Xiaowei Jia
AgroFlux: A Spatial-Temporal Benchmark for Carbon and Nitrogen Flux Prediction in Agricultural Ecosystems
Feb 02, 2026Abstract:Agroecosystem, which heavily influenced by human actions and accounts for a quarter of global greenhouse gas emissions (GHGs), plays a crucial role in mitigating global climate change and securing environmental sustainability. However, we can't manage what we can't measure. Accurately quantifying the pools and fluxes in the carbon, nutrient, and water nexus of the agroecosystem is therefore essential for understanding the underlying drivers of GHG and developing effective mitigation strategies. Conventional approaches like soil sampling, process-based models, and black-box machine learning models are facing challenges such as data sparsity, high spatiotemporal heterogeneity, and complex subsurface biogeochemical and physical processes. Developing new trustworthy approaches such as AI-empowered models, will require the AI-ready benchmark dataset and outlined protocols, which unfortunately do not exist. In this work, we introduce a first-of-its-kind spatial-temporal agroecosystem GHG benchmark dataset that integrates physics-based model simulations from Ecosys and DayCent with real-world observations from eddy covariance flux towers and controlled-environment facilities. We evaluate the performance of various sequential deep learning models on carbon and nitrogen flux prediction, including LSTM-based models, temporal CNN-based model, and Transformer-based models. Furthermore, we explored transfer learning to leverage simulated data to improve the generalization of deep learning models on real-world observations. Our benchmark dataset and evaluation framework contribute to the development of more accurate and scalable AI-driven agroecosystem models, advancing our understanding of ecosystem-climate interactions.
Learning PDE Solvers with Physics and Data: A Unifying View of Physics-Informed Neural Networks and Neural Operators
Jan 20, 2026Abstract:Partial differential equations (PDEs) are central to scientific modeling. Modern workflows increasingly rely on learning-based components to support model reuse, inference, and integration across large computational processes. Despite the emergence of various physics-aware data-driven approaches, the field still lacks a unified perspective to uncover their relationships, limitations, and appropriate roles in scientific workflows. To this end, we propose a unifying perspective to place two dominant paradigms: Physics-Informed Neural Networks (PINNs) and Neural Operators (NOs), within a shared design space. We organize existing methods from three fundamental dimensions: what is learned, how physical structures are integrated into the learning process, and how the computational load is amortized across problem instances. In this way, many challenges can be best understood as consequences of these structural properties of learning PDEs. By analyzing advances through this unifying view, our survey aims to facilitate the development of reliable learning-based PDE solvers and catalyze a synthesis of physics and data.
Accelerating MHC-II Epitope Discovery via Multi-Scale Prediction in Antigen Presentation
Dec 16, 2025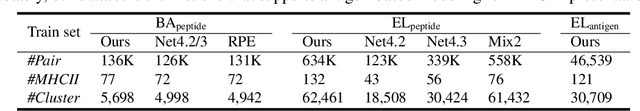
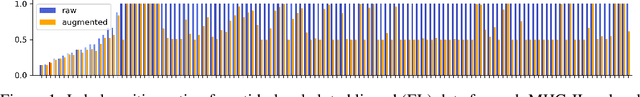
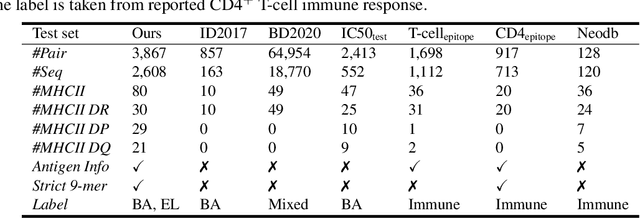
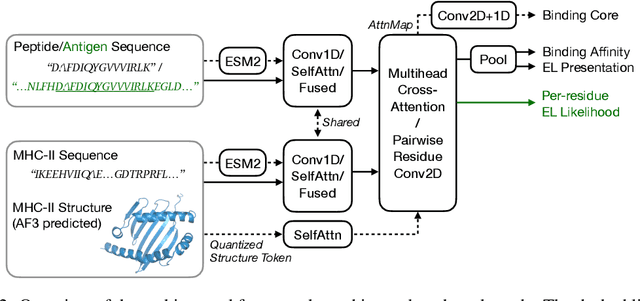
Abstract:Antigenic epitope presented by major histocompatibility complex II (MHC-II) proteins plays an essential role in immunotherapy. However, compared to the more widely studied MHC-I in computational immunotherapy, the study of MHC-II antigenic epitope poses significantly more challenges due to its complex binding specificity and ambiguous motif patterns. Consequently, existing datasets for MHC-II interactions are smaller and less standardized than those available for MHC-I. To address these challenges, we present a well-curated dataset derived from the Immune Epitope Database (IEDB) and other public sources. It not only extends and standardizes existing peptide-MHC-II datasets, but also introduces a novel antigen-MHC-II dataset with richer biological context. Leveraging this dataset, we formulate three major machine learning (ML) tasks of peptide binding, peptide presentation, and antigen presentation, which progressively capture the broader biological processes within the MHC-II antigen presentation pathway. We further employ a multi-scale evaluation framework to benchmark existing models, along with a comprehensive analysis over various modeling designs to this problem with a modular framework. Overall, this work serves as a valuable resource for advancing computational immunotherapy, providing a foundation for future research in ML guided epitope discovery and predictive modeling of immune responses.
GREAT: Generalizable Representation Enhancement via Auxiliary Transformations for Zero-Shot Environmental Prediction
Nov 17, 2025Abstract:Environmental modeling faces critical challenges in predicting ecosystem dynamics across unmonitored regions due to limited and geographically imbalanced observation data. This challenge is compounded by spatial heterogeneity, causing models to learn spurious patterns that fit only local data. Unlike conventional domain generalization, environmental modeling must preserve invariant physical relationships and temporal coherence during augmentation. In this paper, we introduce Generalizable Representation Enhancement via Auxiliary Transformations (GREAT), a framework that effectively augments available datasets to improve predictions in completely unseen regions. GREAT guides the augmentation process to ensure that the original governing processes can be recovered from the augmented data, and the inclusion of the augmented data leads to improved model generalization. Specifically, GREAT learns transformation functions at multiple layers of neural networks to augment both raw environmental features and temporal influence. They are refined through a novel bi-level training process that constrains augmented data to preserve key patterns of the original source data. We demonstrate GREAT's effectiveness on stream temperature prediction across six ecologically diverse watersheds in the eastern U.S., each containing multiple stream segments. Experimental results show that GREAT significantly outperforms existing methods in zero-shot scenarios. This work provides a practical solution for environmental applications where comprehensive monitoring is infeasible.
Geo-Aware Models for Stream Temperature Prediction across Different Spatial Regions and Scales
Oct 10, 2025Abstract:Understanding environmental ecosystems is vital for the sustainable management of our planet. However,existing physics-based and data-driven models often fail to generalize to varying spatial regions and scales due to the inherent data heterogeneity presented in real environmental ecosystems. This generalization issue is further exacerbated by the limited observation samples available for model training. To address these issues, we propose Geo-STARS, a geo-aware spatio-temporal modeling framework for predicting stream water temperature across different watersheds and spatial scales. The major innovation of Geo-STARS is the introduction of geo-aware embedding, which leverages geographic information to explicitly capture shared principles and patterns across spatial regions and scales. We further integrate the geo-aware embedding into a gated spatio-temporal graph neural network. This design enables the model to learn complex spatial and temporal patterns guided by geographic and hydrological context, even with sparse or no observational data. We evaluate Geo-STARS's efficacy in predicting stream water temperature, which is a master factor for water quality. Using real-world datasets spanning 37 years across multiple watersheds along the eastern coast of the United States, Geo-STARS demonstrates its superior generalization performance across both regions and scales, outperforming state-of-the-art baselines. These results highlight the promise of Geo-STARS for scalable, data-efficient environmental monitoring and decision-making.
Learning to Retrieve for Environmental Knowledge Discovery: An Augmentation-Adaptive Self-Supervised Learning Framework
Sep 18, 2025Abstract:The discovery of environmental knowledge depends on labeled task-specific data, but is often constrained by the high cost of data collection. Existing machine learning approaches usually struggle to generalize in data-sparse or atypical conditions. To this end, we propose an Augmentation-Adaptive Self-Supervised Learning (A$^2$SL) framework, which retrieves relevant observational samples to enhance modeling of the target ecosystem. Specifically, we introduce a multi-level pairwise learning loss to train a scenario encoder that captures varying degrees of similarity among scenarios. These learned similarities drive a retrieval mechanism that supplements a target scenario with relevant data from different locations or time periods. Furthermore, to better handle variable scenarios, particularly under atypical or extreme conditions where traditional models struggle, we design an augmentation-adaptive mechanism that selectively enhances these scenarios through targeted data augmentation. Using freshwater ecosystems as a case study, we evaluate A$^2$SL in modeling water temperature and dissolved oxygen dynamics in real-world lakes. Experimental results show that A$^2$SL significantly improves predictive accuracy and enhances robustness in data-scarce and atypical scenarios. Although this study focuses on freshwater ecosystems, the A$^2$SL framework offers a broadly applicable solution in various scientific domains.
Unveiling Confirmation Bias in Chain-of-Thought Reasoning
Jun 14, 2025



Abstract:Chain-of-thought (CoT) prompting has been widely adopted to enhance the reasoning capabilities of large language models (LLMs). However, the effectiveness of CoT reasoning is inconsistent across tasks with different reasoning types. This work presents a novel perspective to understand CoT behavior through the lens of \textit{confirmation bias} in cognitive psychology. Specifically, we examine how model internal beliefs, approximated by direct question-answering probabilities, affect both reasoning generation ($Q \to R$) and reasoning-guided answer prediction ($QR \to A$) in CoT. By decomposing CoT into a two-stage process, we conduct a thorough correlation analysis in model beliefs, rationale attributes, and stage-wise performance. Our results provide strong evidence of confirmation bias in LLMs, such that model beliefs not only skew the reasoning process but also influence how rationales are utilized for answer prediction. Furthermore, the interplay between task vulnerability to confirmation bias and the strength of beliefs also provides explanations for CoT effectiveness across reasoning tasks and models. Overall, this study provides a valuable insight for the needs of better prompting strategies that mitigate confirmation bias to enhance reasoning performance. Code is available at \textit{https://github.com/yuewan2/biasedcot}.
X-MethaneWet: A Cross-scale Global Wetland Methane Emission Benchmark Dataset for Advancing Science Discovery with AI
May 23, 2025Abstract:Methane (CH$_4$) is the second most powerful greenhouse gas after carbon dioxide and plays a crucial role in climate change due to its high global warming potential. Accurately modeling CH$_4$ fluxes across the globe and at fine temporal scales is essential for understanding its spatial and temporal variability and developing effective mitigation strategies. In this work, we introduce the first-of-its-kind cross-scale global wetland methane benchmark dataset (X-MethaneWet), which synthesizes physics-based model simulation data from TEM-MDM and the real-world observation data from FLUXNET-CH$_4$. This dataset can offer opportunities for improving global wetland CH$_4$ modeling and science discovery with new AI algorithms. To set up AI model baselines for methane flux prediction, we evaluate the performance of various sequential deep learning models on X-MethaneWet. Furthermore, we explore four different transfer learning techniques to leverage simulated data from TEM-MDM to improve the generalization of deep learning models on real-world FLUXNET-CH$_4$ observations. Our extensive experiments demonstrate the effectiveness of these approaches, highlighting their potential for advancing methane emission modeling and contributing to the development of more accurate and scalable AI-driven climate models.
LLM-based Evaluation Policy Extraction for Ecological Modeling
May 20, 2025Abstract:Evaluating ecological time series is critical for benchmarking model performance in many important applications, including predicting greenhouse gas fluxes, capturing carbon-nitrogen dynamics, and monitoring hydrological cycles. Traditional numerical metrics (e.g., R-squared, root mean square error) have been widely used to quantify the similarity between modeled and observed ecosystem variables, but they often fail to capture domain-specific temporal patterns critical to ecological processes. As a result, these methods are often accompanied by expert visual inspection, which requires substantial human labor and limits the applicability to large-scale evaluation. To address these challenges, we propose a novel framework that integrates metric learning with large language model (LLM)-based natural language policy extraction to develop interpretable evaluation criteria. The proposed method processes pairwise annotations and implements a policy optimization mechanism to generate and combine different assessment metrics. The results obtained on multiple datasets for evaluating the predictions of crop gross primary production and carbon dioxide flux have confirmed the effectiveness of the proposed method in capturing target assessment preferences, including both synthetically generated and expert-annotated model comparisons. The proposed framework bridges the gap between numerical metrics and expert knowledge while providing interpretable evaluation policies that accommodate the diverse needs of different ecosystem modeling studies.
Knowledge Guided Encoder-Decoder Framework: Integrating Multiple Physical Models for Agricultural Ecosystem Modeling
May 13, 2025



Abstract:Agricultural monitoring is critical for ensuring food security, maintaining sustainable farming practices, informing policies on mitigating food shortage, and managing greenhouse gas emissions. Traditional process-based physical models are often designed and implemented for specific situations, and their parameters could also be highly uncertain. In contrast, data-driven models often use black-box structures and does not explicitly model the inter-dependence between different ecological variables. As a result, they require extensive training data and lack generalizability to different tasks with data distribution shifts and inconsistent observed variables. To address the need for more universal models, we propose a knowledge-guided encoder-decoder model, which can predict key crop variables by leveraging knowledge of underlying processes from multiple physical models. The proposed method also integrates a language model to process complex and inconsistent inputs and also utilizes it to implement a model selection mechanism for selectively combining the knowledge from different physical models. Our evaluations on predicting carbon and nitrogen fluxes for multiple sites demonstrate the effectiveness and robustness of the proposed model under various scenarios.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge