Yash Sharma
Provably Learning Object-Centric Representations
May 23, 2023



Abstract:Learning structured representations of the visual world in terms of objects promises to significantly improve the generalization abilities of current machine learning models. While recent efforts to this end have shown promising empirical progress, a theoretical account of when unsupervised object-centric representation learning is possible is still lacking. Consequently, understanding the reasons for the success of existing object-centric methods as well as designing new theoretically grounded methods remains challenging. In the present work, we analyze when object-centric representations can provably be learned without supervision. To this end, we first introduce two assumptions on the generative process for scenes comprised of several objects, which we call compositionality and irreducibility. Under this generative process, we prove that the ground-truth object representations can be identified by an invertible and compositional inference model, even in the presence of dependencies between objects. We empirically validate our results through experiments on synthetic data. Finally, we provide evidence that our theory holds predictive power for existing object-centric models by showing a close correspondence between models' compositionality and invertibility and their empirical identifiability.
How Adversarial Robustness Transfers from Pre-training to Downstream Tasks
Aug 07, 2022



Abstract:Given the rise of large-scale training regimes, adapting pre-trained models to a wide range of downstream tasks has become a standard approach in machine learning. While large benefits in empirical performance have been observed, it is not yet well understood how robustness properties transfer from a pre-trained model to a downstream task. We prove that the robustness of a predictor on downstream tasks can be bound by the robustness of its underlying representation, irrespective of the pre-training protocol. Taken together, our results precisely characterize what is required of the representation function for reliable performance upon deployment.
Weakly Supervised Deep Instance Nuclei Detection using Points Annotation in 3D Cardiovascular Immunofluorescent Images
Jul 29, 2022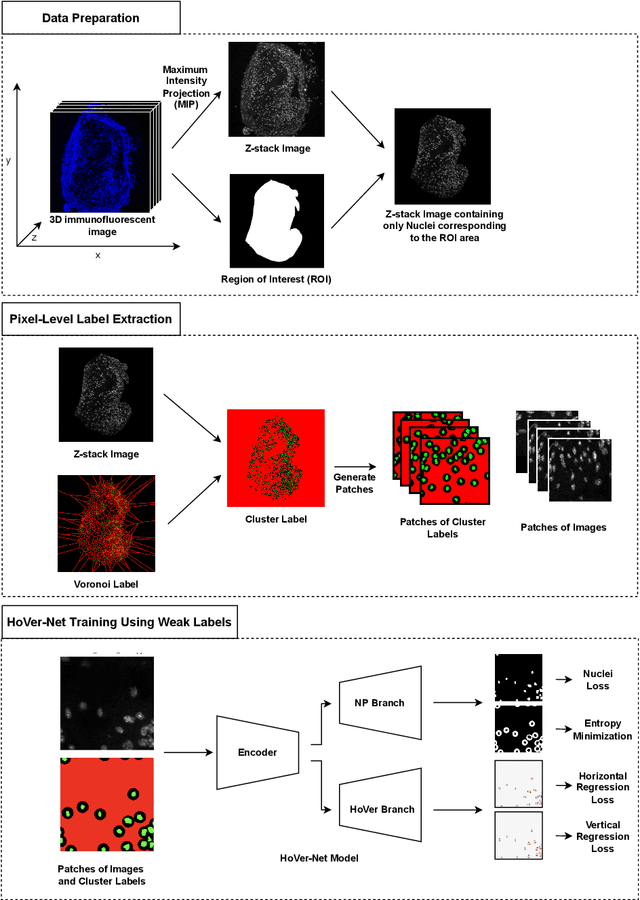
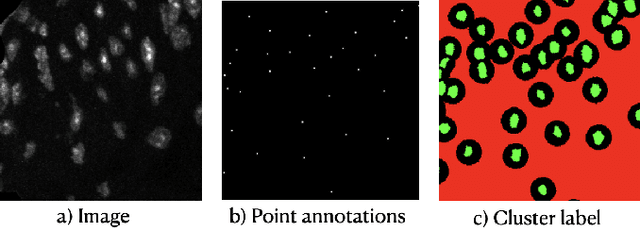


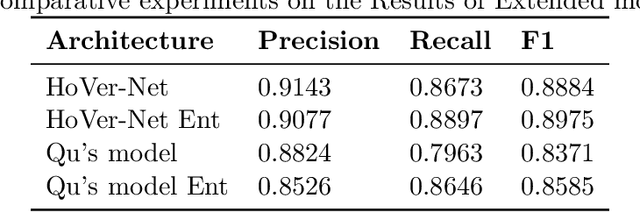
Abstract:Two major causes of death in the United States and worldwide are stroke and myocardial infarction. The underlying cause of both is thrombi released from ruptured or eroded unstable atherosclerotic plaques that occlude vessels in the heart (myocardial infarction) or the brain (stroke). Clinical studies show that plaque composition plays a more important role than lesion size in plaque rupture or erosion events. To determine the plaque composition, various cell types in 3D cardiovascular immunofluorescent images of plaque lesions are counted. However, counting these cells manually is expensive, time-consuming, and prone to human error. These challenges of manual counting motivate the need for an automated approach to localize and count the cells in images. The purpose of this study is to develop an automatic approach to accurately detect and count cells in 3D immunofluorescent images with minimal annotation effort. In this study, we used a weakly supervised learning approach to train the HoVer-Net segmentation model using point annotations to detect nuclei in fluorescent images. The advantage of using point annotations is that they require less effort as opposed to pixel-wise annotation. To train the HoVer-Net model using point annotations, we adopted a popularly used cluster labeling approach to transform point annotations into accurate binary masks of cell nuclei. Traditionally, these approaches have generated binary masks from point annotations, leaving a region around the object unlabeled (which is typically ignored during model training). However, these areas may contain important information that helps determine the boundary between cells. Therefore, we used the entropy minimization loss function in these areas to encourage the model to output more confident predictions on the unlabeled areas. Our comparison studies indicate that the HoVer-Net model trained using our weakly ...
Pixel-level Correspondence for Self-Supervised Learning from Video
Jul 08, 2022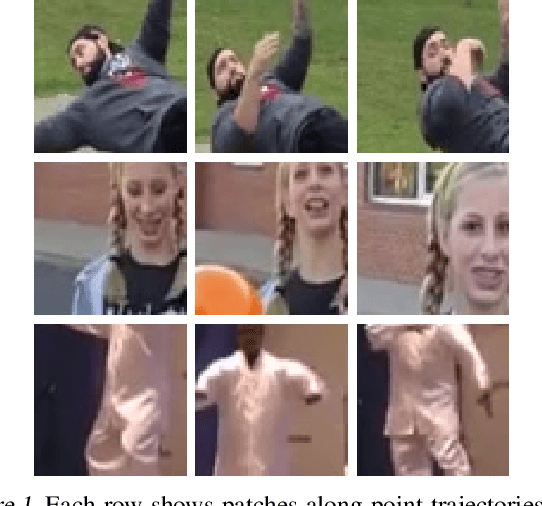
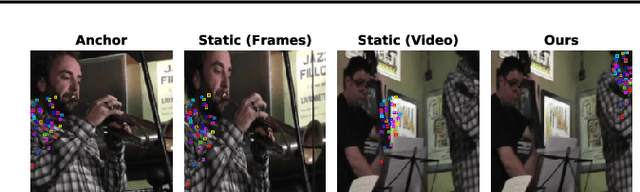
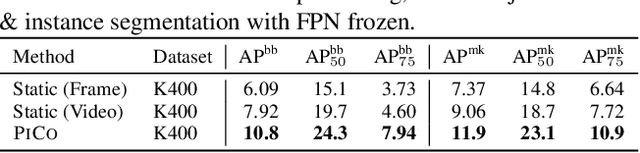
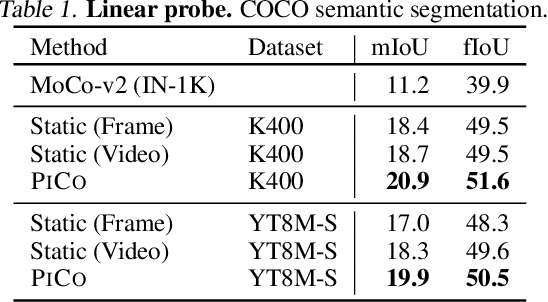


Abstract:While self-supervised learning has enabled effective representation learning in the absence of labels, for vision, video remains a relatively untapped source of supervision. To address this, we propose Pixel-level Correspondence (PiCo), a method for dense contrastive learning from video. By tracking points with optical flow, we obtain a correspondence map which can be used to match local features at different points in time. We validate PiCo on standard benchmarks, outperforming self-supervised baselines on multiple dense prediction tasks, without compromising performance on image classification.
MaNi: Maximizing Mutual Information for Nuclei Cross-Domain Unsupervised Segmentation
Jun 29, 2022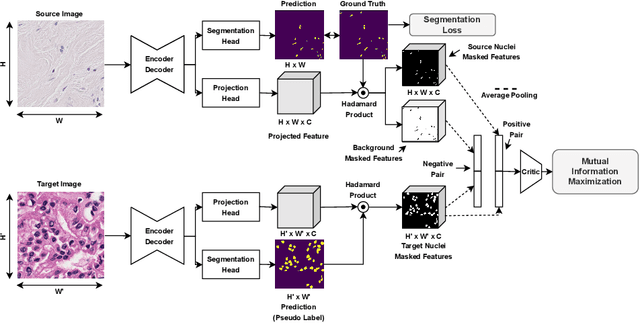
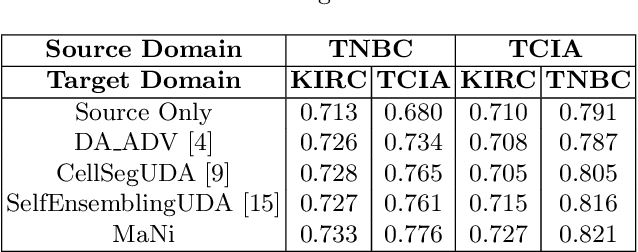
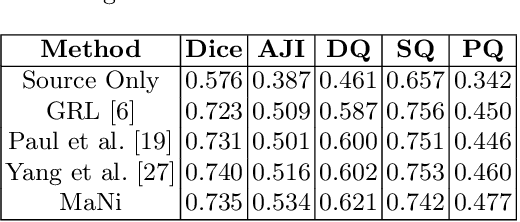
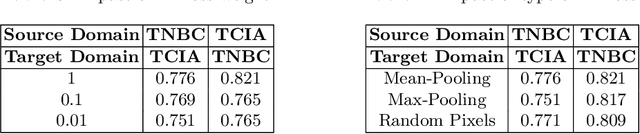
Abstract:In this work, we propose a mutual information (MI) based unsupervised domain adaptation (UDA) method for the cross-domain nuclei segmentation. Nuclei vary substantially in structure and appearances across different cancer types, leading to a drop in performance of deep learning models when trained on one cancer type and tested on another. This domain shift becomes even more critical as accurate segmentation and quantification of nuclei is an essential histopathology task for the diagnosis/ prognosis of patients and annotating nuclei at the pixel level for new cancer types demands extensive effort by medical experts. To address this problem, we maximize the MI between labeled source cancer type data and unlabeled target cancer type data for transferring nuclei segmentation knowledge across domains. We use the Jensen-Shanon divergence bound, requiring only one negative pair per positive pair for MI maximization. We evaluate our set-up for multiple modeling frameworks and on different datasets comprising of over 20 cancer-type domain shifts and demonstrate competitive performance. All the recently proposed approaches consist of multiple components for improving the domain adaptation, whereas our proposed module is light and can be easily incorporated into other methods (Implementation: https://github.com/YashSharma/MaNi ).
Encoding Cardiopulmonary Exercise Testing Time Series as Images for Classification using Convolutional Neural Network
Apr 26, 2022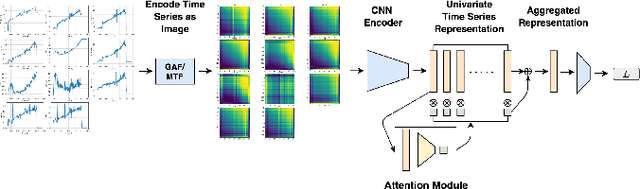
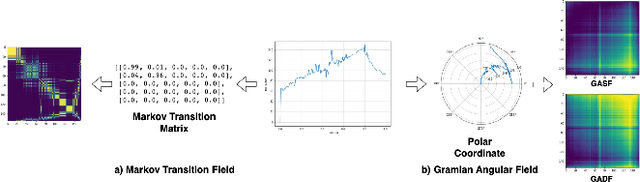
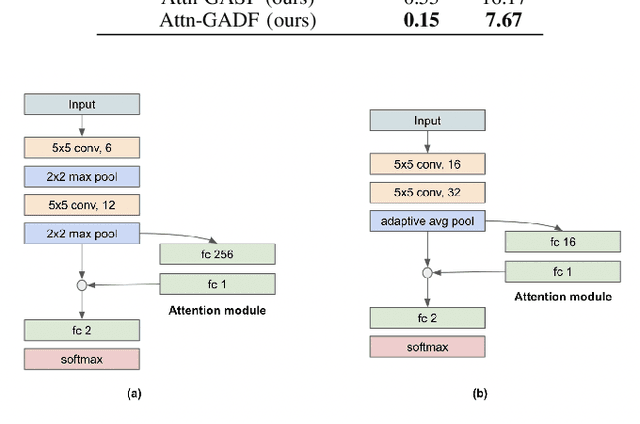
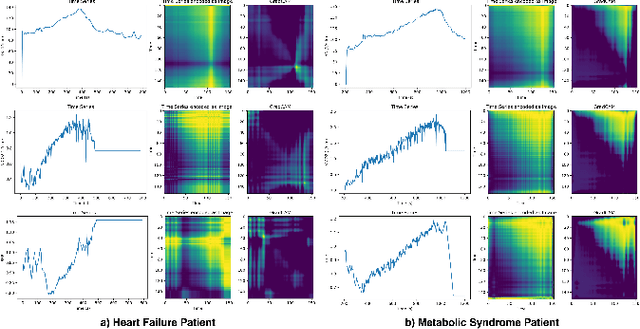
Abstract:Exercise testing has been available for more than a half-century and is a remarkably versatile tool for diagnostic and prognostic information of patients for a range of diseases, especially cardiovascular and pulmonary. With rapid advancements in technology, wearables, and learning algorithm in the last decade, its scope has evolved. Specifically, Cardiopulmonary exercise testing (CPX) is one of the most commonly used laboratory tests for objective evaluation of exercise capacity and performance levels in patients. CPX provides a non-invasive, integrative assessment of the pulmonary, cardiovascular, and skeletal muscle systems involving the measurement of gas exchanges. However, its assessment is challenging, requiring the individual to process multiple time series data points, leading to simplification to peak values and slopes. But this simplification can discard the valuable trend information present in these time series. In this work, we encode the time series as images using the Gramian Angular Field and Markov Transition Field and use it with a convolutional neural network and attention pooling approach for the classification of heart failure and metabolic syndrome patients. Using GradCAMs, we highlight the discriminative features identified by the model.
Unsupervised Learning of Compositional Energy Concepts
Nov 04, 2021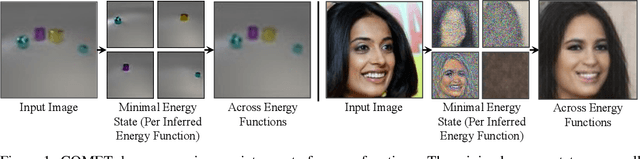
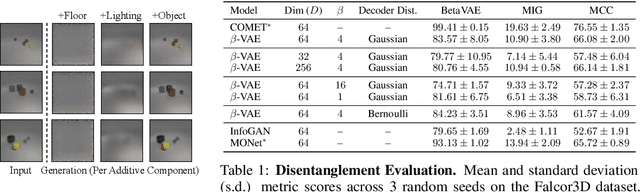
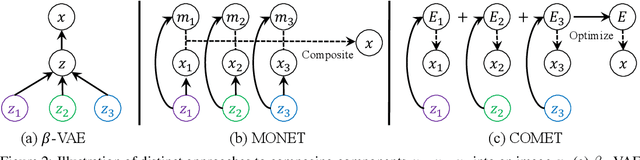
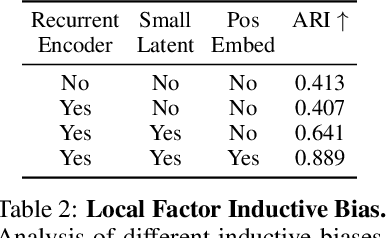
Abstract:Humans are able to rapidly understand scenes by utilizing concepts extracted from prior experience. Such concepts are diverse, and include global scene descriptors, such as the weather or lighting, as well as local scene descriptors, such as the color or size of a particular object. So far, unsupervised discovery of concepts has focused on either modeling the global scene-level or the local object-level factors of variation, but not both. In this work, we propose COMET, which discovers and represents concepts as separate energy functions, enabling us to represent both global concepts as well as objects under a unified framework. COMET discovers energy functions through recomposing the input image, which we find captures independent factors without additional supervision. Sample generation in COMET is formulated as an optimization process on underlying energy functions, enabling us to generate images with permuted and composed concepts. Finally, discovered visual concepts in COMET generalize well, enabling us to compose concepts between separate modalities of images as well as with other concepts discovered by a separate instance of COMET trained on a different dataset. Code and data available at https://energy-based-model.github.io/comet/.
HistoTransfer: Understanding Transfer Learning for Histopathology
Jun 13, 2021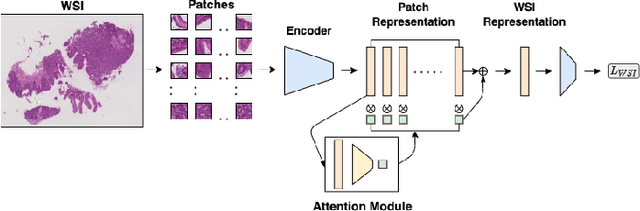
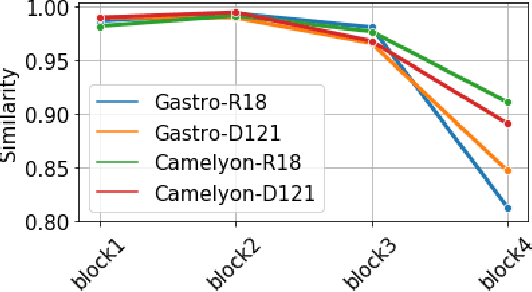
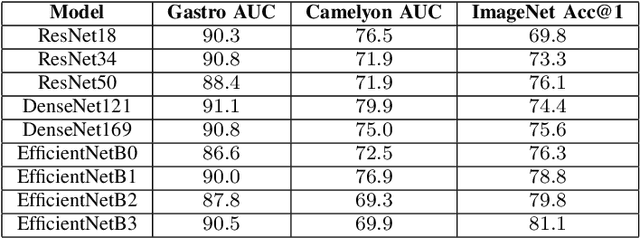

Abstract:Advancement in digital pathology and artificial intelligence has enabled deep learning-based computer vision techniques for automated disease diagnosis and prognosis. However, WSIs present unique computational and algorithmic challenges. WSIs are gigapixel-sized, making them infeasible to be used directly for training deep neural networks. Hence, for modeling, a two-stage approach is adopted: Patch representations are extracted first, followed by the aggregation for WSI prediction. These approaches require detailed pixel-level annotations for training the patch encoder. However, obtaining these annotations is time-consuming and tedious for medical experts. Transfer learning is used to address this gap and deep learning architectures pre-trained on ImageNet are used for generating patch-level representation. Even though ImageNet differs significantly from histopathology data, pre-trained networks have been shown to perform impressively on histopathology data. Also, progress in self-supervised and multi-task learning coupled with the release of multiple histopathology data has led to the release of histopathology-specific networks. In this work, we compare the performance of features extracted from networks trained on ImageNet and histopathology data. We use an attention pooling network over these extracted features for slide-level aggregation. We investigate if features learned using more complex networks lead to gain in performance. We use a simple top-k sampling approach for fine-tuning framework and study the representation similarity between frozen and fine-tuned networks using Centered Kernel Alignment. Further, to examine if intermediate block representation is better suited for feature extraction and ImageNet architectures are unnecessarily large for histopathology, we truncate the blocks of ResNet18 and DenseNet121 and examine the performance.
Self-Supervised Learning with Data Augmentations Provably Isolates Content from Style
Jun 08, 2021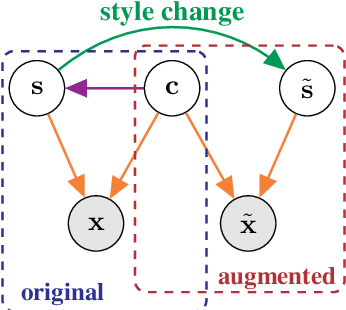
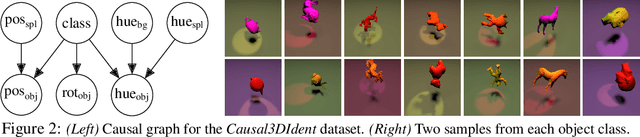
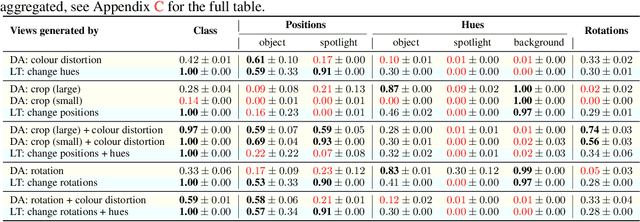

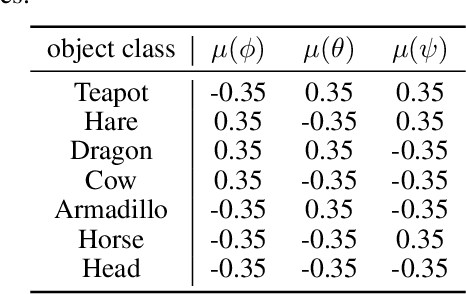
Abstract:Self-supervised representation learning has shown remarkable success in a number of domains. A common practice is to perform data augmentation via hand-crafted transformations intended to leave the semantics of the data invariant. We seek to understand the empirical success of this approach from a theoretical perspective. We formulate the augmentation process as a latent variable model by postulating a partition of the latent representation into a content component, which is assumed invariant to augmentation, and a style component, which is allowed to change. Unlike prior work on disentanglement and independent component analysis, we allow for both nontrivial statistical and causal dependencies in the latent space. We study the identifiability of the latent representation based on pairs of views of the observations and prove sufficient conditions that allow us to identify the invariant content partition up to an invertible mapping in both generative and discriminative settings. We find numerical simulations with dependent latent variables are consistent with our theory. Lastly, we introduce Causal3DIdent, a dataset of high-dimensional, visually complex images with rich causal dependencies, which we use to study the effect of data augmentations performed in practice.
Cluster-to-Conquer: A Framework for End-to-End Multi-Instance Learning for Whole Slide Image Classification
Mar 19, 2021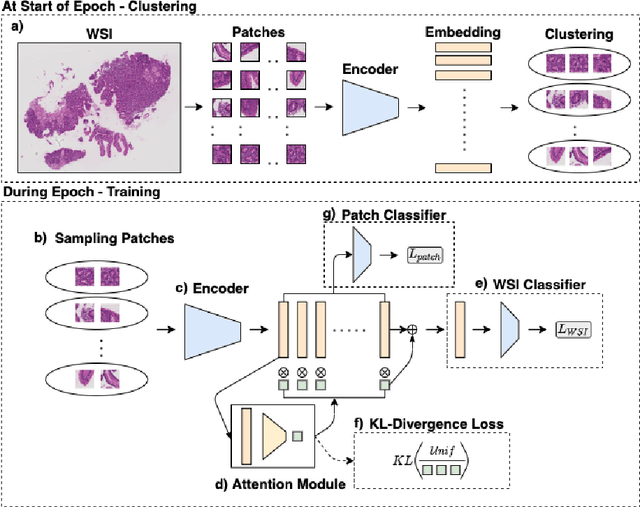
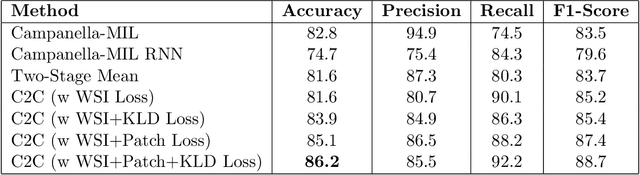
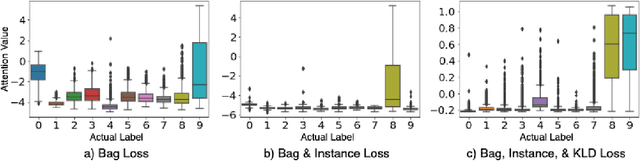
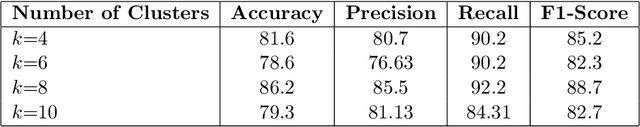
Abstract:In recent years, the availability of digitized Whole Slide Images (WSIs) has enabled the use of deep learning-based computer vision techniques for automated disease diagnosis. However, WSIs present unique computational and algorithmic challenges. WSIs are gigapixel-sized ($\sim$100K pixels), making them infeasible to be used directly for training deep neural networks. Also, often only slide-level labels are available for training as detailed annotations are tedious and can be time-consuming for experts. Approaches using multiple-instance learning (MIL) frameworks have been shown to overcome these challenges. Current state-of-the-art approaches divide the learning framework into two decoupled parts: a convolutional neural network (CNN) for encoding the patches followed by an independent aggregation approach for slide-level prediction. In this approach, the aggregation step has no bearing on the representations learned by the CNN encoder. We have proposed an end-to-end framework that clusters the patches from a WSI into ${k}$-groups, samples ${k}'$ patches from each group for training, and uses an adaptive attention mechanism for slide level prediction; Cluster-to-Conquer (C2C). We have demonstrated that dividing a WSI into clusters can improve the model training by exposing it to diverse discriminative features extracted from the patches. We regularized the clustering mechanism by introducing a KL-divergence loss between the attention weights of patches in a cluster and the uniform distribution. The framework is optimized end-to-end on slide-level cross-entropy, patch-level cross-entropy, and KL-divergence loss (Implementation: https://github.com/YashSharma/C2C).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge