Yiyu Shi
A Semi-Decoupled Approach to Fast and Optimal Hardware-Software Co-Design of Neural Accelerators
Mar 25, 2022
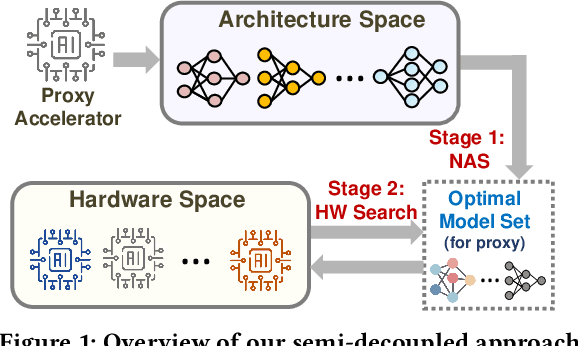
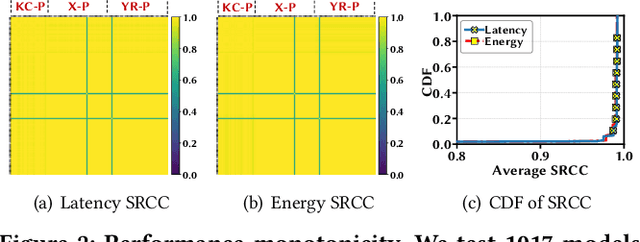
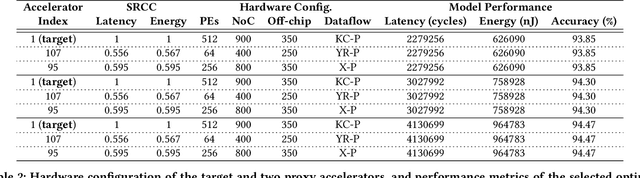
Abstract:In view of the performance limitations of fully-decoupled designs for neural architectures and accelerators, hardware-software co-design has been emerging to fully reap the benefits of flexible design spaces and optimize neural network performance. Nonetheless, such co-design also enlarges the total search space to practically infinity and presents substantial challenges. While the prior studies have been focusing on improving the search efficiency (e.g., via reinforcement learning), they commonly rely on co-searches over the entire architecture-accelerator design space. In this paper, we propose a \emph{semi}-decoupled approach to reduce the size of the total design space by orders of magnitude, yet without losing optimality. We first perform neural architecture search to obtain a small set of optimal architectures for one accelerator candidate. Importantly, this is also the set of (close-to-)optimal architectures for other accelerator designs based on the property that neural architectures' ranking orders in terms of inference latency and energy consumption on different accelerator designs are highly similar. Then, instead of considering all the possible architectures, we optimize the accelerator design only in combination with this small set of architectures, thus significantly reducing the total search cost. We validate our approach by conducting experiments on various architecture spaces for accelerator designs with different dataflows. Our results highlight that we can obtain the optimal design by only navigating over the reduced search space. The source code of this work is at \url{https://github.com/Ren-Research/CoDesign}.
FairPrune: Achieving Fairness Through Pruning for Dermatological Disease Diagnosis
Mar 04, 2022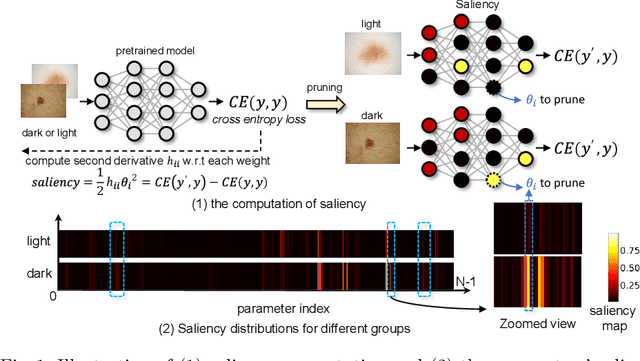
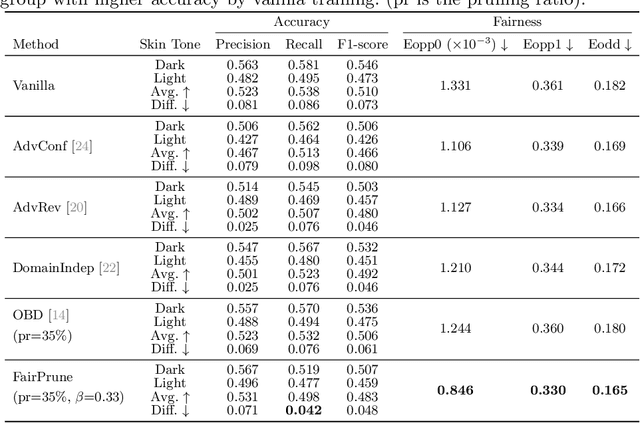
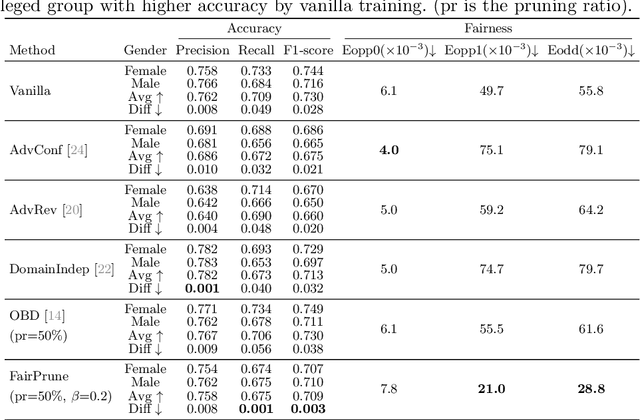
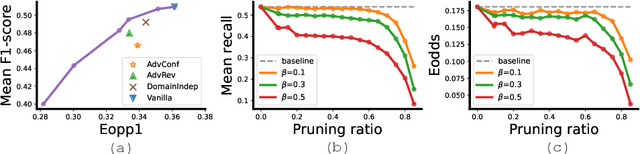
Abstract:Many works have shown that deep learning-based medical image classification models can exhibit bias toward certain demographic attributes like race, gender, and age. Existing bias mitigation methods primarily focus on learning debiased models, which may not necessarily guarantee all sensitive information can be removed and usually comes with considerable accuracy degradation on both privileged and unprivileged groups. To tackle this issue, we propose a method, FairPrune, that achieves fairness by pruning. Conventionally, pruning is used to reduce the model size for efficient inference. However, we show that pruning can also be a powerful tool to achieve fairness. Our observation is that during pruning, each parameter in the model has different importance for different groups' accuracy. By pruning the parameters based on this importance difference, we can reduce the accuracy difference between the privileged group and the unprivileged group to improve fairness without a large accuracy drop. To this end, we use the second derivative of the parameters of a pre-trained model to quantify the importance of each parameter with respect to the model accuracy for each group. Experiments on two skin lesion diagnosis datasets over multiple sensitive attributes demonstrate that our method can greatly improve fairness while keeping the average accuracy of both groups as high as possible.
The Larger The Fairer? Small Neural Networks Can Achieve Fairness for Edge Devices
Feb 23, 2022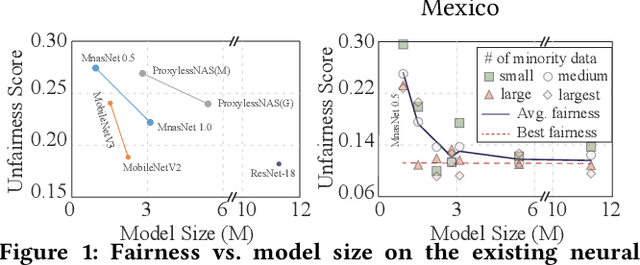
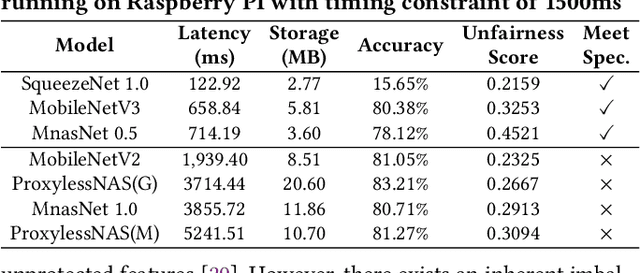
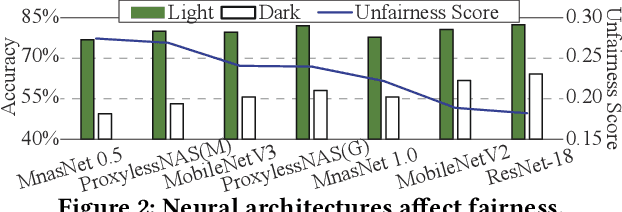
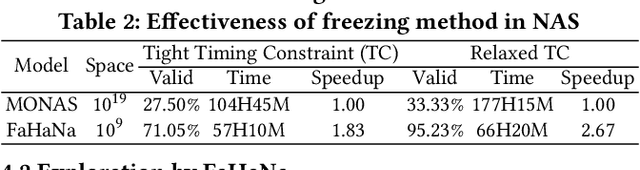
Abstract:Along with the progress of AI democratization, neural networks are being deployed more frequently in edge devices for a wide range of applications. Fairness concerns gradually emerge in many applications, such as face recognition and mobile medical. One fundamental question arises: what will be the fairest neural architecture for edge devices? By examining the existing neural networks, we observe that larger networks typically are fairer. But, edge devices call for smaller neural architectures to meet hardware specifications. To address this challenge, this work proposes a novel Fairness- and Hardware-aware Neural architecture search framework, namely FaHaNa. Coupled with a model freezing approach, FaHaNa can efficiently search for neural networks with balanced fairness and accuracy, while guaranteed to meet hardware specifications. Results show that FaHaNa can identify a series of neural networks with higher fairness and accuracy on a dermatology dataset. Target edge devices, FaHaNa finds a neural architecture with slightly higher accuracy, 5.28x smaller size, 15.14% higher fairness score, compared with MobileNetV2; meanwhile, on Raspberry PI and Odroid XU-4, it achieves 5.75x and 5.79x speedup.
SWIM: Selective Write-Verify for Computing-in-Memory Neural Accelerators
Feb 17, 2022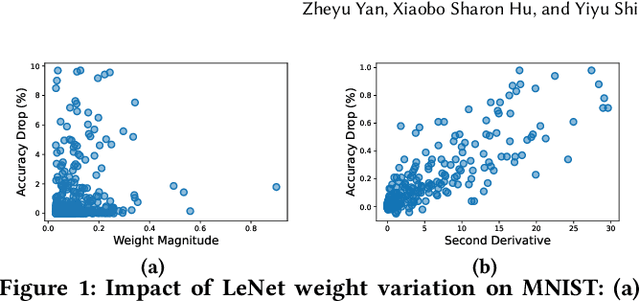
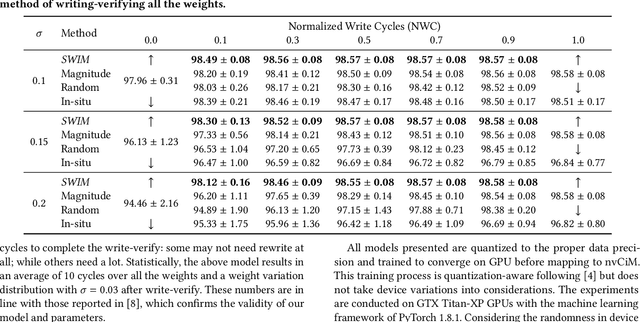
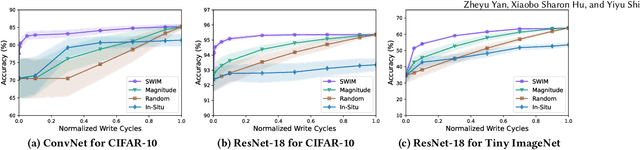
Abstract:Computing-in-Memory architectures based on non-volatile emerging memories have demonstrated great potential for deep neural network (DNN) acceleration thanks to their high energy efficiency. However, these emerging devices can suffer from significant variations during the mapping process i.e., programming weights to the devices), and if left undealt with, can cause significant accuracy degradation. The non-ideality of weight mapping can be compensated by iterative programming with a write-verify scheme, i.e., reading the conductance and rewriting if necessary. In all existing works, such a practice is applied to every single weight of a DNN as it is being mapped, which requires extensive programming time. In this work, we show that it is only necessary to select a small portion of the weights for write-verify to maintain the DNN accuracy, thus achieving significant speedup. We further introduce a second derivative based technique SWIM, which only requires a single pass of forward and backpropagation, to efficiently select the weights that need write-verify. Experimental results on various DNN architectures for different datasets show that SWIM can achieve up to 10x programming speedup compared with conventional full-blown write-verify while attaining a comparable accuracy.
Learn by Challenging Yourself: Contrastive Visual Representation Learning with Hard Sample Generation
Feb 14, 2022Abstract:Contrastive learning (CL), a self-supervised learning approach, can effectively learn visual representations from unlabeled data. However, CL requires learning on vast quantities of diverse data to achieve good performance, without which the performance of CL will greatly degrade. To tackle this problem, we propose a framework with two approaches to improve the data efficiency of CL training by generating beneficial samples and joint learning. The first approach generates hard samples for the main model. The generator is jointly learned with the main model to dynamically customize hard samples based on the training state of the main model. With the progressively growing knowledge of the main model, the generated samples also become harder to constantly encourage the main model to learn better representations. Besides, a pair of data generators are proposed to generate similar but distinct samples as positive pairs. In joint learning, the hardness of a positive pair is progressively increased by decreasing their similarity. In this way, the main model learns to cluster hard positives by pulling the representations of similar yet distinct samples together, by which the representations of similar samples are well-clustered and better representations can be learned. Comprehensive experiments show superior accuracy and data efficiency of the proposed methods over the state-of-the-art on multiple datasets. For example, about 5% accuracy improvement on ImageNet-100 and CIFAR-10, and more than 6% accuracy improvement on CIFAR-100 are achieved for linear classification. Besides, up to 2x data efficiency for linear classification and up to 5x data efficiency for transfer learning are achieved.
Federated Contrastive Learning for Dermatological Disease Diagnosis via On-device Learning
Feb 14, 2022



Abstract:Deep learning models have been deployed in an increasing number of edge and mobile devices to provide healthcare. These models rely on training with a tremendous amount of labeled data to achieve high accuracy. However, for medical applications such as dermatological disease diagnosis, the private data collected by mobile dermatology assistants exist on distributed mobile devices of patients, and each device only has a limited amount of data. Directly learning from limited data greatly deteriorates the performance of learned models. Federated learning (FL) can train models by using data distributed on devices while keeping the data local for privacy. Existing works on FL assume all the data have ground-truth labels. However, medical data often comes without any accompanying labels since labeling requires expertise and results in prohibitively high labor costs. The recently developed self-supervised learning approach, contrastive learning (CL), can leverage the unlabeled data to pre-train a model, after which the model is fine-tuned on limited labeled data for dermatological disease diagnosis. However, simply combining CL with FL as federated contrastive learning (FCL) will result in ineffective learning since CL requires diverse data for learning but each device only has limited data. In this work, we propose an on-device FCL framework for dermatological disease diagnosis with limited labels. Features are shared in the FCL pre-training process to provide diverse and accurate contrastive information. After that, the pre-trained model is fine-tuned with local labeled data independently on each device or collaboratively with supervised federated learning on all devices. Experiments on dermatological disease datasets show that the proposed framework effectively improves the recall and precision of dermatological disease diagnosis compared with state-of-the-art methods.
Distributed Unsupervised Visual Representation Learning with Fused Features
Nov 21, 2021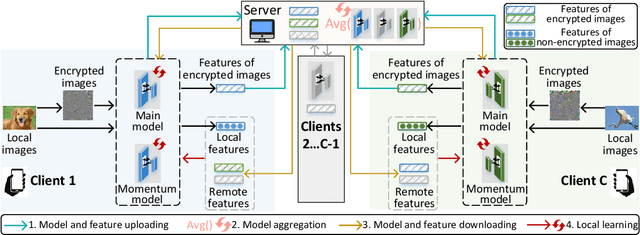
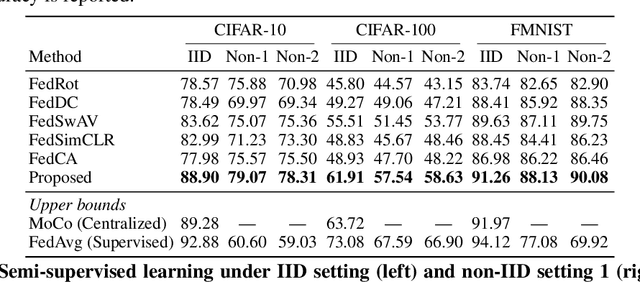
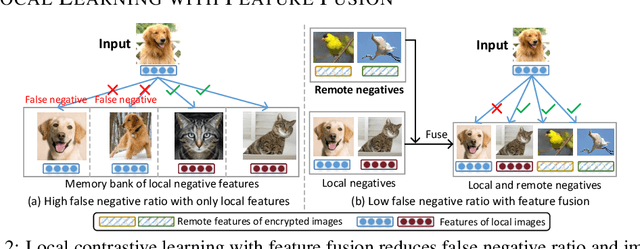
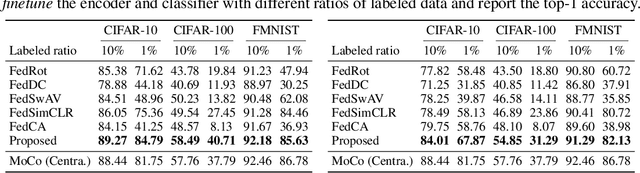
Abstract:Federated learning (FL) enables distributed clients to learn a shared model for prediction while keeping the training data local on each client. However, existing FL requires fully-labeled data for training, which is inconvenient or sometimes infeasible to obtain due to the high labeling cost and the requirement of expertise. The lack of labels makes FL impractical in many realistic settings. Self-supervised learning can address this challenge by learning from unlabeled data such that FL can be widely used. Contrastive learning (CL), a self-supervised learning approach, can effectively learn data representations from unlabeled data. However, the distributed data collected on clients are usually not independent and identically distributed (non-IID) among clients, and each client may only have few classes of data, which degrades the performance of CL and learned representations. To tackle this problem, we propose a federated contrastive learning framework consisting of two approaches: feature fusion and neighborhood matching, by which a unified feature space among clients is learned for better data representations. Feature fusion provides remote features as accurate contrastive information to each client for better local learning. Neighborhood matching further aligns each client's local features to the remote features such that well-clustered features among clients can be learned. Extensive experiments show the effectiveness of the proposed framework. It outperforms other methods by 11\% on IID data and matches the performance of centralized learning.
One Proxy Device Is Enough for Hardware-Aware Neural Architecture Search
Nov 03, 2021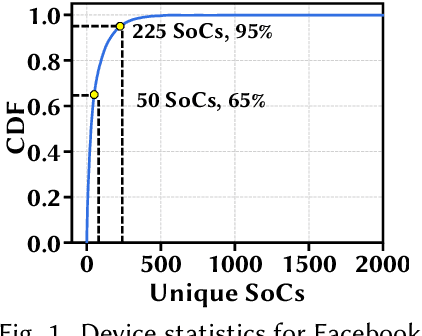
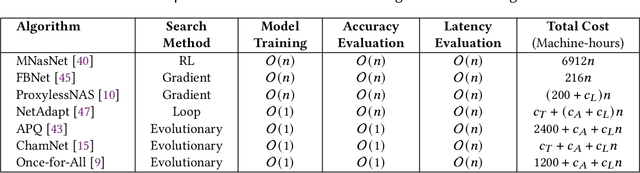
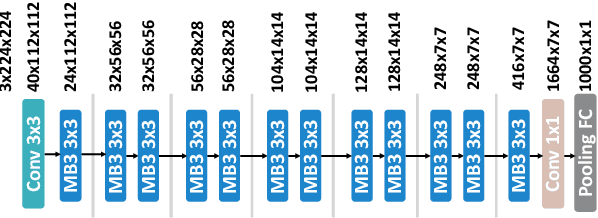
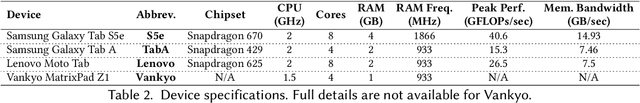
Abstract:Convolutional neural networks (CNNs) are used in numerous real-world applications such as vision-based autonomous driving and video content analysis. To run CNN inference on various target devices, hardware-aware neural architecture search (NAS) is crucial. A key requirement of efficient hardware-aware NAS is the fast evaluation of inference latencies in order to rank different architectures. While building a latency predictor for each target device has been commonly used in state of the art, this is a very time-consuming process, lacking scalability in the presence of extremely diverse devices. In this work, we address the scalability challenge by exploiting latency monotonicity -- the architecture latency rankings on different devices are often correlated. When strong latency monotonicity exists, we can re-use architectures searched for one proxy device on new target devices, without losing optimality. In the absence of strong latency monotonicity, we propose an efficient proxy adaptation technique to significantly boost the latency monotonicity. Finally, we validate our approach and conduct experiments with devices of different platforms on multiple mainstream search spaces, including MobileNet-V2, MobileNet-V3, NAS-Bench-201, ProxylessNAS and FBNet. Our results highlight that, by using just one proxy device, we can find almost the same Pareto-optimal architectures as the existing per-device NAS, while avoiding the prohibitive cost of building a latency predictor for each device. GitHub: https://github.com/Ren-Research/OneProxy
* Accepted by the ACM SIGMETRICS 2022. Published in the Proceedings of the ACM on Measurement and Analysis of Computing Systems, vol. 5, no. 3, Article 34, December 2021. GitHub: https://github.com/Ren-Research/OneProxy
"One-Shot" Reduction of Additive Artifacts in Medical Images
Oct 23, 2021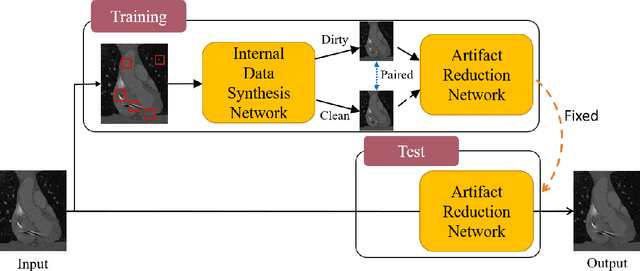
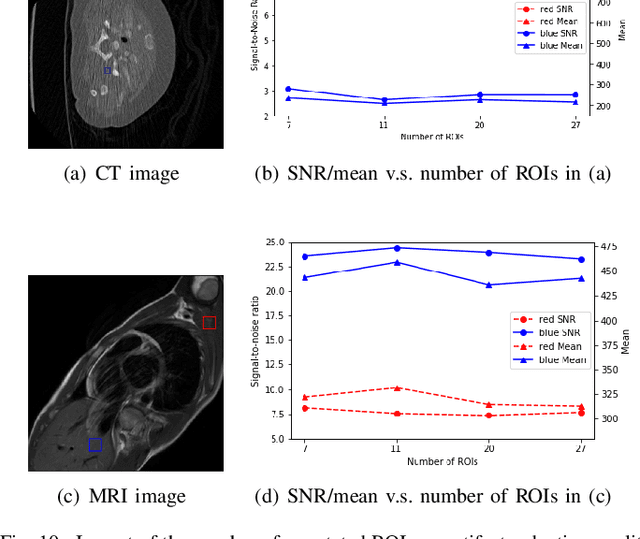
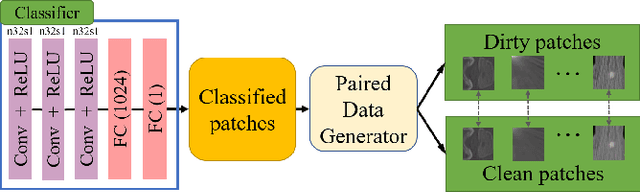

Abstract:Medical images may contain various types of artifacts with different patterns and mixtures, which depend on many factors such as scan setting, machine condition, patients' characteristics, surrounding environment, etc. However, existing deep-learning-based artifact reduction methods are restricted by their training set with specific predetermined artifact types and patterns. As such, they have limited clinical adoption. In this paper, we introduce One-Shot medical image Artifact Reduction (OSAR), which exploits the power of deep learning but without using pre-trained general networks. Specifically, we train a light-weight image-specific artifact reduction network using data synthesized from the input image at test-time. Without requiring any prior large training data set, OSAR can work with almost any medical images that contain varying additive artifacts which are not in any existing data sets. In addition, Computed Tomography (CT) and Magnetic Resonance Imaging (MRI) are used as vehicles and show that the proposed method can reduce artifacts better than state-of-the-art both qualitatively and quantitatively using shorter test time.
Semi-supervised Contrastive Learning for Label-efficient Medical Image Segmentation
Sep 28, 2021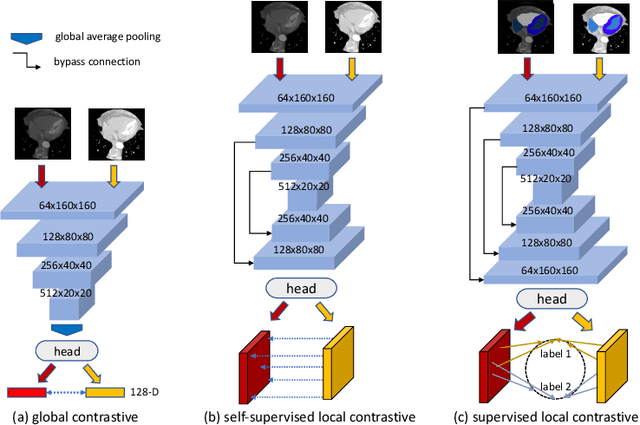
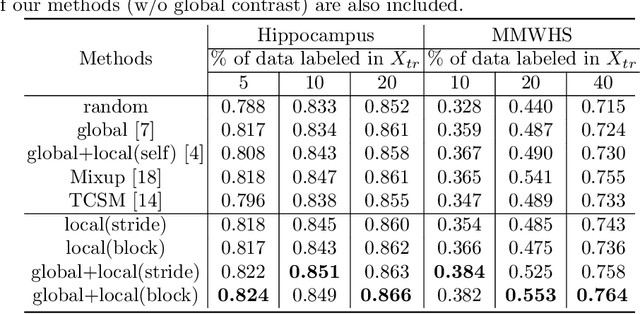
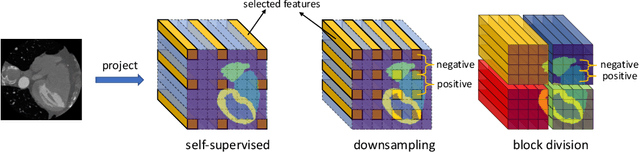
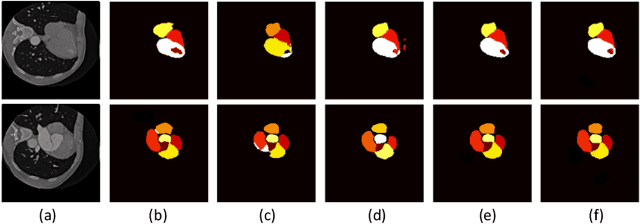
Abstract:The success of deep learning methods in medical image segmentation tasks heavily depends on a large amount of labeled data to supervise the training. On the other hand, the annotation of biomedical images requires domain knowledge and can be laborious. Recently, contrastive learning has demonstrated great potential in learning latent representation of images even without any label. Existing works have explored its application to biomedical image segmentation where only a small portion of data is labeled, through a pre-training phase based on self-supervised contrastive learning without using any labels followed by a supervised fine-tuning phase on the labeled portion of data only. In this paper, we establish that by including the limited label in formation in the pre-training phase, it is possible to boost the performance of contrastive learning. We propose a supervised local contrastive loss that leverages limited pixel-wise annotation to force pixels with the same label to gather around in the embedding space. Such loss needs pixel-wise computation which can be expensive for large images, and we further propose two strategies, downsampling and block division, to address the issue. We evaluate our methods on two public biomedical image datasets of different modalities. With different amounts of labeled data, our methods consistently outperform the state-of-the-art contrast-based methods and other semi-supervised learning techniques.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge