Tristan Naumann
Microsoft Research
Large-Scale Domain-Specific Pretraining for Biomedical Vision-Language Processing
Mar 02, 2023Abstract:Contrastive pretraining on parallel image-text data has attained great success in vision-language processing (VLP), as exemplified by CLIP and related methods. However, prior explorations tend to focus on general domains in the web. Biomedical images and text are rather different, but publicly available datasets are small and skew toward chest X-ray, thus severely limiting progress. In this paper, we conducted by far the largest study on biomedical VLP, using 15 million figure-caption pairs extracted from biomedical research articles in PubMed Central. Our dataset (PMC-15M) is two orders of magnitude larger than existing biomedical image-text datasets such as MIMIC-CXR, and spans a diverse range of biomedical images. The standard CLIP method is suboptimal for the biomedical domain. We propose BiomedCLIP with domain-specific adaptations tailored to biomedical VLP. We conducted extensive experiments and ablation studies on standard biomedical imaging tasks from retrieval to classification to visual question-answering (VQA). BiomedCLIP established new state of the art in a wide range of standard datasets, substantially outperformed prior VLP approaches. Surprisingly, BiomedCLIP even outperformed radiology-specific state-of-the-art models such as BioViL on radiology-specific tasks such as RSNA pneumonia detection, thus highlighting the utility in large-scale pretraining across all biomedical image types. We will release our models at https://aka.ms/biomedclip to facilitate future research in biomedical VLP.
Continual Contrastive Finetuning Improves Low-Resource Relation Extraction
Dec 21, 2022Abstract:Relation extraction (RE), which has relied on structurally annotated corpora for model training, has been particularly challenging in low-resource scenarios and domains. Recent literature has tackled low-resource RE by self-supervised learning, where the solution involves pretraining the relation embedding by RE-based objective and finetuning on labeled data by classification-based objective. However, a critical challenge to this approach is the gap in objectives, which prevents the RE model from fully utilizing the knowledge in pretrained representations. In this paper, we aim at bridging the gap and propose to pretrain and finetune the RE model using consistent objectives of contrastive learning. Since in this kind of representation learning paradigm, one relation may easily form multiple clusters in the representation space, we further propose a multi-center contrastive loss that allows one relation to form multiple clusters to better align with pretraining. Experiments on two document-level RE datasets, BioRED and Re-DocRED, demonstrate the effectiveness of our method. Particularly, when using 1% end-task training data, our method outperforms PLM-based RE classifier by 10.5% and 5.8% on the two datasets, respectively.
Making the Most of Text Semantics to Improve Biomedical Vision--Language Processing
Apr 21, 2022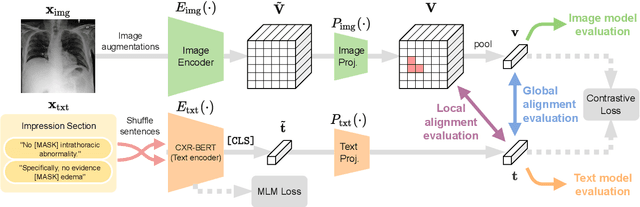
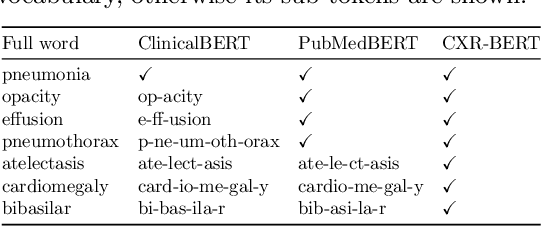
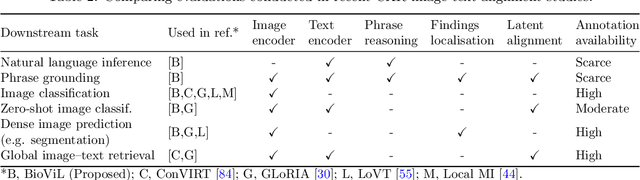

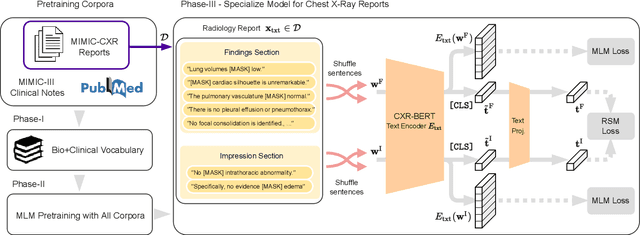

Abstract:Multi-modal data abounds in biomedicine, such as radiology images and reports. Interpreting this data at scale is essential for improving clinical care and accelerating clinical research. Biomedical text with its complex semantics poses additional challenges in vision-language modelling compared to the general domain, and previous work has used insufficiently adapted models that lack domain-specific language understanding. In this paper, we show that principled textual semantic modelling can substantially improve contrastive learning in self-supervised vision--language processing. We release a language model that achieves state-of-the-art results in radiology natural language inference through its improved vocabulary and novel language pretraining objective leveraging semantics and discourse characteristics in radiology reports. Further, we propose a self-supervised joint vision--language approach with a focus on better text modelling. It establishes new state of the art results on a wide range of publicly available benchmarks, in part by leveraging our new domain-specific language model. We release a new dataset with locally-aligned phrase grounding annotations by radiologists to facilitate the study of complex semantic modelling in biomedical vision--language processing. A broad evaluation, including on this new dataset, shows that our contrastive learning approach, aided by textual-semantic modelling, outperforms prior methods in segmentation tasks, despite only using a global-alignment objective.
Towards Structuring Real-World Data at Scale: Deep Learning for Extracting Key Oncology Information from Clinical Text with Patient-Level Supervision
Mar 20, 2022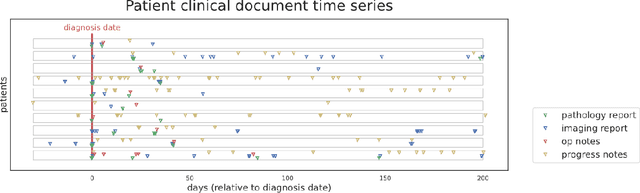
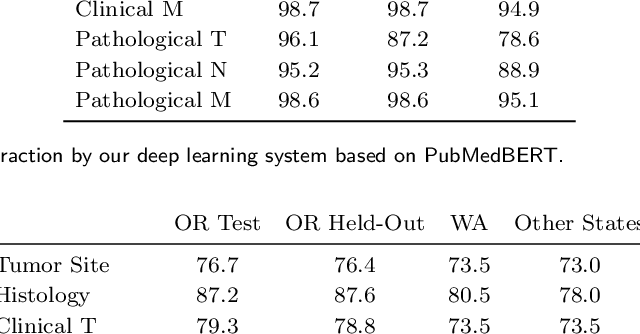
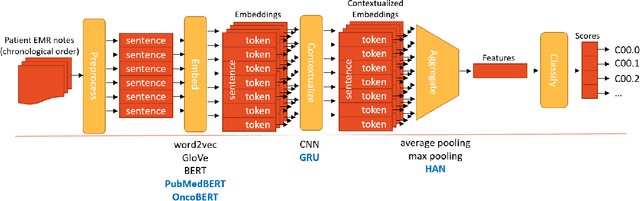
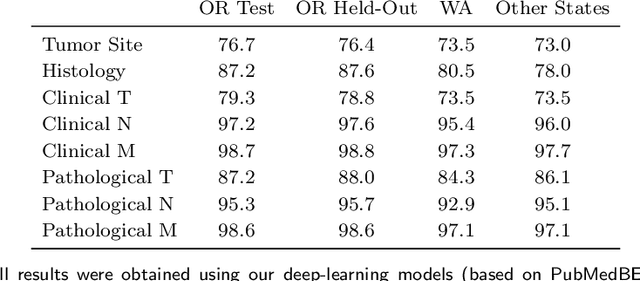
Abstract:Objective: The majority of detailed patient information in real-world data (RWD) is only consistently available in free-text clinical documents. Manual curation is expensive and time-consuming. Developing natural language processing (NLP) methods for structuring RWD is thus essential for scaling real-world evidence generation. Materials and Methods: Traditional rule-based systems are vulnerable to the prevalent linguistic variations and ambiguities in clinical text, and prior applications of machine-learning methods typically require sentence-level or report-level labeled examples that are hard to produce at scale. We propose leveraging patient-level supervision from medical registries, which are often readily available and capture key patient information, for general RWD applications. To combat the lack of sentence-level or report-level annotations, we explore advanced deep-learning methods by combining domain-specific pretraining, recurrent neural networks, and hierarchical attention. Results: We conduct an extensive study on 135,107 patients from the cancer registry of a large integrated delivery network (IDN) comprising healthcare systems in five western US states. Our deep learning methods attain test AUROC of 94-99% for key tumor attributes and comparable performance on held-out data from separate health systems and states. Discussion and Conclusion: Ablation results demonstrate clear superiority of these advanced deep-learning methods over prior approaches. Error analysis shows that our NLP system sometimes even corrects errors in registrar labels. We also conduct a preliminary investigation in accelerating registry curation and general RWD structuring via assisted curation for over 1.2 million cancer patients in this healthcare network.
Knowledge-Rich Self-Supervised Entity Linking
Dec 15, 2021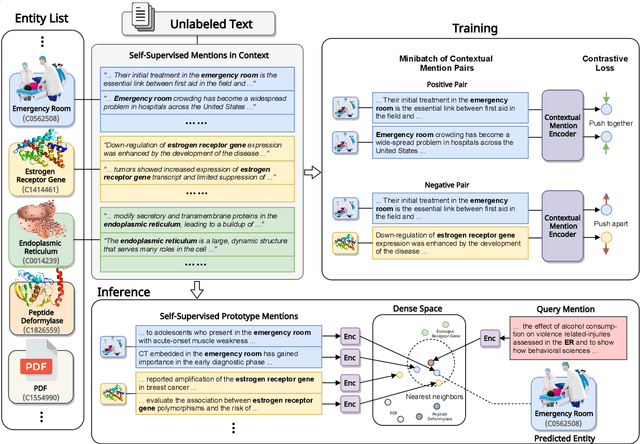
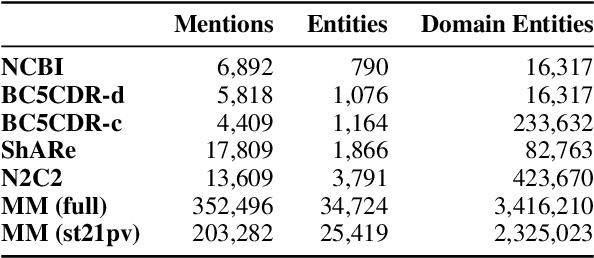
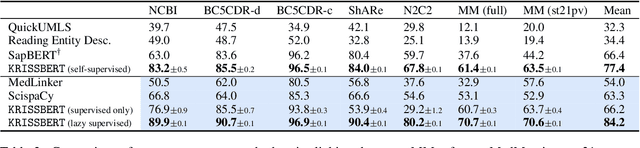
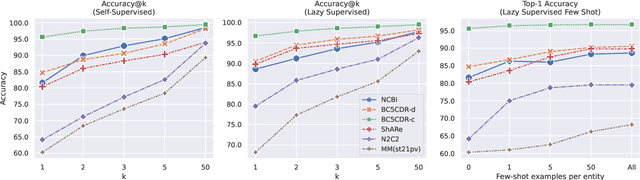
Abstract:Entity linking faces significant challenges, such as prolific variations and prevalent ambiguities, especially in high-value domains with myriad entities. Standard classification approaches suffer from the annotation bottleneck and cannot effectively handle unseen entities. Zero-shot entity linking has emerged as a promising direction for generalizing to new entities, but it still requires example gold entity mentions during training and canonical descriptions for all entities, both of which are rarely available outside of Wikipedia. In this paper, we explore Knowledge-RIch Self-Supervision ($\tt KRISS$) for entity linking, by leveraging readily available domain knowledge. In training, it generates self-supervised mention examples on unlabeled text using a domain ontology and trains a contextual encoder using contrastive learning. For inference, it samples self-supervised mentions as prototypes for each entity and conducts linking by mapping the test mention to the most similar prototype. Our approach subsumes zero-shot and few-shot methods, and can easily incorporate entity descriptions and gold mention labels if available. Using biomedicine as a case study, we conducted extensive experiments on seven standard datasets spanning biomedical literature and clinical notes. Without using any labeled information, our method produces $\tt KRISSBERT$, a universal entity linker for four million UMLS entities, which attains new state of the art, outperforming prior self-supervised methods by as much as over 20 absolute points in accuracy.
Fine-Tuning Large Neural Language Models for Biomedical Natural Language Processing
Dec 15, 2021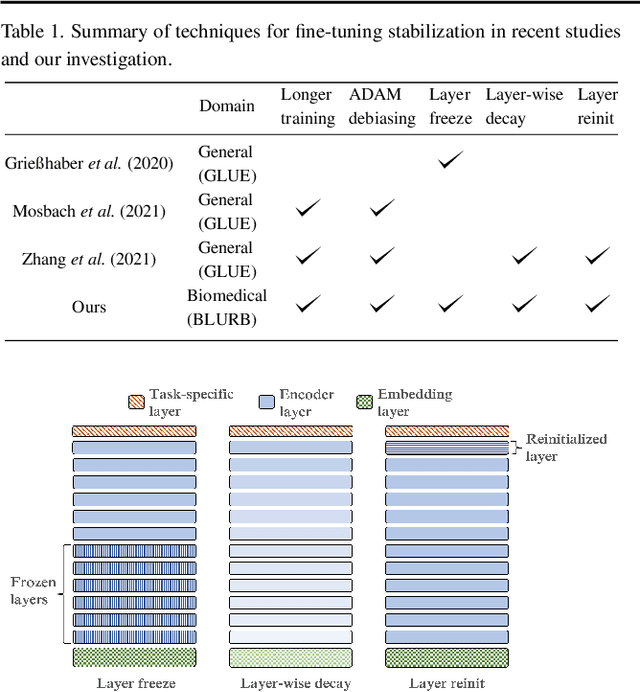
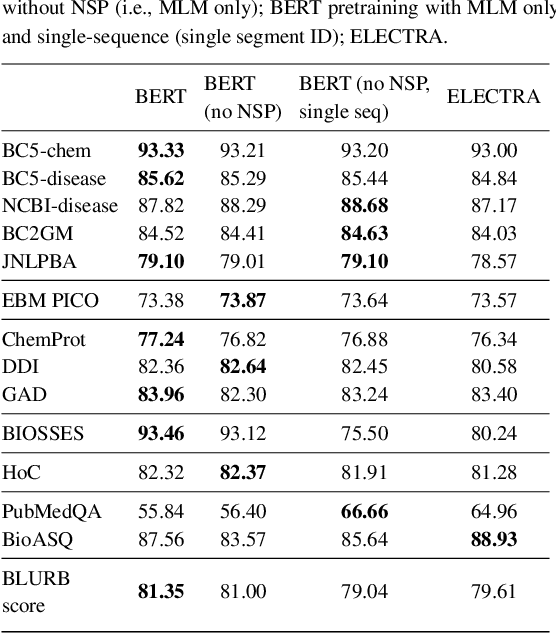
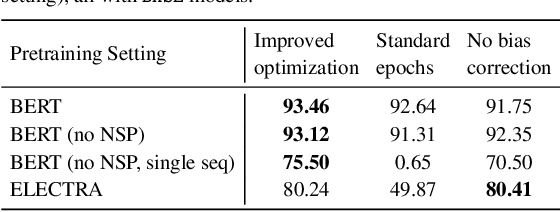
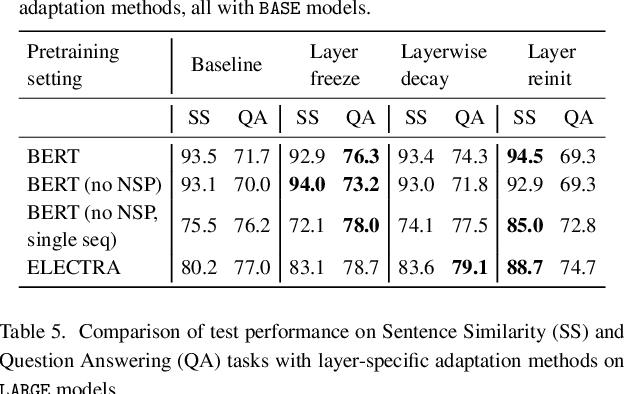
Abstract:Motivation: A perennial challenge for biomedical researchers and clinical practitioners is to stay abreast with the rapid growth of publications and medical notes. Natural language processing (NLP) has emerged as a promising direction for taming information overload. In particular, large neural language models facilitate transfer learning by pretraining on unlabeled text, as exemplified by the successes of BERT models in various NLP applications. However, fine-tuning such models for an end task remains challenging, especially with small labeled datasets, which are common in biomedical NLP. Results: We conduct a systematic study on fine-tuning stability in biomedical NLP. We show that finetuning performance may be sensitive to pretraining settings, especially in low-resource domains. Large models have potential to attain better performance, but increasing model size also exacerbates finetuning instability. We thus conduct a comprehensive exploration of techniques for addressing fine-tuning instability. We show that these techniques can substantially improve fine-tuning performance for lowresource biomedical NLP applications. Specifically, freezing lower layers is helpful for standard BERT-BASE models, while layerwise decay is more effective for BERT-LARGE and ELECTRA models. For low-resource text similarity tasks such as BIOSSES, reinitializing the top layer is the optimal strategy. Overall, domainspecific vocabulary and pretraining facilitate more robust models for fine-tuning. Based on these findings, we establish new state of the art on a wide range of biomedical NLP applications. Availability and implementation: To facilitate progress in biomedical NLP, we release our state-of-the-art pretrained and fine-tuned models: https://aka.ms/BLURB.
Modular Self-Supervision for Document-Level Relation Extraction
Sep 11, 2021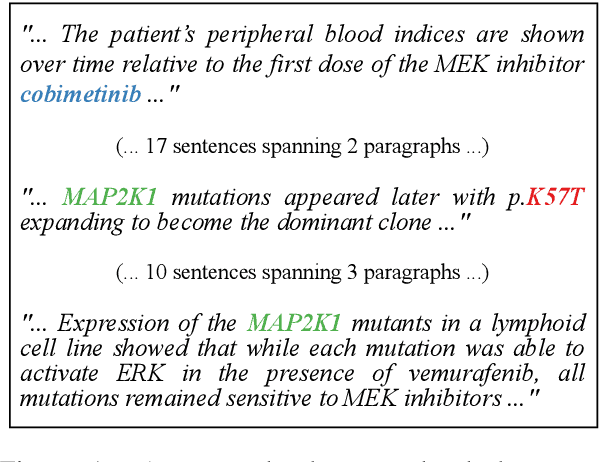
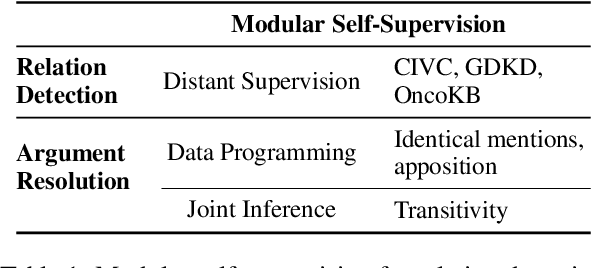
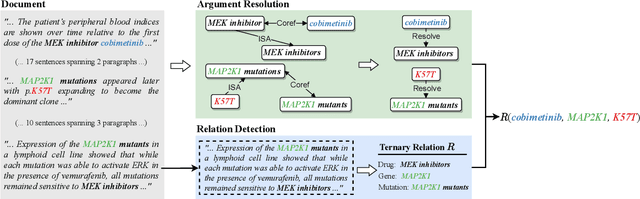
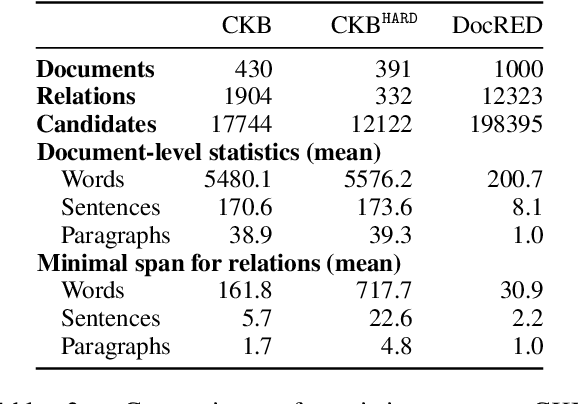
Abstract:Extracting relations across large text spans has been relatively underexplored in NLP, but it is particularly important for high-value domains such as biomedicine, where obtaining high recall of the latest findings is crucial for practical applications. Compared to conventional information extraction confined to short text spans, document-level relation extraction faces additional challenges in both inference and learning. Given longer text spans, state-of-the-art neural architectures are less effective and task-specific self-supervision such as distant supervision becomes very noisy. In this paper, we propose decomposing document-level relation extraction into relation detection and argument resolution, taking inspiration from Davidsonian semantics. This enables us to incorporate explicit discourse modeling and leverage modular self-supervision for each sub-problem, which is less noise-prone and can be further refined end-to-end via variational EM. We conduct a thorough evaluation in biomedical machine reading for precision oncology, where cross-paragraph relation mentions are prevalent. Our method outperforms prior state of the art, such as multi-scale learning and graph neural networks, by over 20 absolute F1 points. The gain is particularly pronounced among the most challenging relation instances whose arguments never co-occur in a paragraph.
Domain-Specific Pretraining for Vertical Search: Case Study on Biomedical Literature
Jun 25, 2021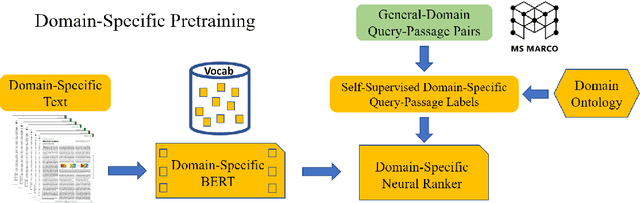
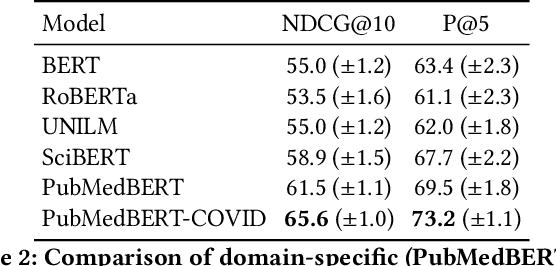
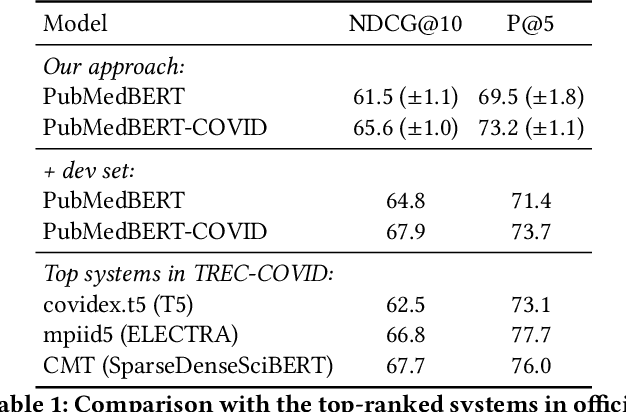

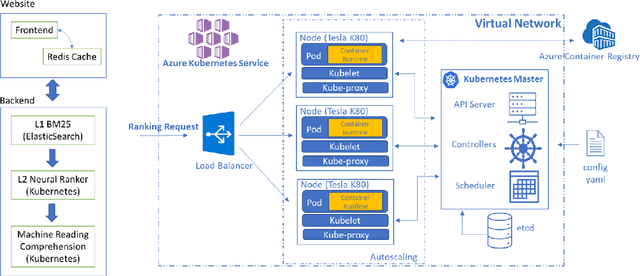
Abstract:Information overload is a prevalent challenge in many high-value domains. A prominent case in point is the explosion of the biomedical literature on COVID-19, which swelled to hundreds of thousands of papers in a matter of months. In general, biomedical literature expands by two papers every minute, totalling over a million new papers every year. Search in the biomedical realm, and many other vertical domains is challenging due to the scarcity of direct supervision from click logs. Self-supervised learning has emerged as a promising direction to overcome the annotation bottleneck. We propose a general approach for vertical search based on domain-specific pretraining and present a case study for the biomedical domain. Despite being substantially simpler and not using any relevance labels for training or development, our method performs comparably or better than the best systems in the official TREC-COVID evaluation, a COVID-related biomedical search competition. Using distributed computing in modern cloud infrastructure, our system can scale to tens of millions of articles on PubMed and has been deployed as Microsoft Biomedical Search, a new search experience for biomedical literature: https://aka.ms/biomedsearch.
Domain-Specific Language Model Pretraining for Biomedical Natural Language Processing
Aug 20, 2020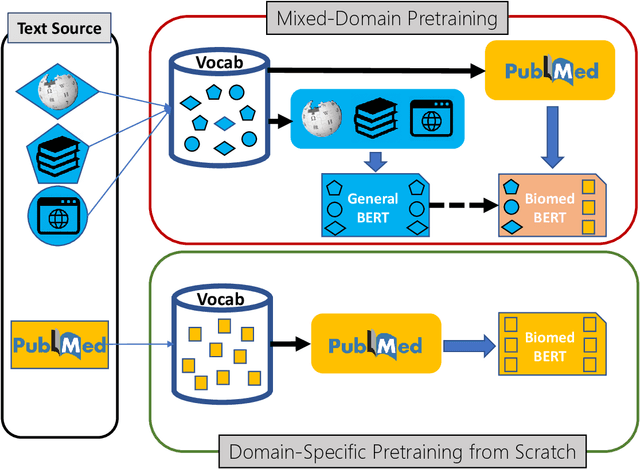
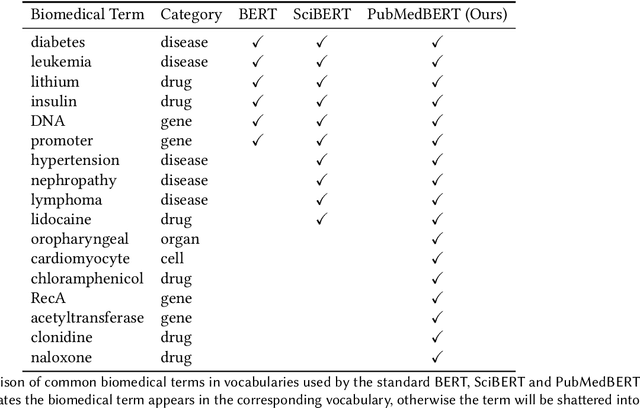
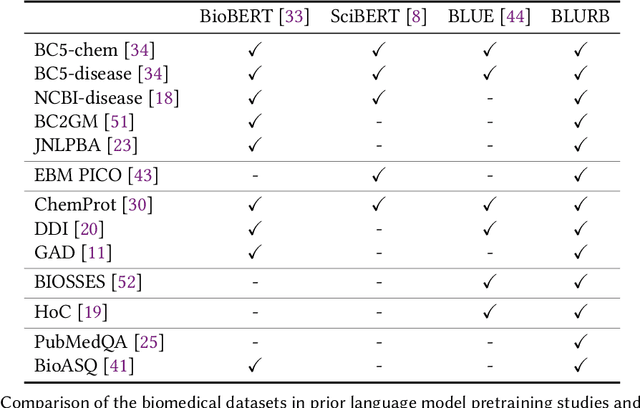
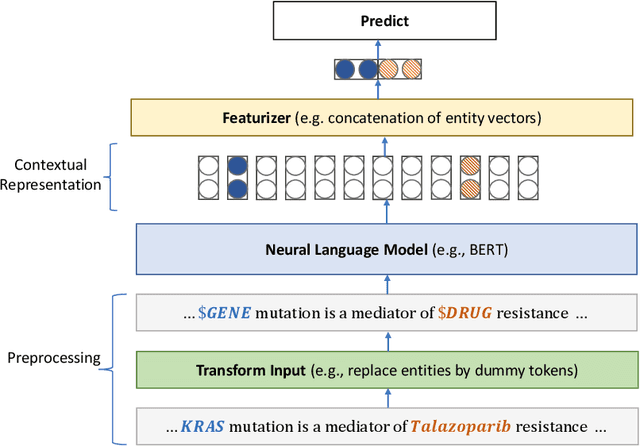
Abstract:Pretraining large neural language models, such as BERT, has led to impressive gains on many natural language processing (NLP) tasks. However, most pretraining efforts focus on general domain corpora, such as newswire and Web. A prevailing assumption is that even domain-specific pretraining can benefit by starting from general-domain language models. In this paper, we challenge this assumption by showing that for domains with abundant unlabeled text, such as biomedicine, pretraining language models from scratch results in substantial gains over continual pretraining of general-domain language models. To facilitate this investigation, we compile a comprehensive biomedical NLP benchmark from publicly-available datasets. Our experiments show that domain-specific pretraining serves as a solid foundation for a wide range of biomedical NLP tasks, leading to new state-of-the-art results across the board. Further, in conducting a thorough evaluation of modeling choices, both for pretraining and task-specific fine-tuning, we discover that some common practices are unnecessary with BERT models, such as using complex tagging schemes in named entity recognition (NER). To help accelerate research in biomedical NLP, we have released our state-of-the-art pretrained and task-specific models for the community, and created a leaderboard featuring our BLURB benchmark (short for Biomedical Language Understanding & Reasoning Benchmark) at https://aka.ms/BLURB.
ML4H Abstract Track 2019
Feb 05, 2020Abstract:A collection of the accepted abstracts for the Machine Learning for Health (ML4H) workshop at NeurIPS 2019. This index is not complete, as some accepted abstracts chose to opt-out of inclusion.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge