Yumin Zhang
TeachPro: Multi-Label Qualitative Teaching Evaluation via Cross-View Graph Synergy and Semantic Anchored Evidence Encoding
Jan 14, 2026Abstract:Standardized Student Evaluation of Teaching often suffer from low reliability, restricted response options, and response distortion. Existing machine learning methods that mine open-ended comments usually reduce feedback to binary sentiment, which overlooks concrete concerns such as content clarity, feedback timeliness, and instructor demeanor, and provides limited guidance for instructional improvement.We propose TeachPro, a multi-label learning framework that systematically assesses five key teaching dimensions: professional expertise, instructional behavior, pedagogical efficacy, classroom experience, and other performance metrics. We first propose a Dimension-Anchored Evidence Encoder, which integrates three core components: (i) a pre-trained text encoder that transforms qualitative feedback annotations into contextualized embeddings; (ii) a prompt module that represents five teaching dimensions as learnable semantic anchors; and (iii) a cross-attention mechanism that aligns evidence with pedagogical dimensions within a structured semantic space. We then propose a Cross-View Graph Synergy Network to represent student comments. This network comprises two components: (i) a Syntactic Branch that extracts explicit grammatical dependencies from parse trees, and (ii) a Semantic Branch that models latent conceptual relations derived from BERT-based similarity graphs. BiAffine fusion module aligns syntactic and semantic units, while a differential regularizer disentangles embeddings to encourage complementary representations. Finally, a cross-attention mechanism bridges the dimension-anchored evidence with the multi-view comment representations. We also contribute a novel benchmark dataset featuring expert qualitative annotations and multi-label scores. Extensive experiments demonstrate that TeachPro offers superior diagnostic granularity and robustness across diverse evaluation settings.
A Visual-Inertial Motion Prior SLAM for Dynamic Environments
Mar 30, 2025Abstract:The Visual-Inertial Simultaneous Localization and Mapping (VI-SLAM) algorithms which are mostly based on static assumption are widely used in fields such as robotics, UAVs, VR, and autonomous driving. To overcome the localization risks caused by dynamic landmarks in most VI-SLAM systems, a robust visual-inertial motion prior SLAM system, named (IDY-VINS), is proposed in this paper which effectively handles dynamic landmarks using inertial motion prior for dynamic environments to varying degrees. Specifically, potential dynamic landmarks are preprocessed during the feature tracking phase by the probabilistic model of landmarks' minimum projection errors which are obtained from inertial motion prior and epipolar constraint. Subsequently, a bundle adjustment (BA) residual is proposed considering the minimum projection error prior for dynamic candidate landmarks. This residual is integrated into a sliding window based nonlinear optimization process to estimate camera poses, IMU states and landmark positions while minimizing the impact of dynamic candidate landmarks that deviate from the motion prior. Finally, experimental results demonstrate that our proposed system outperforms state-of-the-art methods in terms of localization accuracy and time cost by robustly mitigating the influence of dynamic landmarks.
FedSCA: Federated Tuning with Similarity-guided Collaborative Aggregation for Heterogeneous Medical Image Segmentation
Mar 19, 2025



Abstract:Transformer-based foundation models (FMs) have recently demonstrated remarkable performance in medical image segmentation. However, scaling these models is challenging due to the limited size of medical image datasets within isolated hospitals, where data centralization is restricted due to privacy concerns. These constraints, combined with the data-intensive nature of FMs, hinder their broader application. Integrating federated learning (FL) with foundation models (FLFM) fine-tuning offers a potential solution to these challenges by enabling collaborative model training without data sharing, thus allowing FMs to take advantage of a diverse pool of sensitive medical image data across hospitals/clients. However, non-independent and identically distributed (non-IID) data among clients, paired with computational and communication constraints in federated environments, presents an additional challenge that limits further performance improvements and remains inadequately addressed in existing studies. In this work, we propose a novel FLFM fine-tuning framework, \underline{\textbf{Fed}}erated tuning with \underline{\textbf{S}}imilarity-guided \underline{\textbf{C}}ollaborative \underline{\textbf{A}}ggregation (FedSCA), encompassing all phases of the FL process. This includes (1) specially designed parameter-efficient fine-tuning (PEFT) for local client training to enhance computational efficiency; (2) partial low-level adapter transmission for communication efficiency; and (3) similarity-guided collaborative aggregation (SGCA) on the server side to address non-IID issues. Extensive experiments on three FL benchmarks for medical image segmentation demonstrate the effectiveness of our proposed FedSCA, establishing new SOTA performance.
Exemplar-condensed Federated Class-incremental Learning
Dec 25, 2024



Abstract:We propose Exemplar-Condensed federated class-incremental learning (ECoral) to distil the training characteristics of real images from streaming data into informative rehearsal exemplars. The proposed method eliminates the limitations of exemplar selection in replay-based approaches for mitigating catastrophic forgetting in federated continual learning (FCL). The limitations particularly related to the heterogeneity of information density of each summarized data. Our approach maintains the consistency of training gradients and the relationship to past tasks for the summarized exemplars to represent the streaming data compared to the original images effectively. Additionally, our approach reduces the information-level heterogeneity of the summarized data by inter-client sharing of the disentanglement generative model. Extensive experiments show that our ECoral outperforms several state-of-the-art methods and can be seamlessly integrated with many existing approaches to enhance performance.
Prototype Correlation Matching and Class-Relation Reasoning for Few-Shot Medical Image Segmentation
Jun 07, 2024Abstract:Few-shot medical image segmentation has achieved great progress in improving accuracy and efficiency of medical analysis in the biomedical imaging field. However, most existing methods cannot explore inter-class relations among base and novel medical classes to reason unseen novel classes. Moreover, the same kind of medical class has large intra-class variations brought by diverse appearances, shapes and scales, thus causing ambiguous visual characterization to degrade generalization performance of these existing methods on unseen novel classes. To address the above challenges, in this paper, we propose a \underline{\textbf{P}}rototype correlation \underline{\textbf{M}}atching and \underline{\textbf{C}}lass-relation \underline{\textbf{R}}easoning (i.e., \textbf{PMCR}) model. The proposed model can effectively mitigate false pixel correlation matches caused by large intra-class variations while reasoning inter-class relations among different medical classes. Specifically, in order to address false pixel correlation match brought by large intra-class variations, we propose a prototype correlation matching module to mine representative prototypes that can characterize diverse visual information of different appearances well. We aim to explore prototype-level rather than pixel-level correlation matching between support and query features via optimal transport algorithm to tackle false matches caused by intra-class variations. Meanwhile, in order to explore inter-class relations, we design a class-relation reasoning module to segment unseen novel medical objects via reasoning inter-class relations between base and novel classes. Such inter-class relations can be well propagated to semantic encoding of local query features to improve few-shot segmentation performance. Quantitative comparisons illustrates the large performance improvement of our model over other baseline methods.
ExactDreamer: High-Fidelity Text-to-3D Content Creation via Exact Score Matching
May 24, 2024



Abstract:Text-to-3D content creation is a rapidly evolving research area. Given the scarcity of 3D data, current approaches often adapt pre-trained 2D diffusion models for 3D synthesis. Among these approaches, Score Distillation Sampling (SDS) has been widely adopted. However, the issue of over-smoothing poses a significant limitation on the high-fidelity generation of 3D models. To address this challenge, LucidDreamer replaces the Denoising Diffusion Probabilistic Model (DDPM) in SDS with the Denoising Diffusion Implicit Model (DDIM) to construct Interval Score Matching (ISM). However, ISM inevitably inherits inconsistencies from DDIM, causing reconstruction errors during the DDIM inversion process. This results in poor performance in the detailed generation of 3D objects and loss of content. To alleviate these problems, we propose a novel method named Exact Score Matching (ESM). Specifically, ESM leverages auxiliary variables to mathematically guarantee exact recovery in the DDIM reverse process. Furthermore, to effectively capture the dynamic changes of the original and auxiliary variables, the LoRA of a pre-trained diffusion model implements these exact paths. Extensive experiments demonstrate the effectiveness of ESM in text-to-3D generation, particularly highlighting its superiority in detailed generation.
From Sora What We Can See: A Survey of Text-to-Video Generation
May 17, 2024



Abstract:With impressive achievements made, artificial intelligence is on the path forward to artificial general intelligence. Sora, developed by OpenAI, which is capable of minute-level world-simulative abilities can be considered as a milestone on this developmental path. However, despite its notable successes, Sora still encounters various obstacles that need to be resolved. In this survey, we embark from the perspective of disassembling Sora in text-to-video generation, and conducting a comprehensive review of literature, trying to answer the question, \textit{From Sora What We Can See}. Specifically, after basic preliminaries regarding the general algorithms are introduced, the literature is categorized from three mutually perpendicular dimensions: evolutionary generators, excellent pursuit, and realistic panorama. Subsequently, the widely used datasets and metrics are organized in detail. Last but more importantly, we identify several challenges and open problems in this domain and propose potential future directions for research and development.
BAMBOO: a predictive and transferable machine learning force field framework for liquid electrolyte development
Apr 12, 2024Abstract:Despite the widespread applications of machine learning force field (MLFF) on solids and small molecules, there is a notable gap in applying MLFF to complex liquid electrolytes. In this work, we introduce BAMBOO (ByteDance AI Molecular Simulation Booster), a novel framework for molecular dynamics (MD) simulations, with a demonstration of its capabilities in the context of liquid electrolytes for lithium batteries. We design a physics-inspired graph equivariant transformer architecture as the backbone of BAMBOO to learn from quantum mechanical simulations. Additionally, we pioneer an ensemble knowledge distillation approach and apply it on MLFFs to improve the stability of MD simulations. Finally, we propose the density alignment algorithm to align BAMBOO with experimental measurements. BAMBOO demonstrates state-of-the-art accuracy in predicting key electrolyte properties such as density, viscosity, and ionic conductivity across various solvents and salt combinations. Our current model, trained on more than 15 chemical species, achieves the average density error of 0.01 g/cm$^3$ on various compositions compared with experimental data. Moreover, our model demonstrates transferability to molecules not included in the quantum mechanical dataset. We envision this work as paving the way to a "universal MLFF" capable of simulating properties of common organic liquids.
Crucial Semantic Classifier-based Adversarial Learning for Unsupervised Domain Adaptation
Feb 03, 2023



Abstract:Unsupervised Domain Adaptation (UDA), which aims to explore the transferrable features from a well-labeled source domain to a related unlabeled target domain, has been widely progressed. Nevertheless, as one of the mainstream, existing adversarial-based methods neglect to filter the irrelevant semantic knowledge, hindering adaptation performance improvement. Besides, they require an additional domain discriminator that strives extractor to generate confused representations, but discrete designing may cause model collapse. To tackle the above issues, we propose Crucial Semantic Classifier-based Adversarial Learning (CSCAL), which pays more attention to crucial semantic knowledge transferring and leverages the classifier to implicitly play the role of domain discriminator without extra network designing. Specifically, in intra-class-wise alignment, a Paired-Level Discrepancy (PLD) is designed to transfer crucial semantic knowledge. Additionally, based on classifier predictions, a Nuclear Norm-based Discrepancy (NND) is formed that considers inter-class-wise information and improves the adaptation performance. Moreover, CSCAL can be effortlessly merged into different UDA methods as a regularizer and dramatically promote their performance.
Consecutive Knowledge Meta-Adaptation Learning for Unsupervised Medical Diagnosis
Sep 21, 2022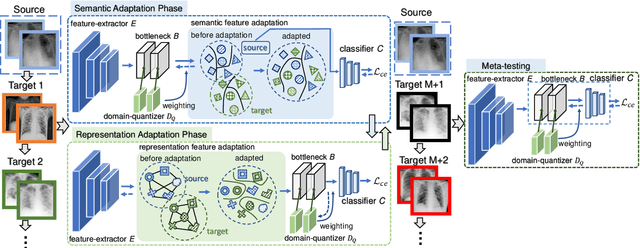
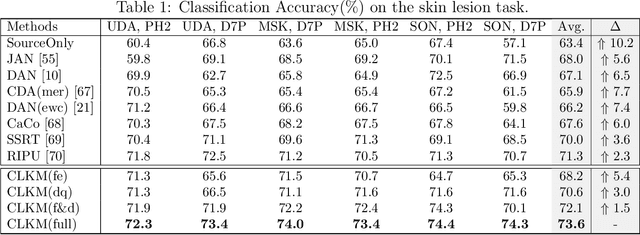
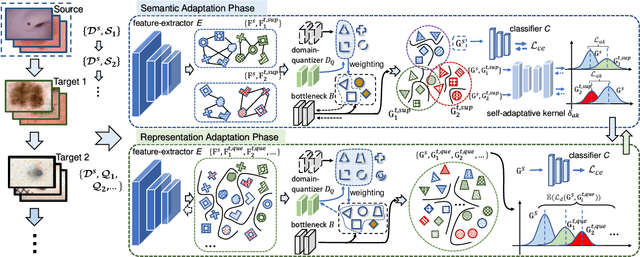
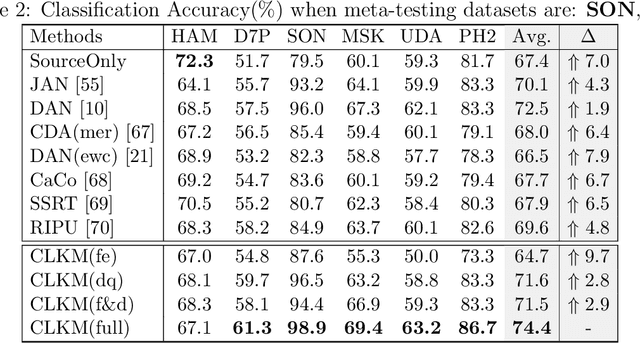
Abstract:Deep learning-based Computer-Aided Diagnosis (CAD) has attracted appealing attention in academic researches and clinical applications. Nevertheless, the Convolutional Neural Networks (CNNs) diagnosis system heavily relies on the well-labeled lesion dataset, and the sensitivity to the variation of data distribution also restricts the potential application of CNNs in CAD. Unsupervised Domain Adaptation (UDA) methods are developed to solve the expensive annotation and domain gaps problem and have achieved remarkable success in medical image analysis. Yet existing UDA approaches only adapt knowledge learned from the source lesion domain to a single target lesion domain, which is against the clinical scenario: the new unlabeled target domains to be diagnosed always arrive in an online and continual manner. Moreover, the performance of existing approaches degrades dramatically on previously learned target lesion domains, due to the newly learned knowledge overwriting the previously learned knowledge (i.e., catastrophic forgetting). To deal with the above issues, we develop a meta-adaptation framework named Consecutive Lesion Knowledge Meta-Adaptation (CLKM), which mainly consists of Semantic Adaptation Phase (SAP) and Representation Adaptation Phase (RAP) to learn the diagnosis model in an online and continual manner. In the SAP, the semantic knowledge learned from the source lesion domain is transferred to consecutive target lesion domains. In the RAP, the feature-extractor is optimized to align the transferable representation knowledge across the source and multiple target lesion domains.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge