Vince Lyzinski
Information Recovery in Shuffled Graphs via Graph Matching
Sep 27, 2017
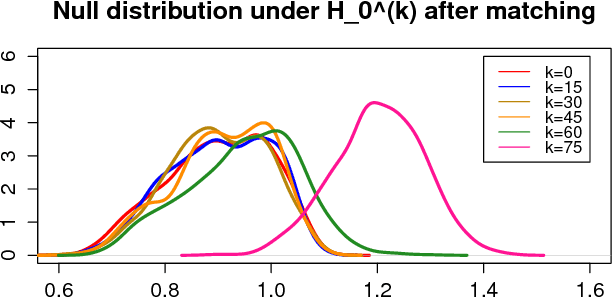
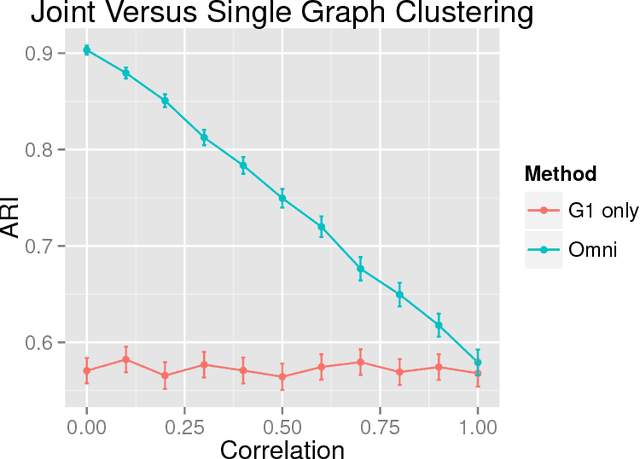
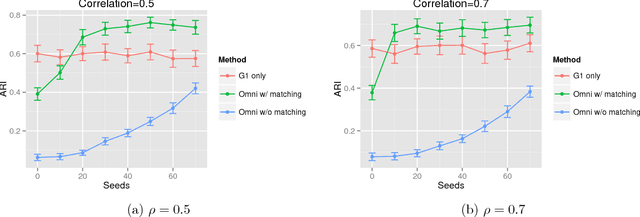
Abstract:While many multiple graph inference methodologies operate under the implicit assumption that an explicit vertex correspondence is known across the vertex sets of the graphs, in practice these correspondences may only be partially or errorfully known. Herein, we provide an information theoretic foundation for understanding the practical impact that errorfully observed vertex correspondences can have on subsequent inference, and the capacity of graph matching methods to recover the lost vertex alignment and inferential performance. Working in the correlated stochastic blockmodel setting, we establish a duality between the loss of mutual information due to an errorfully observed vertex correspondence and the ability of graph matching algorithms to recover the true correspondence across graphs. In the process, we establish a phase transition for graph matchability in terms of the correlation across graphs, and we conjecture the analogous phase transition for the relative information loss due to shuffling vertex labels. We demonstrate the practical effect that graph shuffling---and matching---can have on subsequent inference, with examples from two sample graph hypothesis testing and joint spectral graph clustering.
Statistical inference on random dot product graphs: a survey
Sep 16, 2017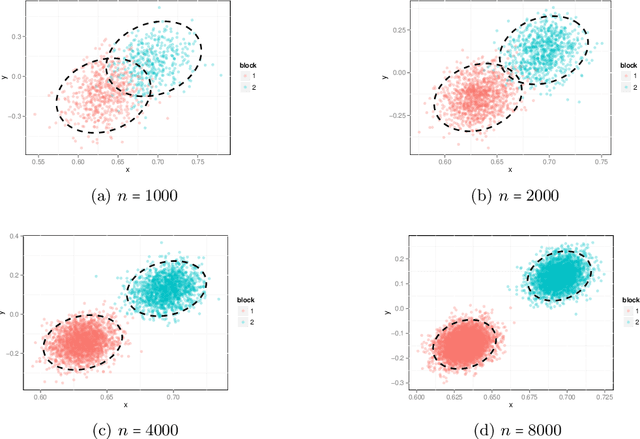
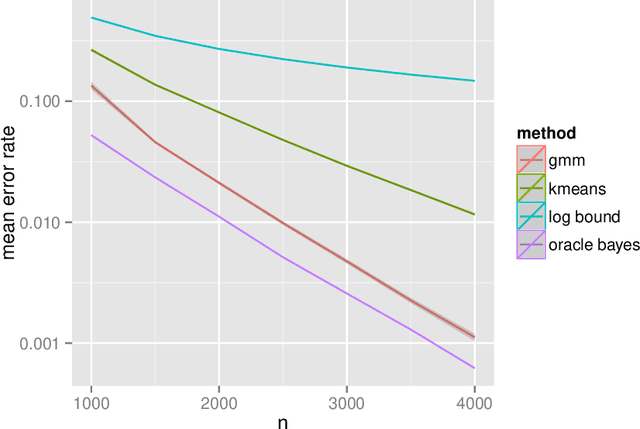

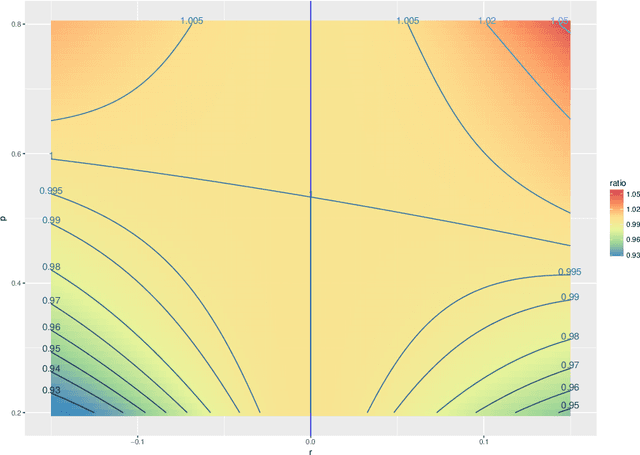
Abstract:The random dot product graph (RDPG) is an independent-edge random graph that is analytically tractable and, simultaneously, either encompasses or can successfully approximate a wide range of random graphs, from relatively simple stochastic block models to complex latent position graphs. In this survey paper, we describe a comprehensive paradigm for statistical inference on random dot product graphs, a paradigm centered on spectral embeddings of adjacency and Laplacian matrices. We examine the analogues, in graph inference, of several canonical tenets of classical Euclidean inference: in particular, we summarize a body of existing results on the consistency and asymptotic normality of the adjacency and Laplacian spectral embeddings, and the role these spectral embeddings can play in the construction of single- and multi-sample hypothesis tests for graph data. We investigate several real-world applications, including community detection and classification in large social networks and the determination of functional and biologically relevant network properties from an exploratory data analysis of the Drosophila connectome. We outline requisite background and current open problems in spectral graph inference.
* An expository survey paper on a comprehensive paradigm for inference for random dot product graphs, centered on graph adjacency and Laplacian spectral embeddings. Paper outlines requisite background; summarizes theory, methodology, and applications from previous and ongoing work; and closes with a discussion of several open problems
Vertex Nomination Via Local Neighborhood Matching
Jul 22, 2017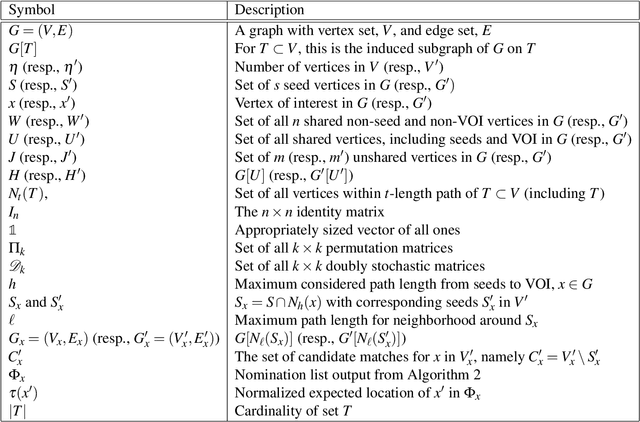
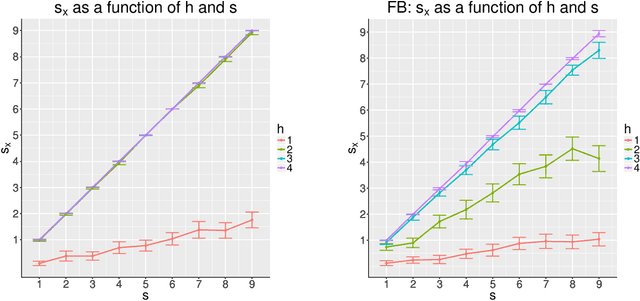
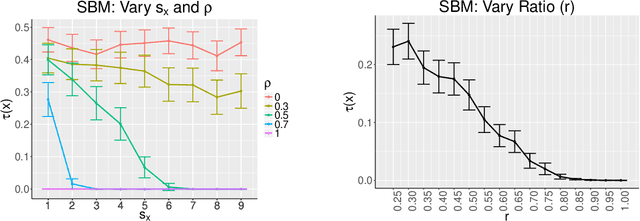
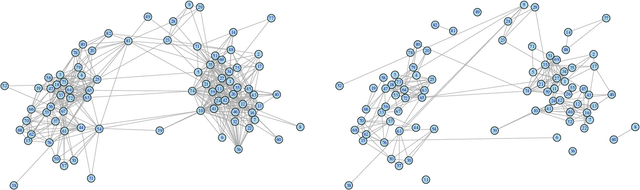
Abstract:Consider two networks on overlapping, non-identical vertex sets. Given vertices of interest in the first network, we seek to identify the corresponding vertices, if any exist, in the second network. While in moderately sized networks graph matching methods can be applied directly to recover the missing correspondences, herein we present a principled methodology appropriate for situations in which the networks are too large for brute-force graph matching. Our methodology identifies vertices in a local neighborhood of the vertices of interest in the first network that have verifiable corresponding vertices in the second network. Leveraging these known correspondences, referred to as seeds, we match the induced subgraphs in each network generated by the neighborhoods of these verified seeds, and rank the vertices of the second network in terms of the most likely matches to the original vertices of interest. We demonstrate the applicability of our methodology through simulations and real data examples.
Semiparametric spectral modeling of the Drosophila connectome
May 09, 2017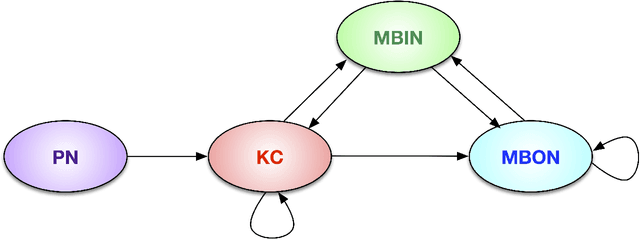
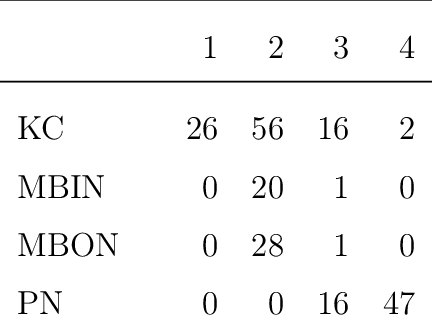
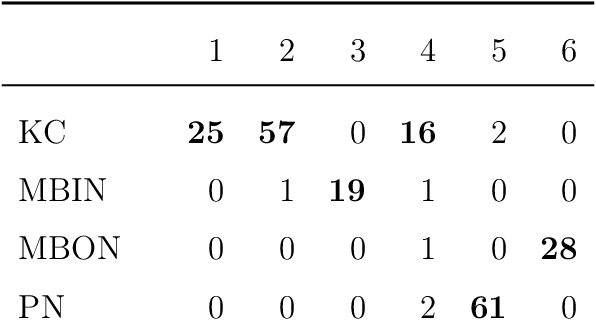
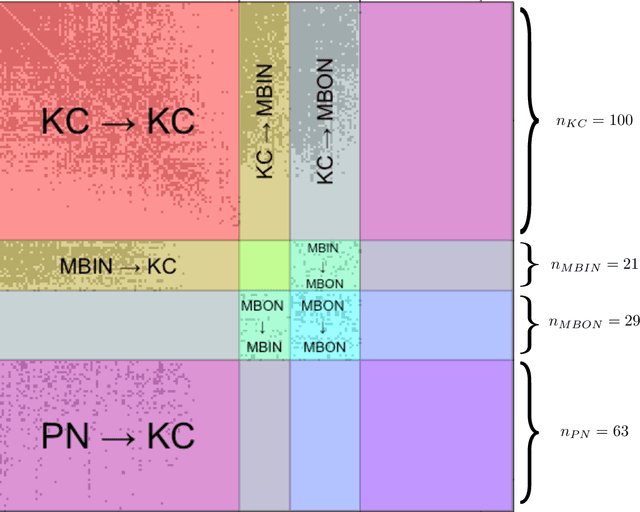

Abstract:We present semiparametric spectral modeling of the complete larval Drosophila mushroom body connectome. Motivated by a thorough exploratory data analysis of the network via Gaussian mixture modeling (GMM) in the adjacency spectral embedding (ASE) representation space, we introduce the latent structure model (LSM) for network modeling and inference. LSM is a generalization of the stochastic block model (SBM) and a special case of the random dot product graph (RDPG) latent position model, and is amenable to semiparametric GMM in the ASE representation space. The resulting connectome code derived via semiparametric GMM composed with ASE captures latent connectome structure and elucidates biologically relevant neuronal properties.
Fast Embedding for JOFC Using the Raw Stress Criterion
Oct 31, 2016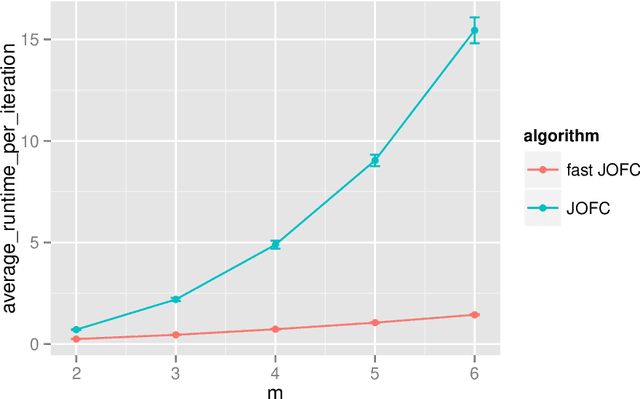
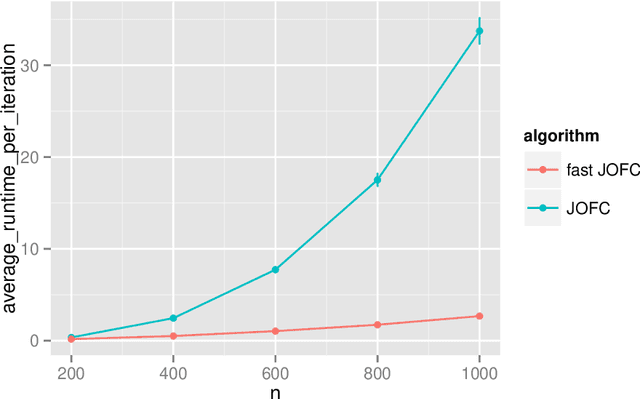
Abstract:The Joint Optimization of Fidelity and Commensurability (JOFC) manifold matching methodology embeds an omnibus dissimilarity matrix consisting of multiple dissimilarities on the same set of objects. One approach to this embedding optimizes the preservation of fidelity to each individual dissimilarity matrix together with commensurability of each given observation across modalities via iterative majorization of a raw stress error criterion by successive Guttman transforms. In this paper, we exploit the special structure inherent to JOFC to exactly and efficiently compute the successive Guttman transforms, and as a result we are able to greatly speed up the JOFC procedure for both in-sample and out-of-sample embedding. We demonstrate the scalability of our implementation on both real and simulated data examples.
On the Consistency of the Likelihood Maximization Vertex Nomination Scheme: Bridging the Gap Between Maximum Likelihood Estimation and Graph Matching
Aug 27, 2016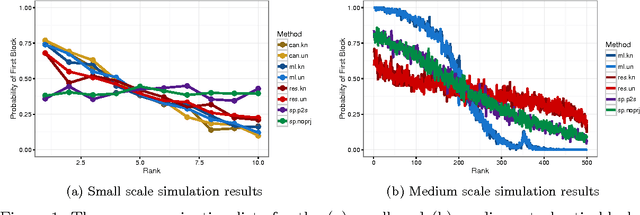

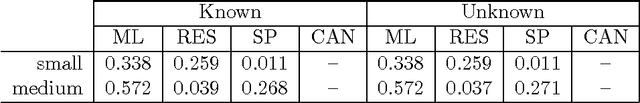

Abstract:Given a graph in which a few vertices are deemed interesting a priori, the vertex nomination task is to order the remaining vertices into a nomination list such that there is a concentration of interesting vertices at the top of the list. Previous work has yielded several approaches to this problem, with theoretical results in the setting where the graph is drawn from a stochastic block model (SBM), including a vertex nomination analogue of the Bayes optimal classifier. In this paper, we prove that maximum likelihood (ML)-based vertex nomination is consistent, in the sense that the performance of the ML-based scheme asymptotically matches that of the Bayes optimal scheme. We prove theorems of this form both when model parameters are known and unknown. Additionally, we introduce and prove consistency of a related, more scalable restricted-focus ML vertex nomination scheme. Finally, we incorporate vertex and edge features into ML-based vertex nomination and briefly explore the empirical effectiveness of this approach.
Community Detection and Classification in Hierarchical Stochastic Blockmodels
Aug 26, 2016
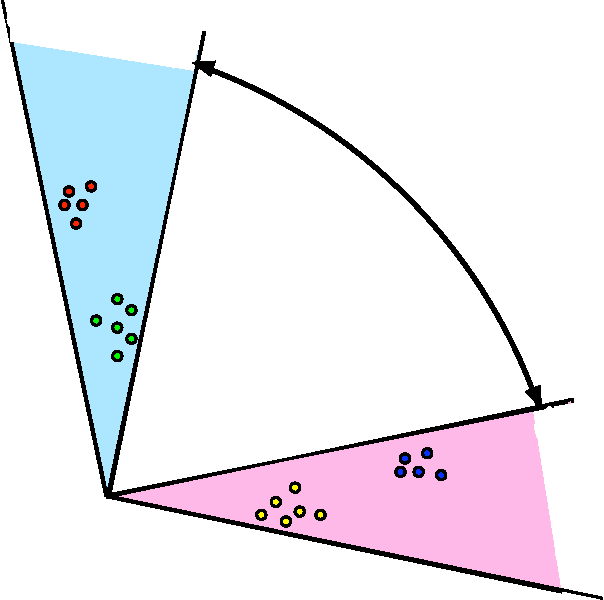
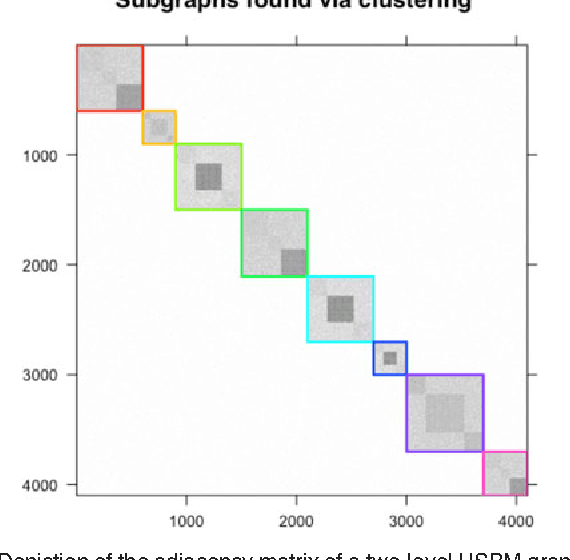
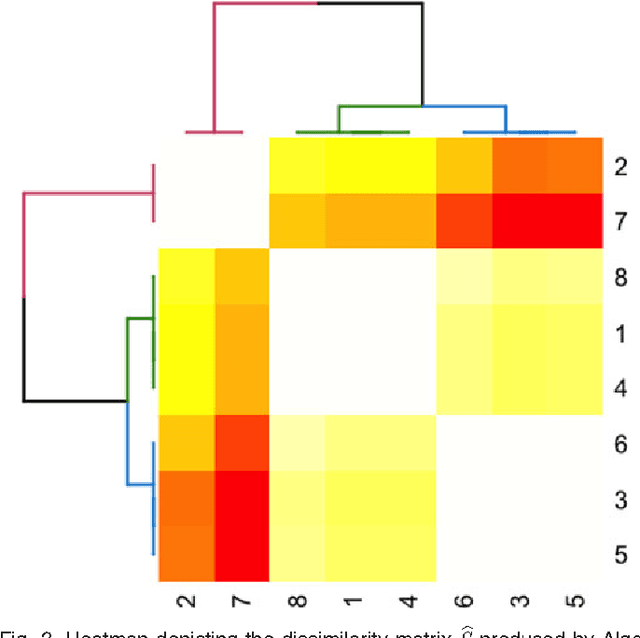
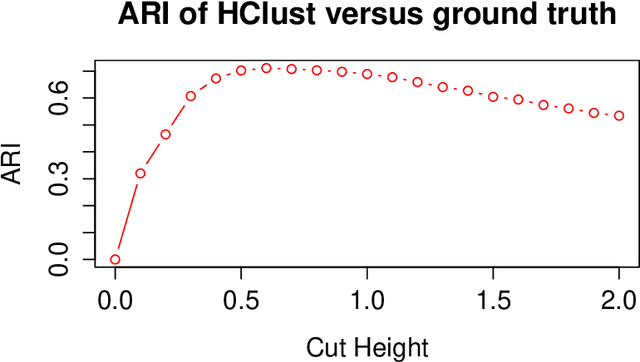
Abstract:We propose a robust, scalable, integrated methodology for community detection and community comparison in graphs. In our procedure, we first embed a graph into an appropriate Euclidean space to obtain a low-dimensional representation, and then cluster the vertices into communities. We next employ nonparametric graph inference techniques to identify structural similarity among these communities. These two steps are then applied recursively on the communities, allowing us to detect more fine-grained structure. We describe a hierarchical stochastic blockmodel---namely, a stochastic blockmodel with a natural hierarchical structure---and establish conditions under which our algorithm yields consistent estimates of model parameters and motifs, which we define to be stochastically similar groups of subgraphs. Finally, we demonstrate the effectiveness of our algorithm in both simulated and real data. Specifically, we address the problem of locating similar subcommunities in a partially reconstructed Drosophila connectome and in the social network Friendster.
Laplacian Eigenmaps from Sparse, Noisy Similarity Measurements
Aug 16, 2016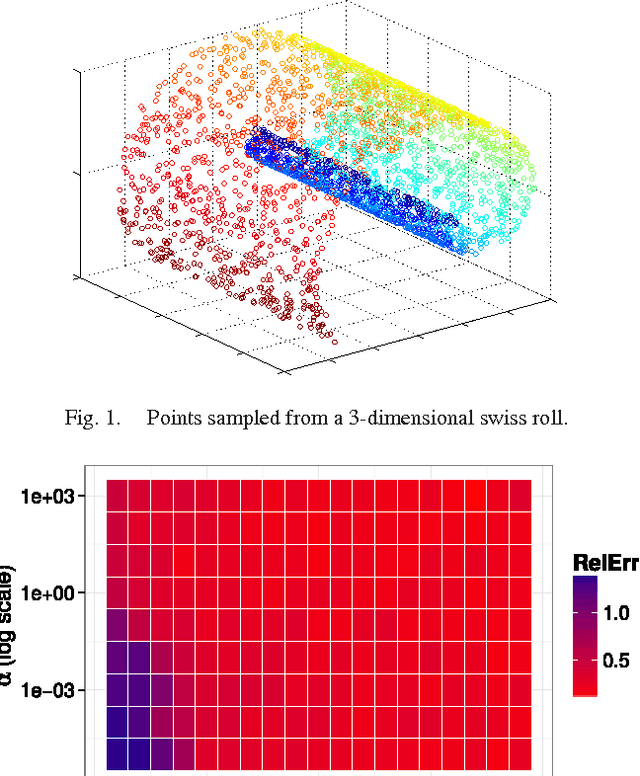
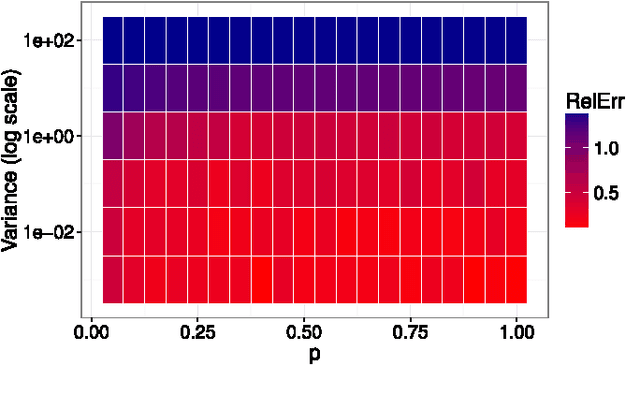
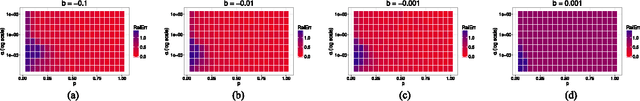
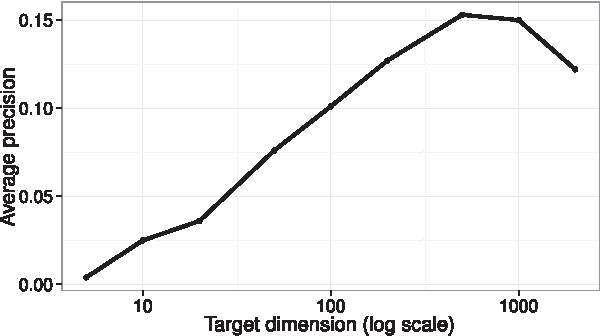
Abstract:Manifold learning and dimensionality reduction techniques are ubiquitous in science and engineering, but can be computationally expensive procedures when applied to large data sets or when similarities are expensive to compute. To date, little work has been done to investigate the tradeoff between computational resources and the quality of learned representations. We present both theoretical and experimental explorations of this question. In particular, we consider Laplacian eigenmaps embeddings based on a kernel matrix, and explore how the embeddings behave when this kernel matrix is corrupted by occlusion and noise. Our main theoretical result shows that under modest noise and occlusion assumptions, we can (with high probability) recover a good approximation to the Laplacian eigenmaps embedding based on the uncorrupted kernel matrix. Our results also show how regularization can aid this approximation. Experimentally, we explore the effects of noise and occlusion on Laplacian eigenmaps embeddings of two real-world data sets, one from speech processing and one from neuroscience, as well as a synthetic data set.
Scalable Out-of-Sample Extension of Graph Embeddings Using Deep Neural Networks
Jun 14, 2016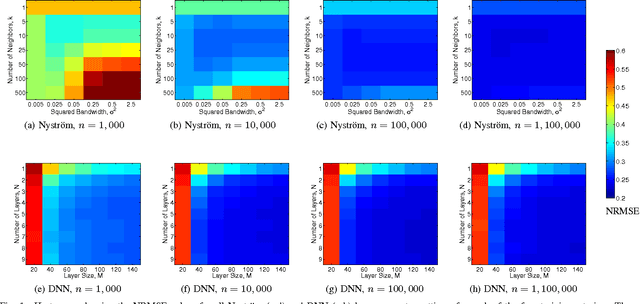
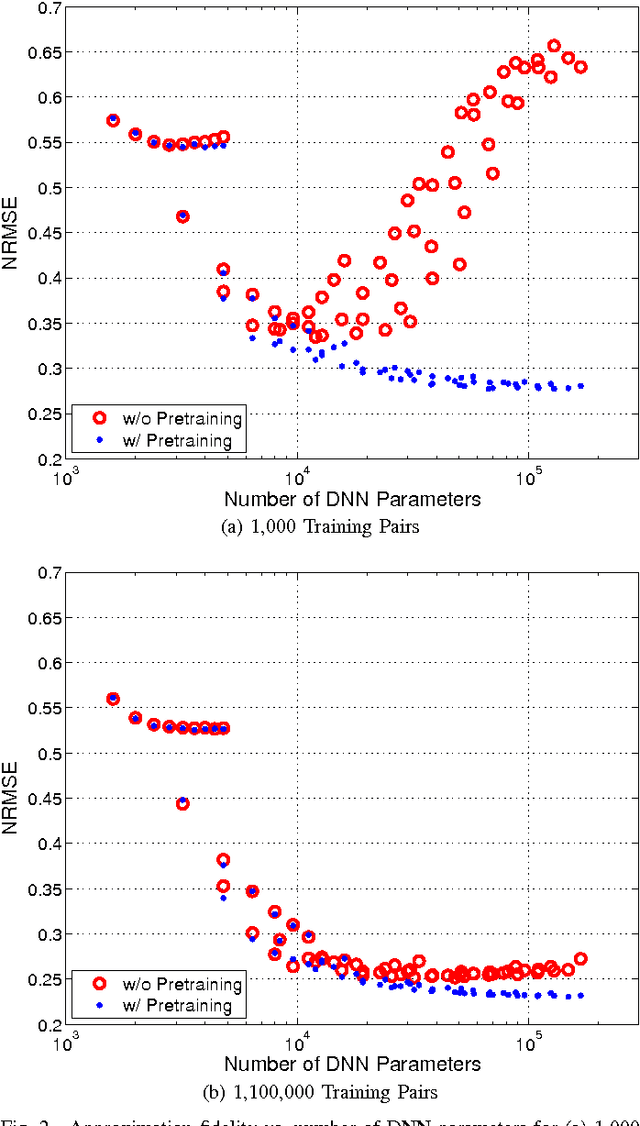
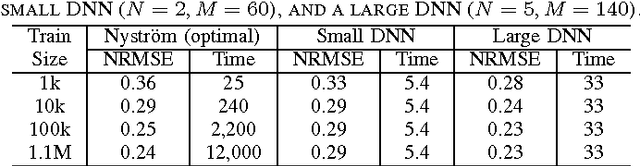
Abstract:Several popular graph embedding techniques for representation learning and dimensionality reduction rely on performing computationally expensive eigendecompositions to derive a nonlinear transformation of the input data space. The resulting eigenvectors encode the embedding coordinates for the training samples only, and so the embedding of novel data samples requires further costly computation. In this paper, we present a method for the out-of-sample extension of graph embeddings using deep neural networks (DNN) to parametrically approximate these nonlinear maps. Compared with traditional nonparametric out-of-sample extension methods, we demonstrate that the DNNs can generalize with equal or better fidelity and require orders of magnitude less computation at test time. Moreover, we find that unsupervised pretraining of the DNNs improves optimization for larger network sizes, thus removing sensitivity to model selection.
Spectral Clustering for Divide-and-Conquer Graph Matching
Mar 12, 2015Abstract:We present a parallelized bijective graph matching algorithm that leverages seeds and is designed to match very large graphs. Our algorithm combines spectral graph embedding with existing state-of-the-art seeded graph matching procedures. We justify our approach by proving that modestly correlated, large stochastic block model random graphs are correctly matched utilizing very few seeds through our divide-and-conquer procedure. We also demonstrate the effectiveness of our approach in matching very large graphs in simulated and real data examples, showing up to a factor of 8 improvement in runtime with minimal sacrifice in accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge