirectional image translation.
Disentangled Uncertainty and Out of Distribution Detection in Medical Generative Models
Nov 11, 2022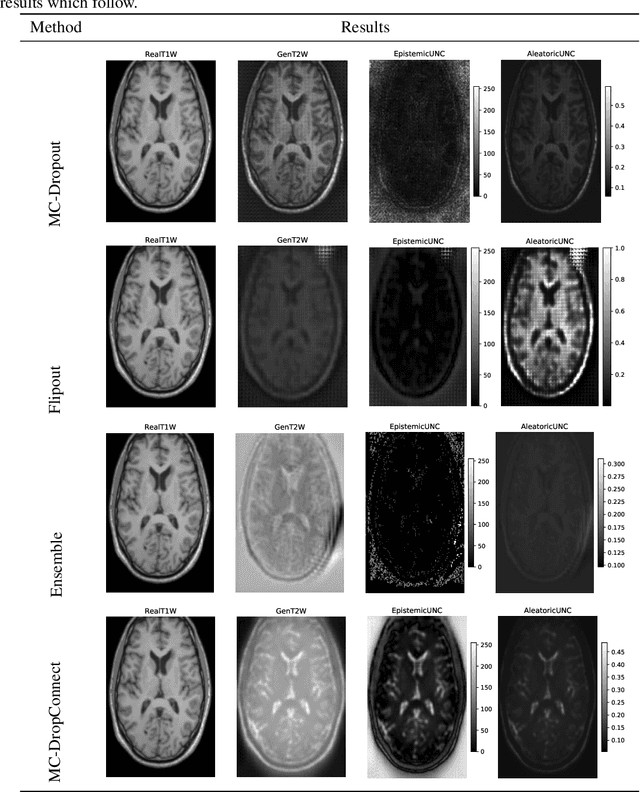
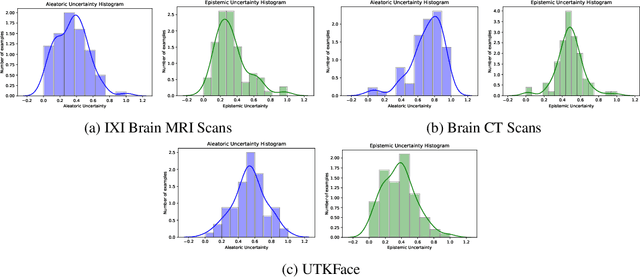
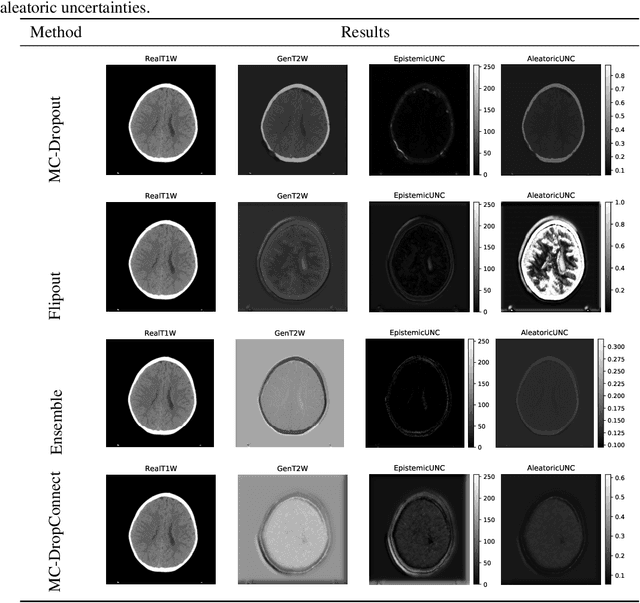
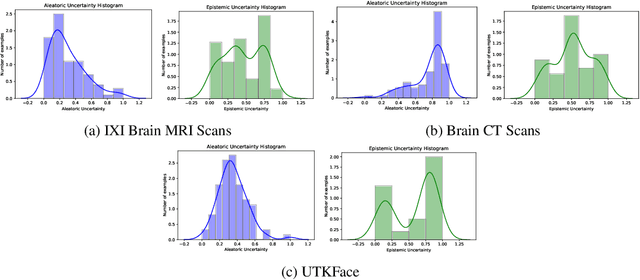
Trusting the predictions of deep learning models in safety critical settings such as the medical domain is still not a viable option. Distentangled uncertainty quantification in the field of medical imaging has received little attention. In this paper, we study disentangled uncertainties in image to image translation tasks in the medical domain. We compare multiple uncertainty quantification methods, namely Ensembles, Flipout, Dropout, and DropConnect, while using CycleGAN to convert T1-weighted brain MRI scans to T2-weighted brain MRI scans. We further evaluate uncertainty behavior in the presence of out of distribution data (Brain CT and RGB Face Images), showing that epistemic uncertainty can be used to detect out of distribution inputs, which should increase reliability of model outputs.
Towards Realistic Underwater Dataset Generation and Color Restoration
Nov 27, 2022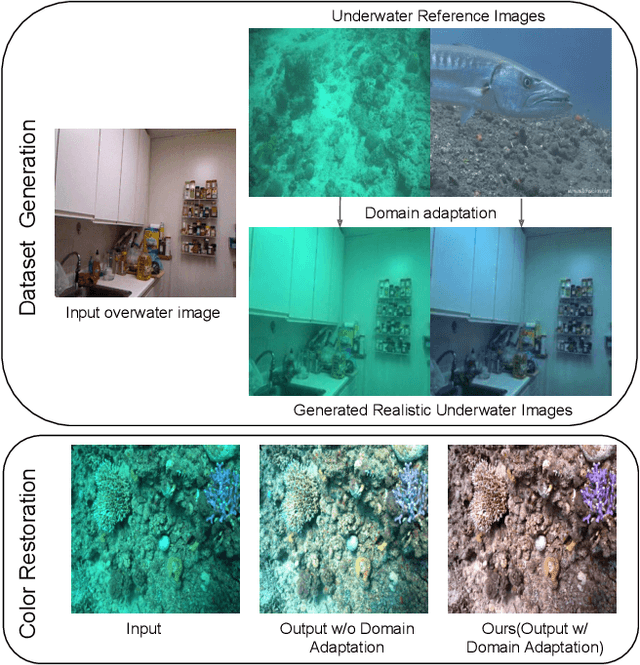
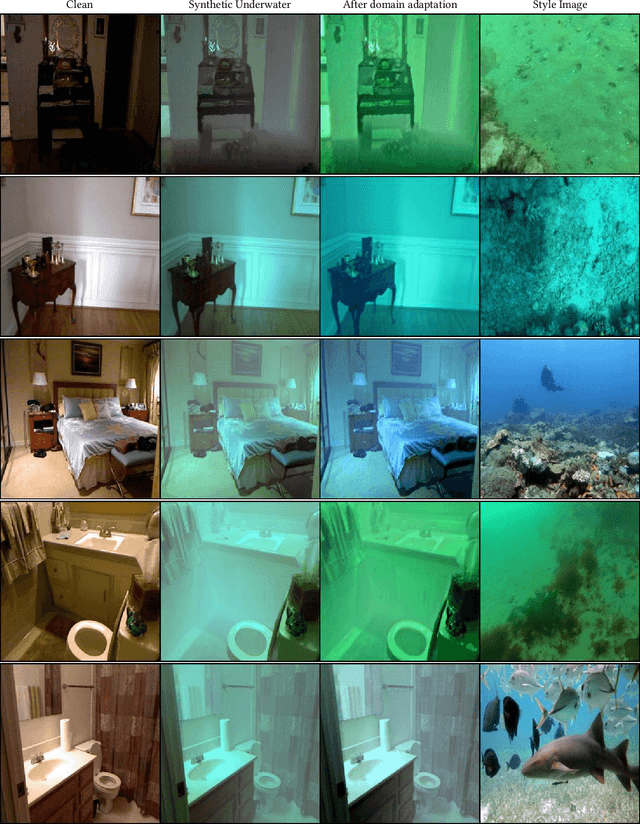
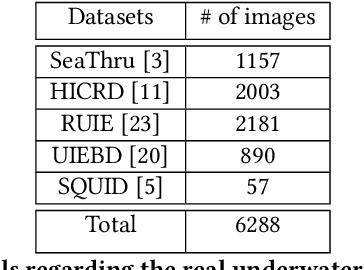

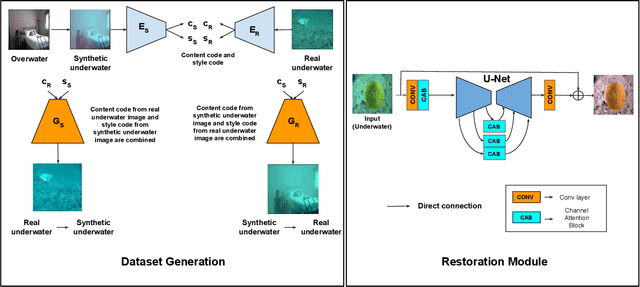
Recovery of true color from underwater images is an ill-posed problem. This is because the wide-band attenuation coefficients for the RGB color channels depend on object range, reflectance, etc. which are difficult to model. Also, there is backscattering due to suspended particles in water. Thus, most existing deep-learning based color restoration methods, which are trained on synthetic underwater datasets, do not perform well on real underwater data. This can be attributed to the fact that synthetic data cannot accurately represent real conditions. To address this issue, we use an image to image translation network to bridge the gap between the synthetic and real domains by translating images from synthetic underwater domain to real underwater domain. Using this multimodal domain adaptation technique, we create a dataset that can capture a diverse array of underwater conditions. We then train a simple but effective CNN based network on our domain adapted dataset to perform color restoration. Code and pre-trained models can be accessed at https://github.com/nehamjain10/TRUDGCR
Ultra-high-resolution unpaired stain transformation via Kernelized Instance Normalization
Aug 23, 2022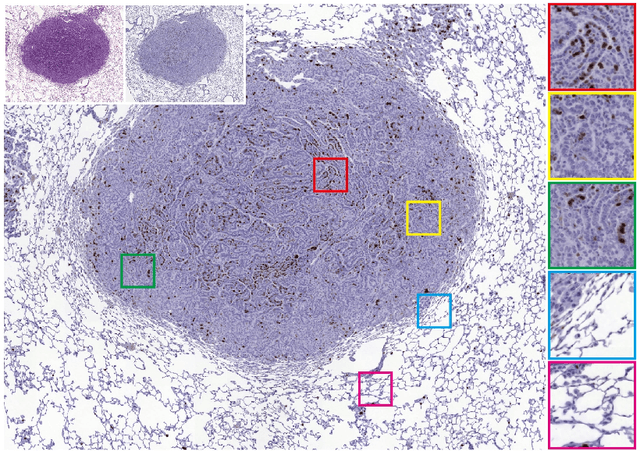
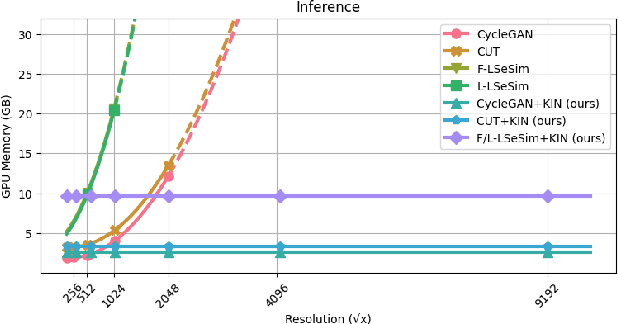
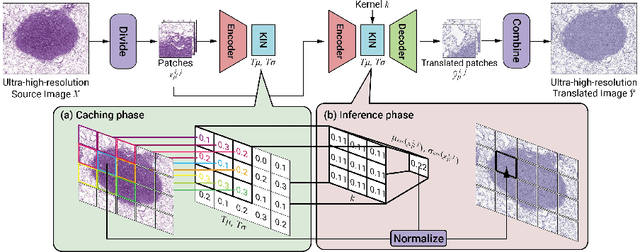
While hematoxylin and eosin (H&E) is a standard staining procedure, immunohistochemistry (IHC) staining further serves as a diagnostic and prognostic method. However, acquiring special staining results requires substantial costs. Hence, we proposed a strategy for ultra-high-resolution unpaired image-to-image translation: Kernelized Instance Normalization (KIN), which preserves local information and successfully achieves seamless stain transformation with constant GPU memory usage. Given a patch, corresponding position, and a kernel, KIN computes local statistics using convolution operation. In addition, KIN can be easily plugged into most currently developed frameworks without re-training. We demonstrate that KIN achieves state-of-the-art stain transformation by replacing instance normalization (IN) layers with KIN layers in three popular frameworks and testing on two histopathological datasets. Furthermore, we manifest the generalizability of KIN with high-resolution natural images. Finally, human evaluation and several objective metrics are used to compare the performance of different approaches. Overall, this is the first successful study for the ultra-high-resolution unpaired image-to-image translation with constant space complexity. Code is available at: https://github.com/Kaminyou/URUST
Is Image-to-Image Translation the Panacea for Multimodal Image Registration? A Comparative Study
Mar 30, 2021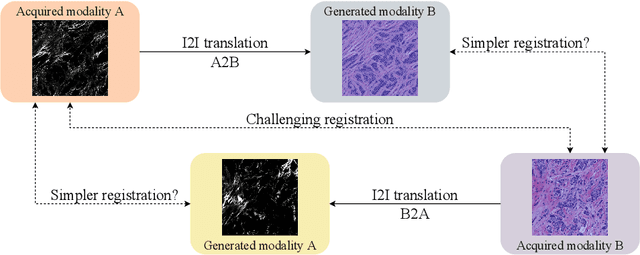
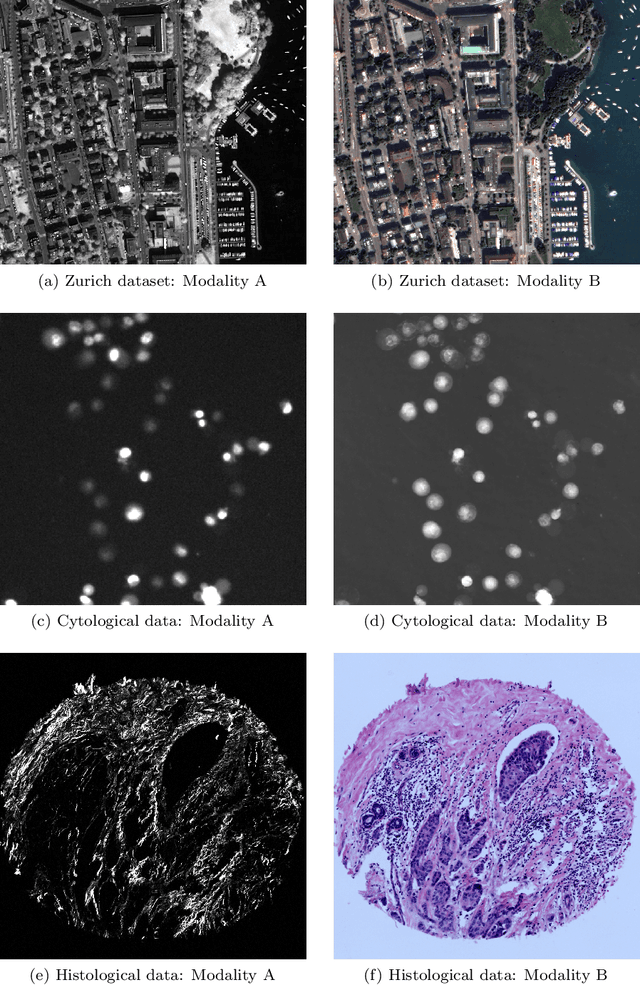
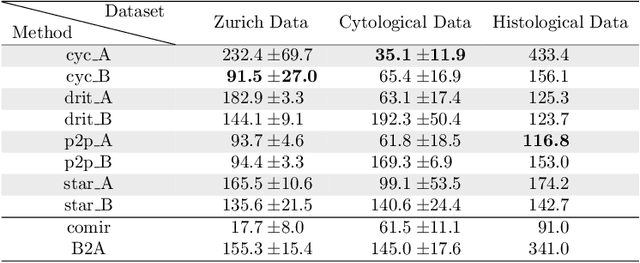

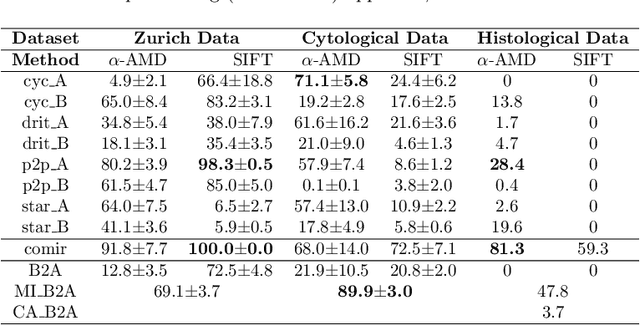
Despite current advancement in the field of biomedical image processing, propelled by the deep learning revolution, multimodal image registration, due to its several challenges, is still often performed manually by specialists. The recent success of image-to-image (I2I) translation in computer vision applications and its growing use in biomedical areas provide a tempting possibility of transforming the multimodal registration problem into a, potentially easier, monomodal one. We conduct an empirical study of the applicability of modern I2I translation methods for the task of multimodal biomedical image registration. We compare the performance of four Generative Adversarial Network (GAN)-based methods and one contrastive representation learning method, subsequently combined with two representative monomodal registration methods, to judge the effectiveness of modality translation for multimodal image registration. We evaluate these method combinations on three publicly available multimodal datasets of increasing difficulty, and compare with the performance of registration by Mutual Information maximisation and one modern data-specific multimodal registration method. Our results suggest that, although I2I translation may be helpful when the modalities to register are clearly correlated, registration of modalities which express distinctly different properties of the sample are not well handled by the I2I translation approach. When less information is shared between the modalities, the I2I translation methods struggle to provide good predictions, which impairs the registration performance. The evaluated representation learning method, which aims to find an in-between representation, manages better, and so does the Mutual Information maximisation approach. We share our complete experimental setup as open-source (https://github.com/Noodles-321/Registration).
Augmented Cyclic Consistency Regularization for Unpaired Image-to-Image Translation
Feb 29, 2020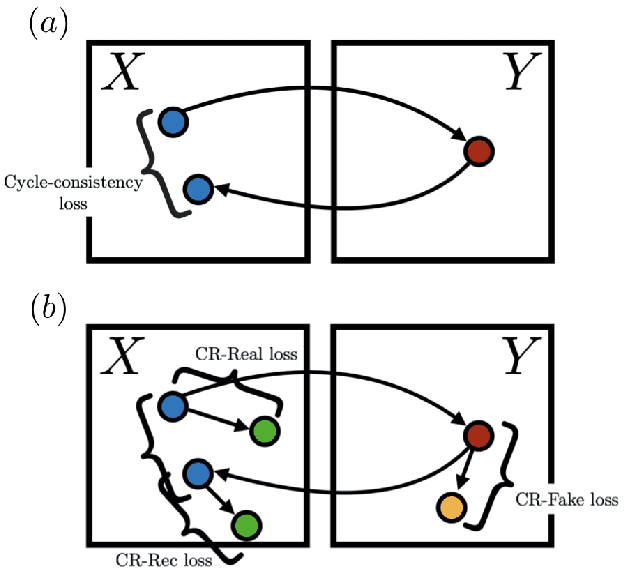
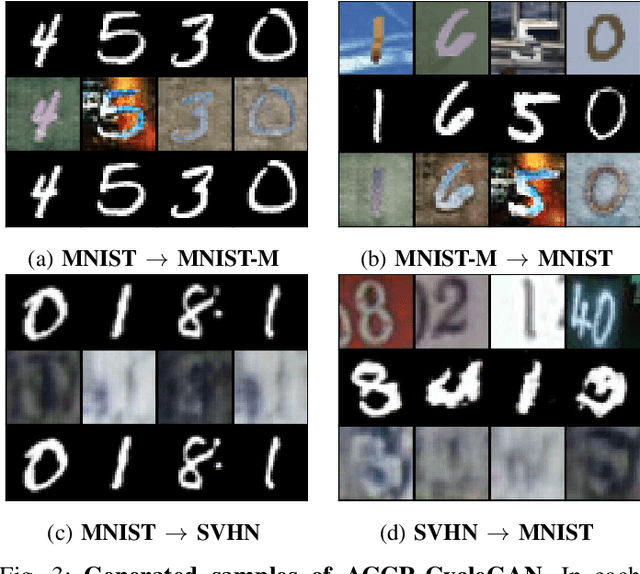
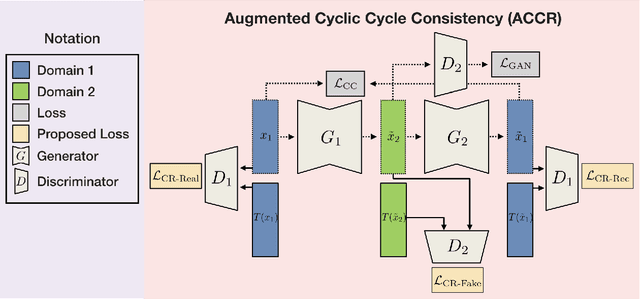


Unpaired image-to-image (I2I) translation has received considerable attention in pattern recognition and computer vision because of recent advancements in generative adversarial networks (GANs). However, due to the lack of explicit supervision, unpaired I2I models often fail to generate realistic images, especially in challenging datasets with different backgrounds and poses. Hence, stabilization is indispensable for real-world applications and GANs. Herein, we propose Augmented Cyclic Consistency Regularization (ACCR), a novel regularization method for unpaired I2I translation. Our main idea is to enforce consistency regularization originating from semi-supervised learning on the discriminators leveraging real, fake, reconstructed, and augmented samples. We regularize the discriminators to output similar predictions when fed pairs of original and perturbed images. We qualitatively clarify the generation property between unpaired I2I models and standard GANs, and explain why consistency regularization on fake and reconstructed samples works well. Quantitatively, our method outperforms the consistency regularized GAN (CR-GAN) in digit translations and demonstrates efficacy against several data augmentation variants and cycle-consistent constraints.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge