Yiqiu Shen
Superclass-Guided Representation Disentanglement for Spurious Correlation Mitigation
Aug 12, 2025



Abstract:To enhance group robustness to spurious correlations, prior work often relies on auxiliary annotations for groups or spurious features and assumes identical sets of groups across source and target domains. These two requirements are both unnatural and impractical in real-world settings. To overcome these limitations, we propose a method that leverages the semantic structure inherent in class labels--specifically, superclass information--to naturally reduce reliance on spurious features. Our model employs gradient-based attention guided by a pre-trained vision-language model to disentangle superclass-relevant and irrelevant features. Then, by promoting the use of all superclass-relevant features for prediction, our approach achieves robustness to more complex spurious correlations without the need to annotate any source samples. Experiments across diverse datasets demonstrate that our method significantly outperforms baselines in domain generalization tasks, with clear improvements in both quantitative metrics and qualitative visualizations.
Fine-Tuning In-House Large Language Models to Infer Differential Diagnosis from Radiology Reports
Oct 11, 2024



Abstract:Radiology reports summarize key findings and differential diagnoses derived from medical imaging examinations. The extraction of differential diagnoses is crucial for downstream tasks, including patient management and treatment planning. However, the unstructured nature of these reports, characterized by diverse linguistic styles and inconsistent formatting, presents significant challenges. Although proprietary large language models (LLMs) such as GPT-4 can effectively retrieve clinical information, their use is limited in practice by high costs and concerns over the privacy of protected health information (PHI). This study introduces a pipeline for developing in-house LLMs tailored to identify differential diagnoses from radiology reports. We first utilize GPT-4 to create 31,056 labeled reports, then fine-tune open source LLM using this dataset. Evaluated on a set of 1,067 reports annotated by clinicians, the proposed model achieves an average F1 score of 92.1\%, which is on par with GPT-4 (90.8\%). Through this study, we provide a methodology for constructing in-house LLMs that: match the performance of GPT, reduce dependence on expensive proprietary models, and enhance the privacy and security of PHI.
BURExtract-Llama: An LLM for Clinical Concept Extraction in Breast Ultrasound Reports
Aug 21, 2024



Abstract:Breast ultrasound is essential for detecting and diagnosing abnormalities, with radiology reports summarizing key findings like lesion characteristics and malignancy assessments. Extracting this critical information is challenging due to the unstructured nature of these reports, with varied linguistic styles and inconsistent formatting. While proprietary LLMs like GPT-4 are effective, they are costly and raise privacy concerns when handling protected health information. This study presents a pipeline for developing an in-house LLM to extract clinical information from radiology reports. We first use GPT-4 to create a small labeled dataset, then fine-tune a Llama3-8B model on it. Evaluated on clinician-annotated reports, our model achieves an average F1 score of 84.6%, which is on par with GPT-4. Our findings demonstrate the feasibility of developing an in-house LLM that not only matches GPT-4's performance but also offers cost reductions and enhanced data privacy.
Understanding differences in applying DETR to natural and medical images
May 27, 2024Abstract:Transformer-based detectors have shown success in computer vision tasks with natural images. These models, exemplified by the Deformable DETR, are optimized through complex engineering strategies tailored to the typical characteristics of natural scenes. However, medical imaging data presents unique challenges such as extremely large image sizes, fewer and smaller regions of interest, and object classes which can be differentiated only through subtle differences. This study evaluates the applicability of these transformer-based design choices when applied to a screening mammography dataset that represents these distinct medical imaging data characteristics. Our analysis reveals that common design choices from the natural image domain, such as complex encoder architectures, multi-scale feature fusion, query initialization, and iterative bounding box refinement, do not improve and sometimes even impair object detection performance in medical imaging. In contrast, simpler and shallower architectures often achieve equal or superior results. This finding suggests that the adaptation of transformer models for medical imaging data requires a reevaluation of standard practices, potentially leading to more efficient and specialized frameworks for medical diagnosis.
Leveraging Transformers to Improve Breast Cancer Classification and Risk Assessment with Multi-modal and Longitudinal Data
Nov 15, 2023



Abstract:Breast cancer screening, primarily conducted through mammography, is often supplemented with ultrasound for women with dense breast tissue. However, existing deep learning models analyze each modality independently, missing opportunities to integrate information across imaging modalities and time. In this study, we present Multi-modal Transformer (MMT), a neural network that utilizes mammography and ultrasound synergistically, to identify patients who currently have cancer and estimate the risk of future cancer for patients who are currently cancer-free. MMT aggregates multi-modal data through self-attention and tracks temporal tissue changes by comparing current exams to prior imaging. Trained on 1.3 million exams, MMT achieves an AUROC of 0.943 in detecting existing cancers, surpassing strong uni-modal baselines. For 5-year risk prediction, MMT attains an AUROC of 0.826, outperforming prior mammography-based risk models. Our research highlights the value of multi-modal and longitudinal imaging in cancer diagnosis and risk stratification.
Multiple Instance Learning via Iterative Self-Paced Supervised Contrastive Learning
Oct 17, 2022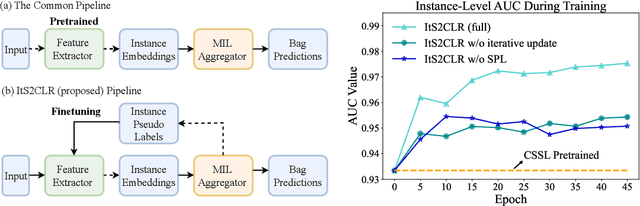
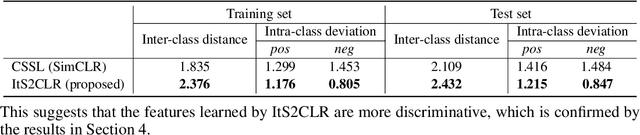
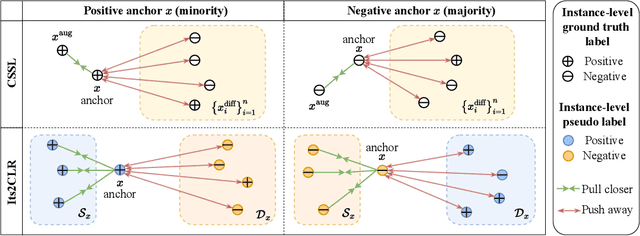

Abstract:Learning representations for individual instances when only bag-level labels are available is a fundamental challenge in multiple instance learning (MIL). Recent works have shown promising results using contrastive self-supervised learning (CSSL), which learns to push apart representations corresponding to two different randomly-selected instances. Unfortunately, in real-world applications such as medical image classification, there is often class imbalance, so randomly-selected instances mostly belong to the same majority class, which precludes CSSL from learning inter-class differences. To address this issue, we propose a novel framework, Iterative Self-paced Supervised Contrastive Learning for MIL Representations (ItS2CLR), which improves the learned representation by exploiting instance-level pseudo labels derived from the bag-level labels. The framework employs a novel self-paced sampling strategy to ensure the accuracy of pseudo labels. We evaluate ItS2CLR on three medical datasets, showing that it improves the quality of instance-level pseudo labels and representations, and outperforms existing MIL methods in terms of both bag and instance level accuracy.
Minimax Supervised Clustering in the Anisotropic Gaussian Mixture Model: A new take on Robust Interpolation
Nov 13, 2021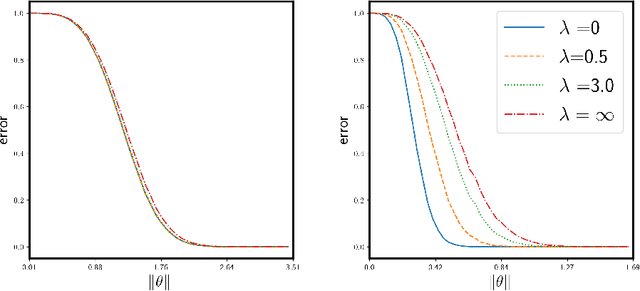
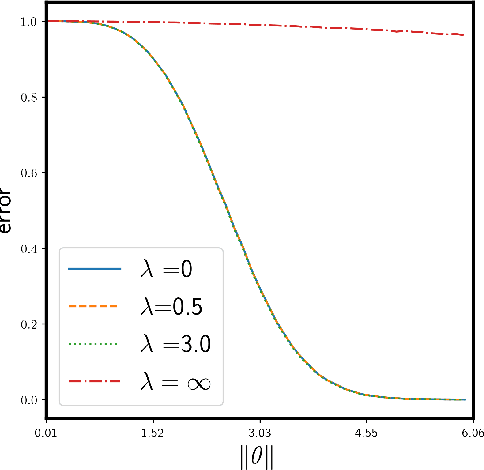
Abstract:We study the supervised clustering problem under the two-component anisotropic Gaussian mixture model in high dimensions and in the non-asymptotic setting. We first derive a lower and a matching upper bound for the minimax risk of clustering in this framework. We also show that in the high-dimensional regime, the linear discriminant analysis (LDA) classifier turns out to be sub-optimal in the minimax sense. Next, we characterize precisely the risk of $\ell_2$-regularized supervised least squares classifiers. We deduce the fact that the interpolating solution may outperform the regularized classifier, under mild assumptions on the covariance structure of the noise. Our analysis also shows that interpolation can be robust to corruption in the covariance of the noise when the signal is aligned with the "clean" part of the covariance, for the properly defined notion of alignment. To the best of our knowledge, this peculiar phenomenon has not yet been investigated in the rapidly growing literature related to interpolation. We conclude that interpolation is not only benign but can also be optimal, and in some cases robust.
Adaptive Early-Learning Correction for Segmentation from Noisy Annotations
Oct 07, 2021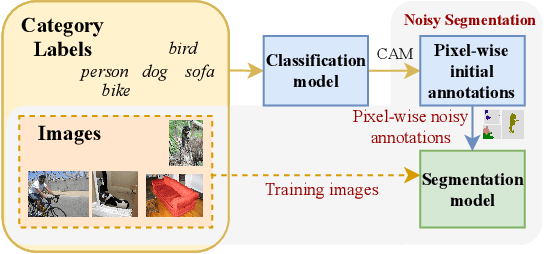

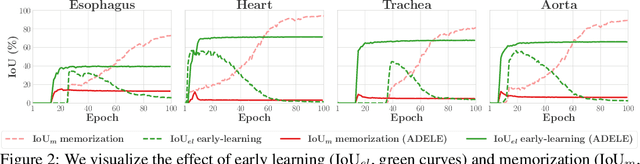

Abstract:Deep learning in the presence of noisy annotations has been studied extensively in classification, but much less in segmentation tasks. In this work, we study the learning dynamics of deep segmentation networks trained on inaccurately-annotated data. We discover a phenomenon that has been previously reported in the context of classification: the networks tend to first fit the clean pixel-level labels during an "early-learning" phase, before eventually memorizing the false annotations. However, in contrast to classification, memorization in segmentation does not arise simultaneously for all semantic categories. Inspired by these findings, we propose a new method for segmentation from noisy annotations with two key elements. First, we detect the beginning of the memorization phase separately for each category during training. This allows us to adaptively correct the noisy annotations in order to exploit early learning. Second, we incorporate a regularization term that enforces consistency across scales to boost robustness against annotation noise. Our method outperforms standard approaches on a medical-imaging segmentation task where noises are synthesized to mimic human annotation errors. It also provides robustness to realistic noisy annotations present in weakly-supervised semantic segmentation, achieving state-of-the-art results on PASCAL VOC 2012.
Weakly-supervised High-resolution Segmentation of Mammography Images for Breast Cancer Diagnosis
Jun 15, 2021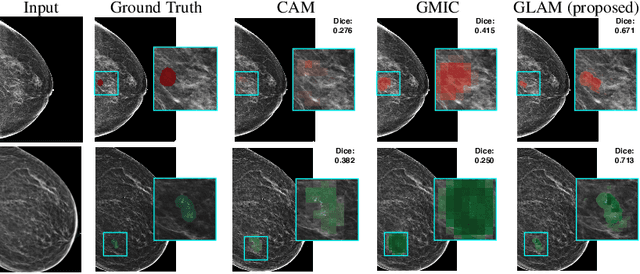
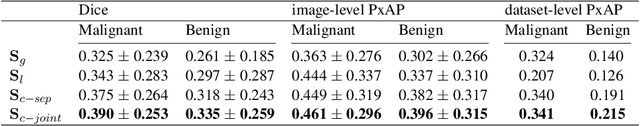
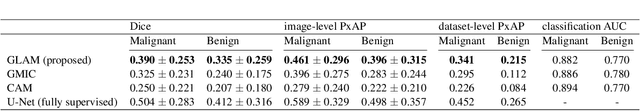
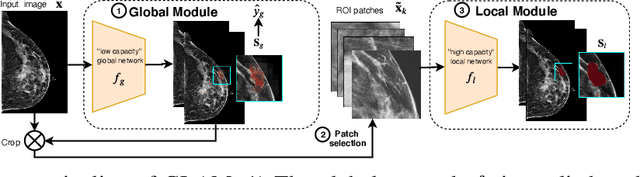

Abstract:In the last few years, deep learning classifiers have shown promising results in image-based medical diagnosis. However, interpreting the outputs of these models remains a challenge. In cancer diagnosis, interpretability can be achieved by localizing the region of the input image responsible for the output, i.e. the location of a lesion. Alternatively, segmentation or detection models can be trained with pixel-wise annotations indicating the locations of malignant lesions. Unfortunately, acquiring such labels is labor-intensive and requires medical expertise. To overcome this difficulty, weakly-supervised localization can be utilized. These methods allow neural network classifiers to output saliency maps highlighting the regions of the input most relevant to the classification task (e.g. malignant lesions in mammograms) using only image-level labels (e.g. whether the patient has cancer or not) during training. When applied to high-resolution images, existing methods produce low-resolution saliency maps. This is problematic in applications in which suspicious lesions are small in relation to the image size. In this work, we introduce a novel neural network architecture to perform weakly-supervised segmentation of high-resolution images. The proposed model selects regions of interest via coarse-level localization, and then performs fine-grained segmentation of those regions. We apply this model to breast cancer diagnosis with screening mammography, and validate it on a large clinically-realistic dataset. Measured by Dice similarity score, our approach outperforms existing methods by a large margin in terms of localization performance of benign and malignant lesions, relatively improving the performance by 39.6% and 20.0%, respectively. Code and the weights of some of the models are available at https://github.com/nyukat/GLAM
Reducing false-positive biopsies with deep neural networks that utilize local and global information in screening mammograms
Sep 19, 2020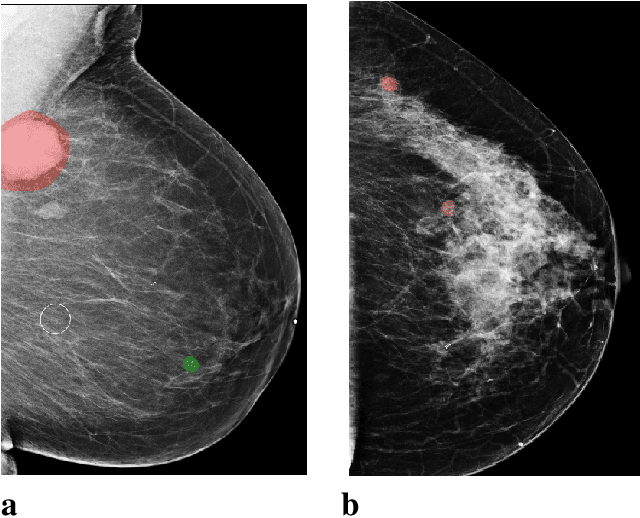
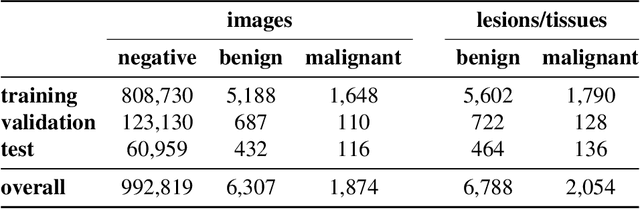
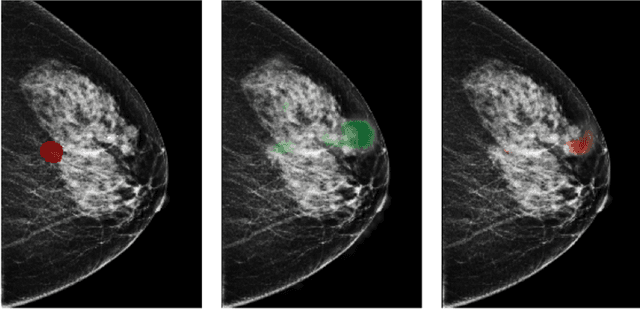
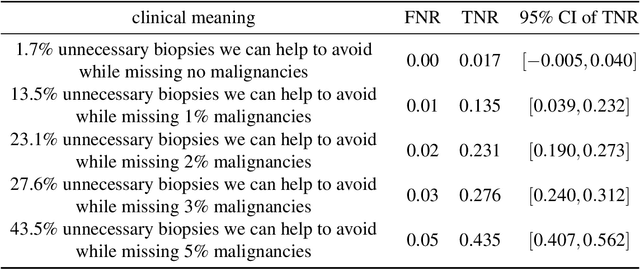
Abstract:Breast cancer is the most common cancer in women, and hundreds of thousands of unnecessary biopsies are done around the world at a tremendous cost. It is crucial to reduce the rate of biopsies that turn out to be benign tissue. In this study, we build deep neural networks (DNNs) to classify biopsied lesions as being either malignant or benign, with the goal of using these networks as second readers serving radiologists to further reduce the number of false positive findings. We enhance the performance of DNNs that are trained to learn from small image patches by integrating global context provided in the form of saliency maps learned from the entire image into their reasoning, similar to how radiologists consider global context when evaluating areas of interest. Our experiments are conducted on a dataset of 229,426 screening mammography exams from 141,473 patients. We achieve an AUC of 0.8 on a test set consisting of 464 benign and 136 malignant lesions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge