Yiyu Shi
Contrastive Learning with Temporal Correlated Medical Images: A Case Study using Lung Segmentation in Chest X-Rays
Sep 16, 2021
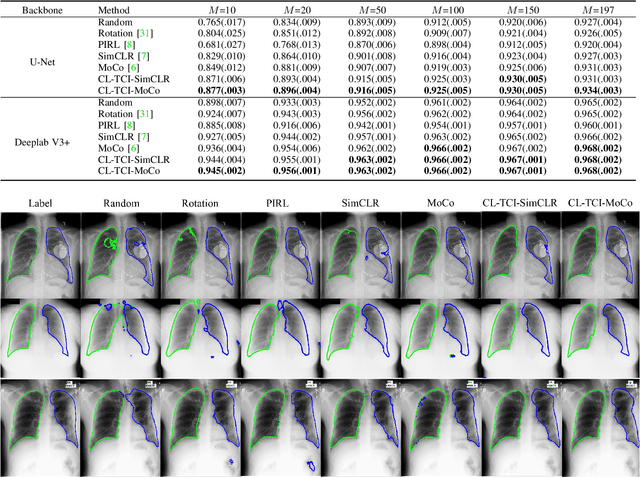
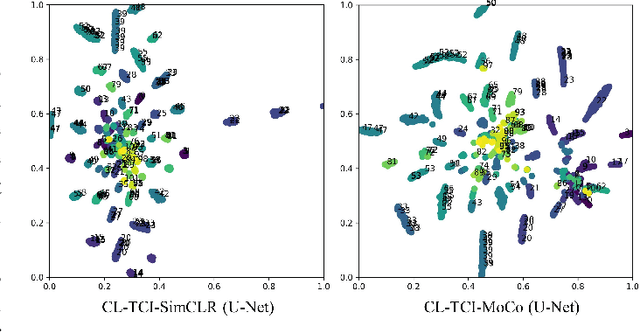
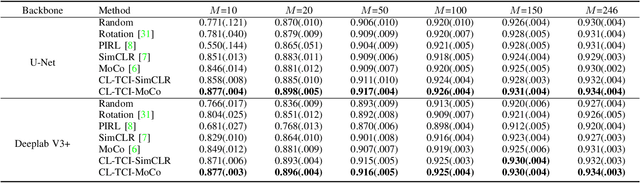
Abstract:Contrastive learning has been proved to be a promising technique for image-level representation learning from unlabeled data. Many existing works have demonstrated improved results by applying contrastive learning in classification and object detection tasks for either natural images or medical images. However, its application to medical image segmentation tasks has been limited. In this work, we use lung segmentation in chest X-rays as a case study and propose a contrastive learning framework with temporal correlated medical images, named CL-TCI, to learn superior encoders for initializing the segmentation network. We adapt CL-TCI from two state-of-the-art contrastive learning methods-MoCo and SimCLR. Experiment results on three chest X-ray datasets show that under two different segmentation backbones, U-Net and Deeplab-V3, CL-TCI can outperform all baselines that do not incorporate any temporal correlation in both semi-supervised learning setting and transfer learning setting with limited annotation. This suggests that information among temporal correlated medical images can indeed improve contrastive learning performance. Between the two variations of CL-TCI, CL-TCI adapted from MoCo outperforms CL-TCI adapted from SimCLR in most settings, indicating that more contrastive samples can benefit the learning process and help the network learn high-quality representations.
Hardware-aware Real-time Myocardial Segmentation Quality Control in Contrast Echocardiography
Sep 14, 2021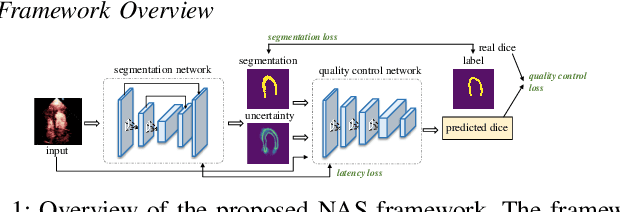
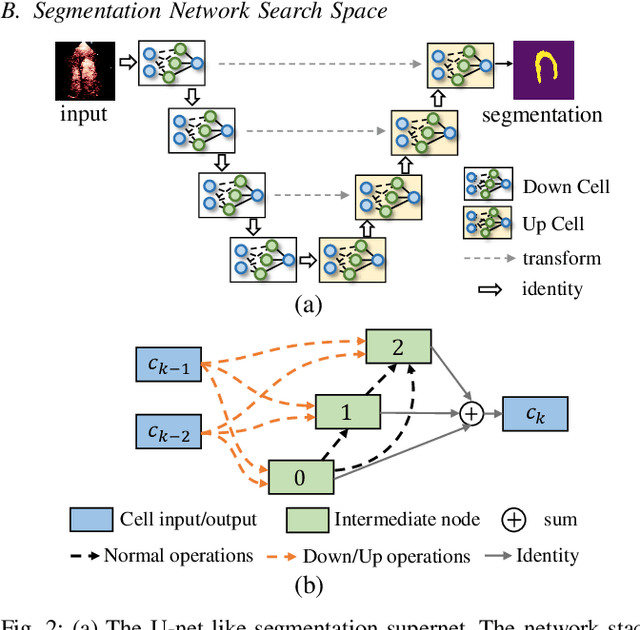
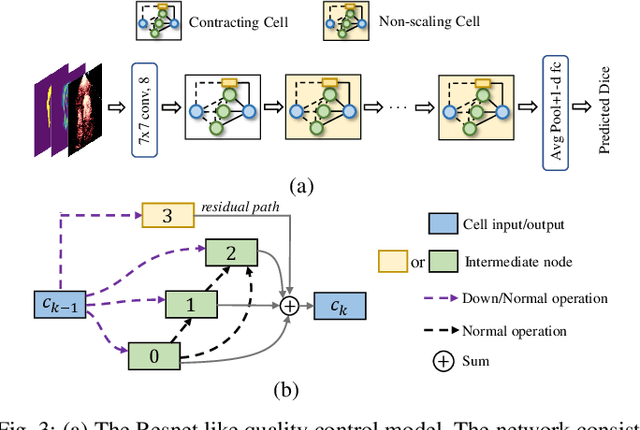
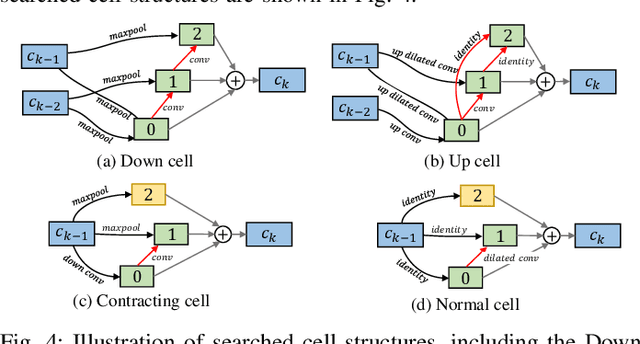
Abstract:Automatic myocardial segmentation of contrast echocardiography has shown great potential in the quantification of myocardial perfusion parameters. Segmentation quality control is an important step to ensure the accuracy of segmentation results for quality research as well as its clinical application. Usually, the segmentation quality control happens after the data acquisition. At the data acquisition time, the operator could not know the quality of the segmentation results. On-the-fly segmentation quality control could help the operator to adjust the ultrasound probe or retake data if the quality is unsatisfied, which can greatly reduce the effort of time-consuming manual correction. However, it is infeasible to deploy state-of-the-art DNN-based models because the segmentation module and quality control module must fit in the limited hardware resource on the ultrasound machine while satisfying strict latency constraints. In this paper, we propose a hardware-aware neural architecture search framework for automatic myocardial segmentation and quality control of contrast echocardiography. We explicitly incorporate the hardware latency as a regularization term into the loss function during training. The proposed method searches the best neural network architecture for the segmentation module and quality prediction module with strict latency.
RADARS: Memory Efficient Reinforcement Learning Aided Differentiable Neural Architecture Search
Sep 13, 2021



Abstract:Differentiable neural architecture search (DNAS) is known for its capacity in the automatic generation of superior neural networks. However, DNAS based methods suffer from memory usage explosion when the search space expands, which may prevent them from running successfully on even advanced GPU platforms. On the other hand, reinforcement learning (RL) based methods, while being memory efficient, are extremely time-consuming. Combining the advantages of both types of methods, this paper presents RADARS, a scalable RL-aided DNAS framework that can explore large search spaces in a fast and memory-efficient manner. RADARS iteratively applies RL to prune undesired architecture candidates and identifies a promising subspace to carry out DNAS. Experiments using a workstation with 12 GB GPU memory show that on CIFAR-10 and ImageNet datasets, RADARS can achieve up to 3.41% higher accuracy with 2.5X search time reduction compared with a state-of-the-art RL-based method, while the two DNAS baselines cannot complete due to excessive memory usage or search time. To the best of the authors' knowledge, this is the first DNAS framework that can handle large search spaces with bounded memory usage.
Exploration of Quantum Neural Architecture by Mixing Quantum Neuron Designs
Sep 08, 2021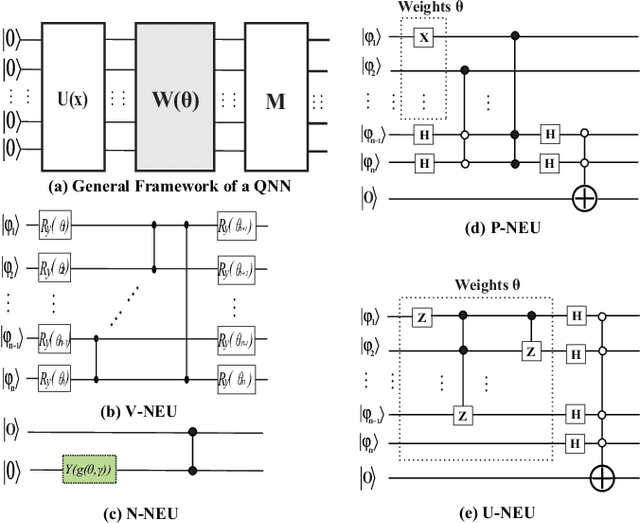
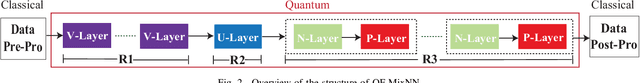
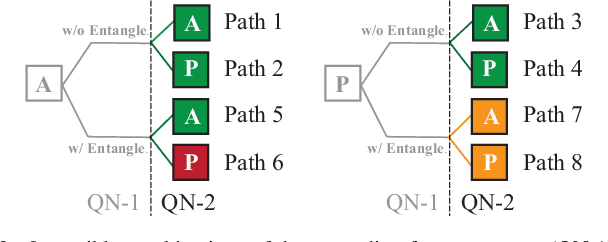
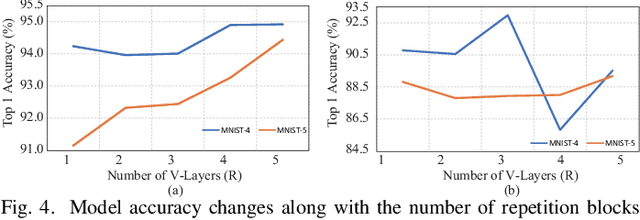
Abstract:With the constant increase of the number of quantum bits (qubits) in the actual quantum computers, implementing and accelerating the prevalent deep learning on quantum computers are becoming possible. Along with this trend, there emerge quantum neural architectures based on different designs of quantum neurons. A fundamental question in quantum deep learning arises: what is the best quantum neural architecture? Inspired by the design of neural architectures for classical computing which typically employs multiple types of neurons, this paper makes the very first attempt to mix quantum neuron designs to build quantum neural architectures. We observe that the existing quantum neuron designs may be quite different but complementary, such as neurons from variation quantum circuits (VQC) and Quantumflow. More specifically, VQC can apply real-valued weights but suffer from being extended to multiple layers, while QuantumFlow can build a multi-layer network efficiently, but is limited to use binary weights. To take their respective advantages, we propose to mix them together and figure out a way to connect them seamlessly without additional costly measurement. We further investigate the design principles to mix quantum neurons, which can provide guidance for quantum neural architecture exploration in the future. Experimental results demonstrate that the identified quantum neural architectures with mixed quantum neurons can achieve 90.62% of accuracy on the MNIST dataset, compared with 52.77% and 69.92% on the VQC and QuantumFlow, respectively.
Can Noise on Qubits Be Learned in Quantum Neural Network? A Case Study on QuantumFlow
Sep 08, 2021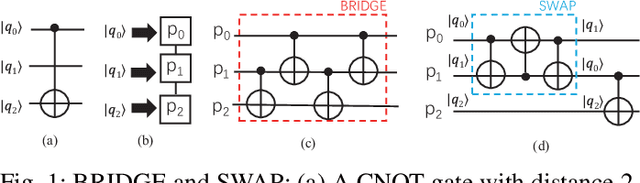
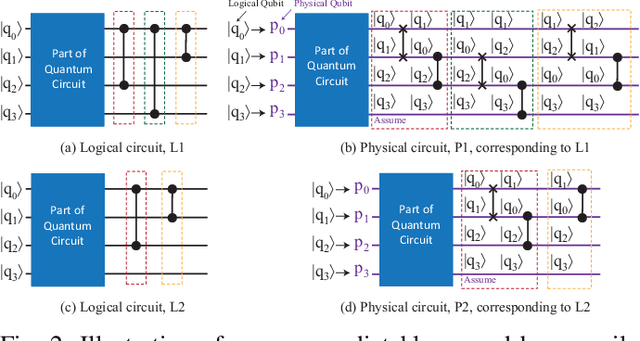
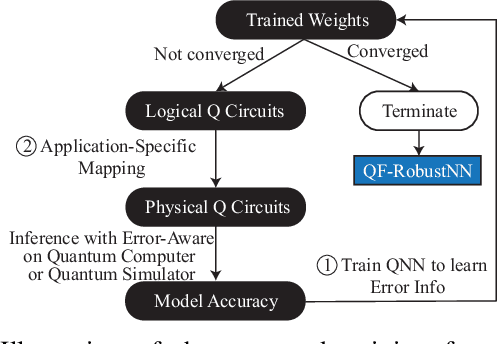
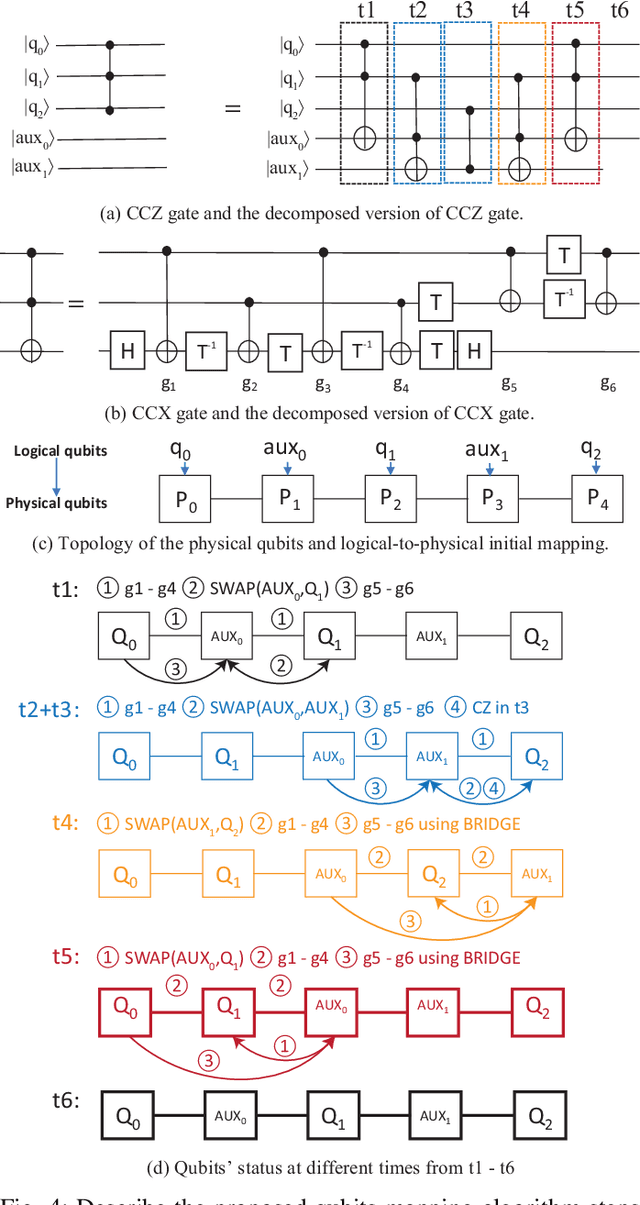
Abstract:In the noisy intermediate-scale quantum (NISQ) era, one of the key questions is how to deal with the high noise level existing in physical quantum bits (qubits). Quantum error correction is promising but requires an extensive number (e.g., over 1,000) of physical qubits to create one "perfect" qubit, exceeding the capacity of the existing quantum computers. This paper aims to tackle the noise issue from another angle: instead of creating perfect qubits for general quantum algorithms, we investigate the potential to mitigate the noise issue for dedicate algorithms. Specifically, this paper targets quantum neural network (QNN), and proposes to learn the errors in the training phase, so that the identified QNN model can be resilient to noise. As a result, the implementation of QNN needs no or a small number of additional physical qubits, which is more realistic for the near-term quantum computers. To achieve this goal, an application-specific compiler is essential: on the one hand, the error cannot be learned if the mapping from logical qubits to physical qubits exists randomness; on the other hand, the compiler needs to be efficient so that the lengthy training procedure can be completed in a reasonable time. In this paper, we utilize the recent QNN framework, QuantumFlow, as a case study. Experimental results show that the proposed approach can optimize QNN models for different errors in qubits, achieving up to 28% accuracy improvement compared with the model obtained by the error-agnostic training.
ImageTBAD: A 3D Computed Tomography Angiography Image Dataset for Automatic Segmentation of Type-B Aortic Dissection
Sep 01, 2021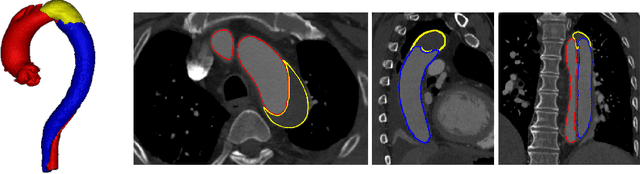
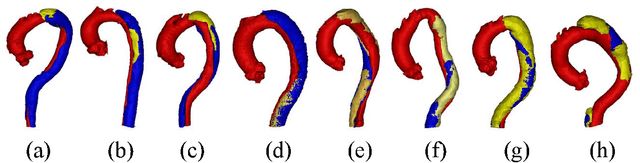
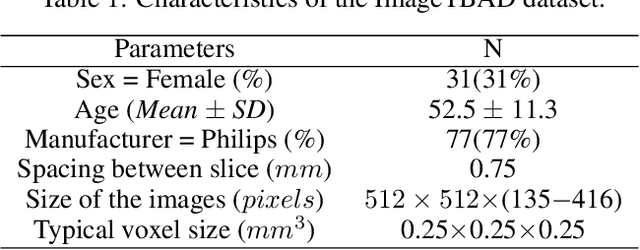

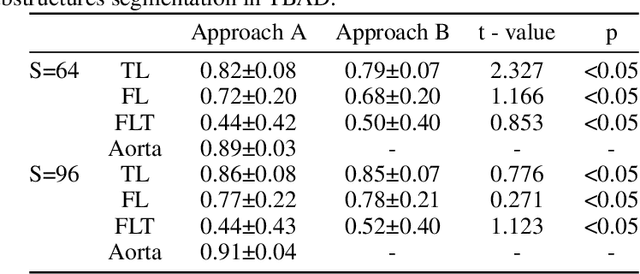
Abstract:Type-B Aortic Dissection (TBAD) is one of the most serious cardiovascular events characterized by a growing yearly incidence,and the severity of disease prognosis. Currently, computed tomography angiography (CTA) has been widely adopted for the diagnosis and prognosis of TBAD. Accurate segmentation of true lumen (TL), false lumen (FL), and false lumen thrombus (FLT) in CTA are crucial for the precise quantification of anatomical features. However, existing works only focus on only TL and FL without considering FLT. In this paper, we propose ImageTBAD, the first 3D computed tomography angiography (CTA) image dataset of TBAD with annotation of TL, FL, and FLT. The proposed dataset contains 100 TBAD CTA images, which is of decent size compared with existing medical imaging datasets. As FLT can appear almost anywhere along the aorta with irregular shapes, segmentation of FLT presents a wide class of segmentation problems where targets exist in a variety of positions with irregular shapes. We further propose a baseline method for automatic segmentation of TBAD. Results show that the baseline method can achieve comparable results with existing works on aorta and TL segmentation. However, the segmentation accuracy of FLT is only 52%, which leaves large room for improvement and also shows the challenge of our dataset. To facilitate further research on this challenging problem, our dataset and codes are released to the public.
Uncertainty Modeling of Emerging Device-based Computing-in-Memory Neural Accelerators with Application to Neural Architecture Search
Jul 06, 2021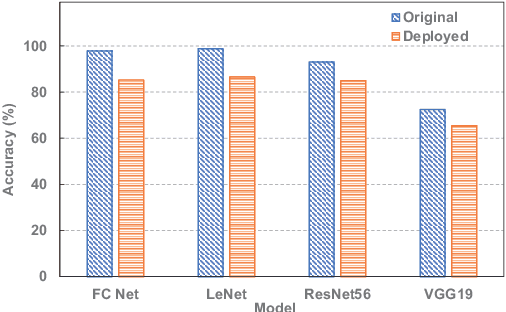
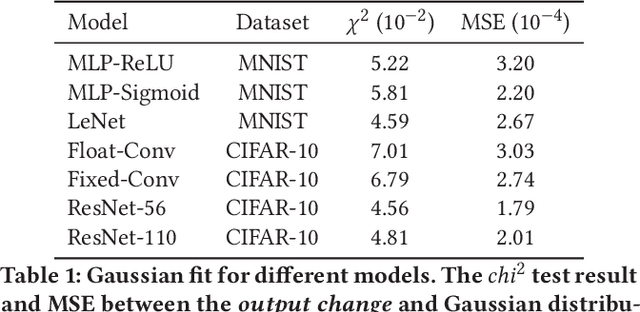
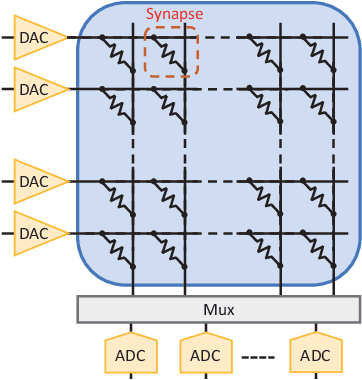
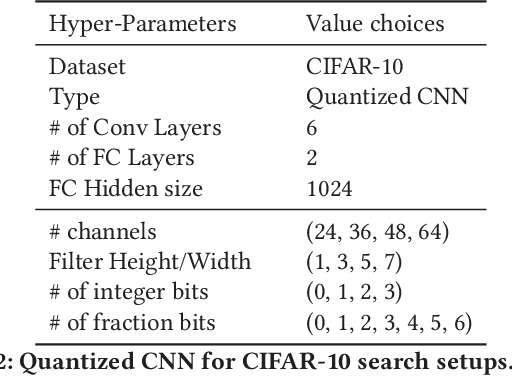
Abstract:Emerging device-based Computing-in-memory (CiM) has been proved to be a promising candidate for high-energy efficiency deep neural network (DNN) computations. However, most emerging devices suffer uncertainty issues, resulting in a difference between actual data stored and the weight value it is designed to be. This leads to an accuracy drop from trained models to actually deployed platforms. In this work, we offer a thorough analysis of the effect of such uncertainties-induced changes in DNN models. To reduce the impact of device uncertainties, we propose UAE, an uncertainty-aware Neural Architecture Search scheme to identify a DNN model that is both accurate and robust against device uncertainties.
Segmentation with Multiple Acceptable Annotations: A Case Study of Myocardial Segmentation in Contrast Echocardiography
Jun 29, 2021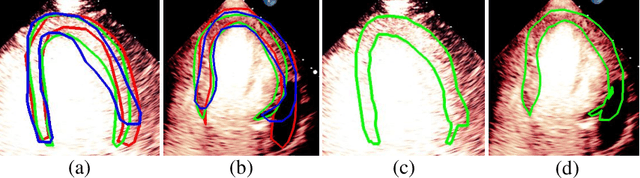
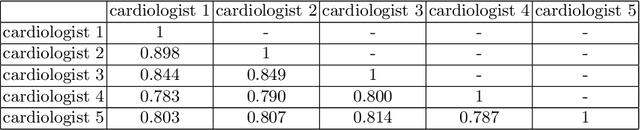
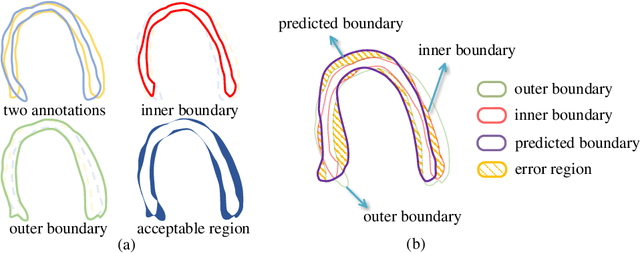
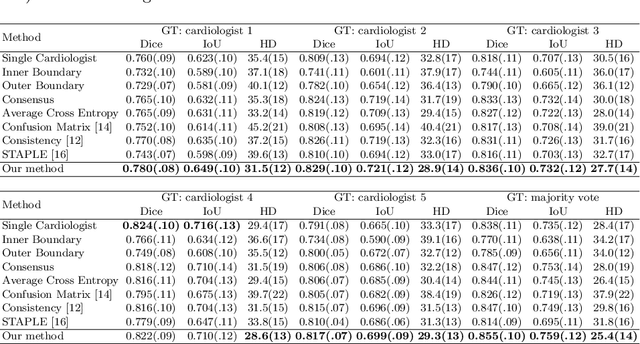
Abstract:Most existing deep learning-based frameworks for image segmentation assume that a unique ground truth is known and can be used for performance evaluation. This is true for many applications, but not all. Myocardial segmentation of Myocardial Contrast Echocardiography (MCE), a critical task in automatic myocardial perfusion analysis, is an example. Due to the low resolution and serious artifacts in MCE data, annotations from different cardiologists can vary significantly, and it is hard to tell which one is the best. In this case, how can we find a good way to evaluate segmentation performance and how do we train the neural network? In this paper, we address the first problem by proposing a new extended Dice to effectively evaluate the segmentation performance when multiple accepted ground truth is available. Then based on our proposed metric, we solve the second problem by further incorporating the new metric into a loss function that enables neural networks to flexibly learn general features of myocardium. Experiment results on our clinical MCE data set demonstrate that the neural network trained with the proposed loss function outperforms those existing ones that try to obtain a unique ground truth from multiple annotations, both quantitatively and qualitatively. Finally, our grading study shows that using extended Dice as an evaluation metric can better identify segmentation results that need manual correction compared with using Dice.
Positional Contrastive Learning for Volumetric Medical Image Segmentation
Jun 18, 2021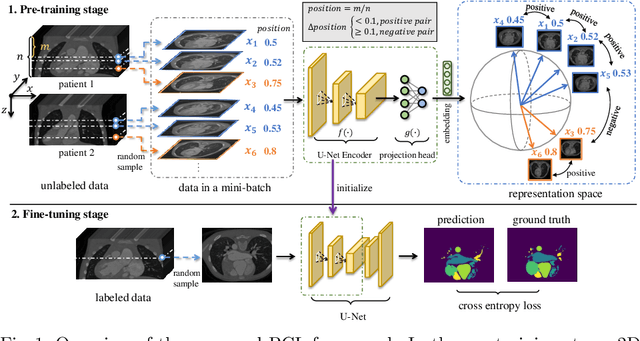
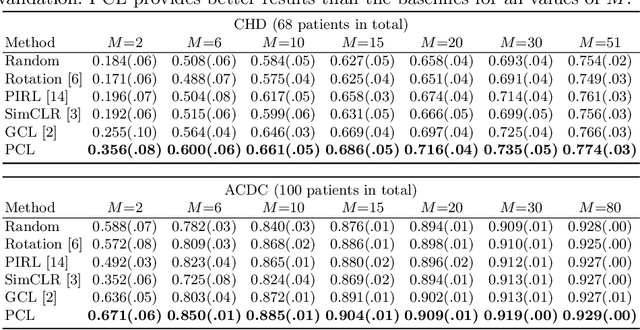
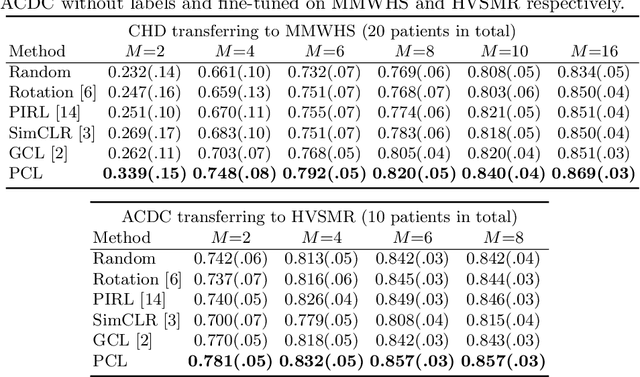
Abstract:The success of deep learning heavily depends on the availability of large labeled training sets. However, it is hard to get large labeled datasets in medical image domain because of the strict privacy concern and costly labeling efforts. Contrastive learning, an unsupervised learning technique, has been proved powerful in learning image-level representations from unlabeled data. The learned encoder can then be transferred or fine-tuned to improve the performance of downstream tasks with limited labels. A critical step in contrastive learning is the generation of contrastive data pairs, which is relatively simple for natural image classification but quite challenging for medical image segmentation due to the existence of the same tissue or organ across the dataset. As a result, when applied to medical image segmentation, most state-of-the-art contrastive learning frameworks inevitably introduce a lot of false-negative pairs and result in degraded segmentation quality. To address this issue, we propose a novel positional contrastive learning (PCL) framework to generate contrastive data pairs by leveraging the position information in volumetric medical images. Experimental results on CT and MRI datasets demonstrate that the proposed PCL method can substantially improve the segmentation performance compared to existing methods in both semi-supervised setting and transfer learning setting.
Enabling On-Device Self-Supervised Contrastive Learning With Selective Data Contrast
Jun 07, 2021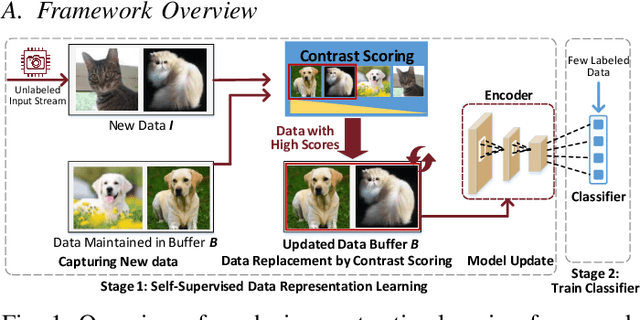
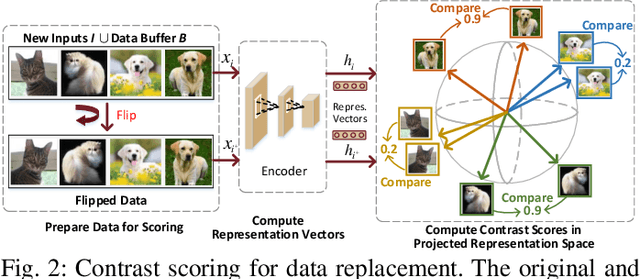
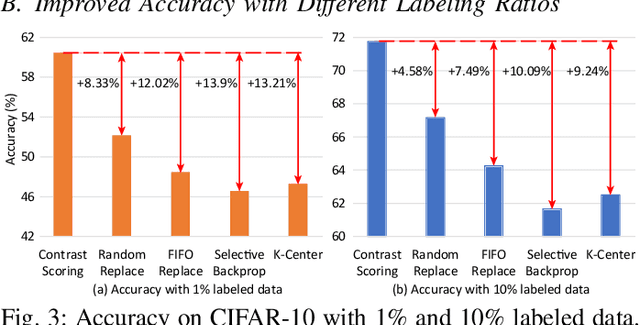
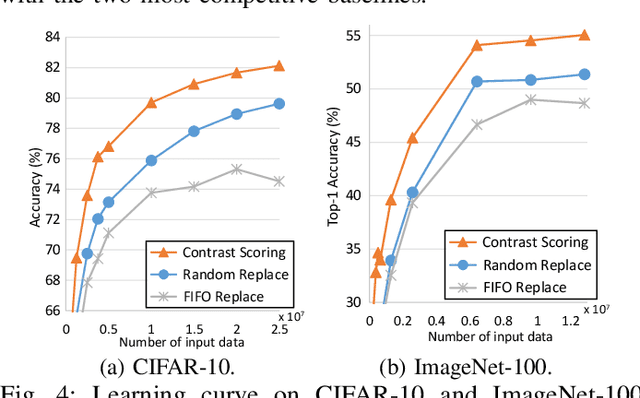
Abstract:After a model is deployed on edge devices, it is desirable for these devices to learn from unlabeled data to continuously improve accuracy. Contrastive learning has demonstrated its great potential in learning from unlabeled data. However, the online input data are usually none independent and identically distributed (non-iid) and storages of edge devices are usually too limited to store enough representative data from different data classes. We propose a framework to automatically select the most representative data from the unlabeled input stream, which only requires a small data buffer for dynamic learning. Experiments show that accuracy and learning speed are greatly improved.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge