Yuxing Li
Relay Buffer Independent Communication over Pooled HBM for Efficient MoE Inference on Ascend
May 07, 2026Abstract:Mixture-of-Experts (MoE) inference requires large-scale token exchange across devices, making dispatch and combine major bottlenecks in both prefill and decode. Beyond network transfer, routing-driven layout transformation, temporary relay, and output restoration can add substantial overhead. Existing MoE communication paths are often buffer-centric, using explicit inter-process relay and reordering buffers around collective transfer. This report presents a relay-buffer-free communication design for MoE inference acceleration on Ascend systems. The design reorganizes dispatch and combine around direct placement into destination expert windows and direct reading from remote expert windows. Built on globally pooled high-bandwidth memory and symmetric-memory allocation, it removes most intermediate relay and reordering buffers while retaining only lightweight control state, including counts, offsets, and synchronization metadata. We instantiate the design as two schedules for the main phases of MoE inference: a prefill schedule with richer planning state for throughput-oriented execution, and a compact decode schedule for latency-sensitive execution. Experiments on Ascend-based MoE workloads show reduced dispatch and combine latency in both settings. At the serving level, the implementation improves time to first token (TTFT), preserves competitive time per output token (TPOT), and enlarges the feasible scheduling space under practical latency constraints. These results indicate that, on platforms with globally addressable device memory, reducing intermediate buffering and output restoration around expert execution is an effective direction for accelerating MoE inference.
Deep Speckle Holography Redefines Label-free Nanoparticle Phenotyping
May 03, 2026Abstract:Nanoparticle metrology has long been constrained by the assumption that, in mixed and unprocessed fluids, particle size, morphology, composition, and species-specific abundance cannot be resolved simultaneously from a single label-free measurement. Here, we revisit this long-standing limitation by showing that complex forward speckle-holographic fields define an information-rich optical space for multidimensional particle signatures. We report deep speckle holography, a physics-informed generative framework that profiles particle identity, size, morphology, and species-resolved abundance from a single non-contact optical measurement. Across purified suspensions, mixed particle populations, environmental waters, human urine, and other unprocessed native fluids, the method enables direct nanoparticle inference without purification, labeling, or destructive preprocessing, delivering concurrent multidimensional readouts in 0.9 s over a dynamic range spanning 10 orders of magnitude. Deep speckle holography establishes a route toward direct label-free nanoparticle phenotyping in real-world fluids, moving nanoscale measurement beyond isolated-particle characterization toward multidimensional inference in complex mixtures, and expanding the scope of questions nanoscale measurement can address, from real-time tracking of nanoparticle transformations in living and environmental systems to non-invasive quality control of nanomedicine formulations, and beyond.
EPOFusion: Exposure aware Progressive Optimization Method for Infrared and Visible Image Fusion
Mar 17, 2026Abstract:Overexposure frequently occurs in practical scenarios, causing the loss of critical visual information. However, existing infrared and visible fusion methods still exhibit unsatisfactory performance in highly bright regions. To address this, we propose EPOFusion, an exposure-aware fusion model. Specifically, a guidance module is introduced to facilitate the encoder in extracting fine-grained infrared features from overexposed regions. Meanwhile, an iterative decoder incorporating a multiscale context fusion module is designed to progressively enhance the fused image, ensuring consistent details and superior visual quality. Finally, an adaptive loss function dynamically constrains the fusion process, enabling an effective balance between the modalities under varying exposure conditions. To achieve better exposure awareness, we construct the first infrared and visible overexposure dataset (IVOE) with high quality infrared guided annotations for overexposed regions. Extensive experiments show that EPOFusion outperforms existing methods. It maintains infrared cues in overexposed regions while achieving visually faithful fusion in non-overexposed areas, thereby enhancing both visual fidelity and downstream task performance. Code, fusion results and IVOE dataset will be made available at https://github.com/warren-wzw/EPOFusion.git.
Personalized Federated Learning with Residual Fisher Information for Medical Image Segmentation
Mar 16, 2026Abstract:Federated learning enables multiple clients (institutions) to collaboratively train machine learning models without sharing their private data. To address the challenge of data heterogeneity across clients, personalized federated learning (pFL) aims to learn customized models for each client. In this work, we propose pFL-ResFIM, a novel pFL framework that achieves client-adaptive personalization at the parameter level. Specifically, we introduce a new metric, Residual Fisher Information Matrix (ResFIM), to quantify the sensitivity of model parameters to domain discrepancies. To estimate ResFIM for each client model under privacy constraints, we employ a spectral transfer strategy that generates simulated data reflecting the domain styles of different clients. Based on the estimated ResFIM, we partition model parameters into domain-sensitive and domain-invariant components. A personalized model for each client is then constructed by aggregating only the domain-invariant parameters on the server. Extensive experiments on public datasets demonstrate that pFL-ResFIM consistently outperforms state-of-the-art methods, validating its effectiveness.
Joint Segmentation and Grading with Iterative Optimization for Multimodal Glaucoma Diagnosis
Mar 15, 2026Abstract:Accurate diagnosis of glaucoma is challenging, as early-stage changes are subtle and often lack clear structural or appearance cues. Most existing approaches rely on a single modality, such as fundus or optical coherence tomography (OCT), capturing only partial pathological information and often missing early disease progression. In this paper, we propose an iterative multimodal optimization model (IMO) for joint segmentation and grading. IMO integrates fundus and OCT features through a mid-level fusion strategy, enhanced by a cross-modal feature alignment (CMFA) module to reduce modality discrepancies. An iterative refinement decoder progressively optimizes the multimodal features through a denoising diffusion mechanism, enabling fine-grained segmentation of the optic disc and cup while supporting accurate glaucoma grading. Extensive experiments show that our method effectively integrates multimodal features, providing a comprehensive and clinically significant approach to glaucoma assessment. Source codes are available at https://github.com/warren-wzw/IMO.git.
FL-MedSegBench: A Comprehensive Benchmark for Federated Learning on Medical Image Segmentation
Mar 12, 2026Abstract:Federated learning (FL) offers a privacy-preserving paradigm for collaborative medical image analysis without sharing raw data. However, the absence of standardized benchmarks for medical image segmentation hinders fair and comprehensive evaluation of FL methods. To address this gap, we introduce FL-MedSegBench, the first comprehensive benchmark for federated learning on medical image segmentation. Our benchmark encompasses nine segmentation tasks across ten imaging modalities, covering both 2D and 3D formats with realistic clinical heterogeneity. We systematically evaluate eight generic FL (gFL) and five personalized FL (pFL) methods across multiple dimensions: segmentation accuracy, fairness, communication efficiency, convergence behavior, and generalization to unseen domains. Extensive experiments reveal several key insights: (i) pFL methods, particularly those with client-specific batch normalization (\textit{e.g.}, FedBN), consistently outperform generic approaches; (ii) No single method universally dominates, with performance being dataset-dependent; (iii) Communication frequency analysis shows normalization-based personalization methods exhibit remarkable robustness to reduced communication frequency; (iv) Fairness evaluation identifies methods like Ditto and FedRDN that protect underperforming clients; (v) A method's generalization to unseen domains is strongly tied to its ability to perform well across participating clients. We will release an open-source toolkit to foster reproducible research and accelerate clinically applicable FL solutions, providing empirically grounded guidelines for real-world clinical deployment. The source code is available at https://github.com/meiluzhu/FL-MedSegBench.
Knowledge Transfer between Datasets for Learning-based Tissue Microstructure Estimation
Oct 24, 2019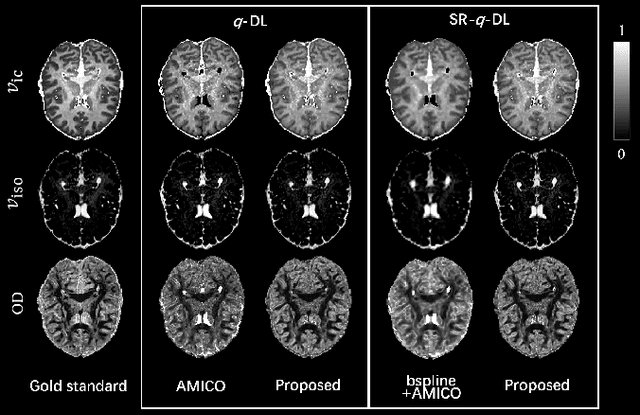
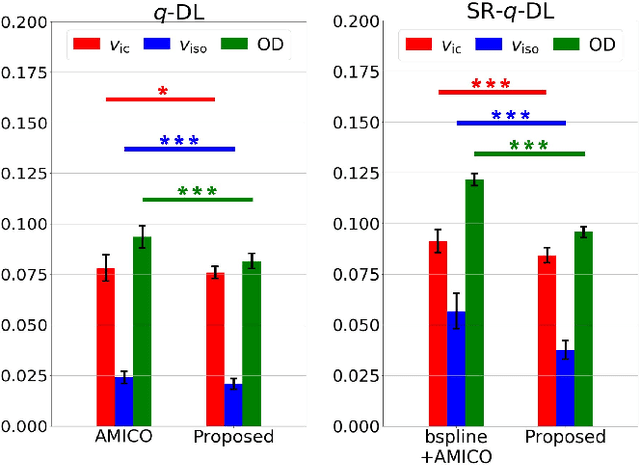
Abstract:Learning-based approaches, especially those based on deep networks, have enabled high-quality estimation of tissue microstructure from low-quality diffusion magnetic resonance imaging (dMRI) scans, which are acquired with a limited number of diffusion gradients and a relatively poor spatial resolution. These learning-based approaches to tissue microstructure estimation require acquisitions of training dMRI scans with high-quality diffusion signals, which are densely sampled in the q-space and have a high spatial resolution. However, the acquisition of training scans may not be available for all datasets. Therefore, we explore knowledge transfer between different dMRI datasets so that learning-based tissue microstructure estimation can be applied for datasets where training scans are not acquired. Specifically, for a target dataset of interest, where only low-quality diffusion signals are acquired without training scans, we exploit the information in a source dMRI dataset acquired with high-quality diffusion signals. We interpolate the diffusion signals in the source dataset in the q-space using a dictionary-based signal representation, so that the interpolated signals match the acquisition scheme of the target dataset. Then, the interpolated signals are used together with the high-quality tissue microstructure computed from the source dataset to train deep networks that perform tissue microstructure estimation for the target dataset. Experiments were performed on brain dMRI scans with low-quality diffusion signals, where the benefit of the proposed strategy is demonstrated.
Semi-Supervised Brain Lesion Segmentation with an Adapted Mean Teacher Model
Mar 04, 2019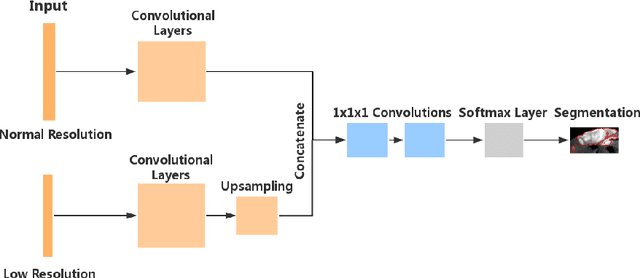
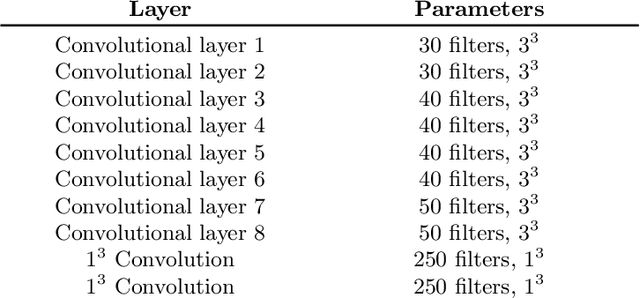
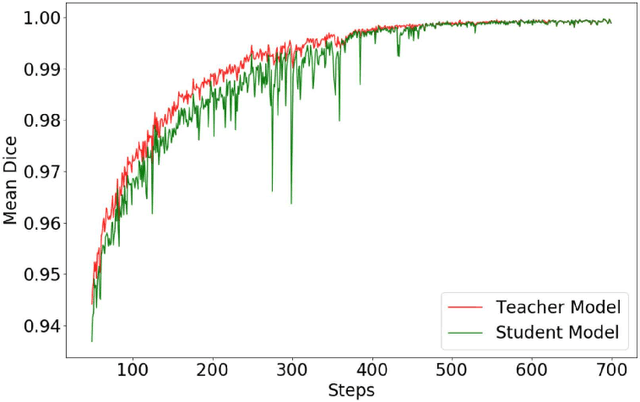
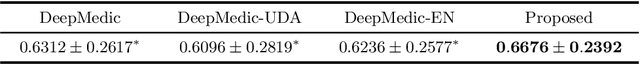
Abstract:Automated brain lesion segmentation provides valuable information for the analysis and intervention of patients. In particular, methods based on convolutional neural networks (CNNs) have achieved state-of-the-art segmentation performance. However, CNNs usually require a decent amount of annotated data, which may be costly and time-consuming to obtain. Since unannotated data is generally abundant, it is desirable to use unannotated data to improve the segmentation performance for CNNs when limited annotated data is available. In this work, we propose a semi-supervised learning (SSL) approach to brain lesion segmentation, where unannotated data is incorporated into the training of CNNs. We adapt the mean teacher model, which is originally developed for SSL-based image classification, for brain lesion segmentation. Assuming that the network should produce consistent outputs for similar inputs, a loss of segmentation consistency is designed and integrated into a self-ensembling framework. Specifically, we build a student model and a teacher model, which share the same CNN architecture for segmentation. The student and teacher models are updated alternately. At each step, the student model learns from the teacher model by minimizing the weighted sum of the segmentation loss computed from annotated data and the segmentation consistency loss between the teacher and student models computed from unannotated data. Then, the teacher model is updated by combining the updated student model with the historical information of teacher models using an exponential moving average strategy. For demonstration, the proposed approach was evaluated on ischemic stroke lesion segmentation, where it improves stroke lesion segmentation with the incorporation of unannotated data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge