Srijay Deshpande
Synthesis of Annotated Colorectal Cancer Tissue Images from Gland Layout
May 08, 2023Abstract:Generating annotated pairs of realistic tissue images along with their annotations is a challenging task in computational histopathology. Such synthetic images and their annotations can be useful in training and evaluation of algorithms in the domain of computational pathology. To address this, we present an interactive framework to generate pairs of realistic colorectal cancer histology images with corresponding tissue component masks from the input gland layout. The framework shows the ability to generate realistic qualitative tissue images preserving morphological characteristics including stroma, goblet cells and glandular lumen. We show the appearance of glands can be controlled by user inputs such as number of glands, their locations and sizes. We also validate the quality of generated annotated pair with help of the gland segmentation algorithm.
CoNIC Challenge: Pushing the Frontiers of Nuclear Detection, Segmentation, Classification and Counting
Mar 14, 2023



Abstract:Nuclear detection, segmentation and morphometric profiling are essential in helping us further understand the relationship between histology and patient outcome. To drive innovation in this area, we setup a community-wide challenge using the largest available dataset of its kind to assess nuclear segmentation and cellular composition. Our challenge, named CoNIC, stimulated the development of reproducible algorithms for cellular recognition with real-time result inspection on public leaderboards. We conducted an extensive post-challenge analysis based on the top-performing models using 1,658 whole-slide images of colon tissue. With around 700 million detected nuclei per model, associated features were used for dysplasia grading and survival analysis, where we demonstrated that the challenge's improvement over the previous state-of-the-art led to significant boosts in downstream performance. Our findings also suggest that eosinophils and neutrophils play an important role in the tumour microevironment. We release challenge models and WSI-level results to foster the development of further methods for biomarker discovery.
SynCLay: Interactive Synthesis of Histology Images from Bespoke Cellular Layouts
Dec 28, 2022



Abstract:Automated synthesis of histology images has several potential applications in computational pathology. However, no existing method can generate realistic tissue images with a bespoke cellular layout or user-defined histology parameters. In this work, we propose a novel framework called SynCLay (Synthesis from Cellular Layouts) that can construct realistic and high-quality histology images from user-defined cellular layouts along with annotated cellular boundaries. Tissue image generation based on bespoke cellular layouts through the proposed framework allows users to generate different histological patterns from arbitrary topological arrangement of different types of cells. SynCLay generated synthetic images can be helpful in studying the role of different types of cells present in the tumor microenvironmet. Additionally, they can assist in balancing the distribution of cellular counts in tissue images for designing accurate cellular composition predictors by minimizing the effects of data imbalance. We train SynCLay in an adversarial manner and integrate a nuclear segmentation and classification model in its training to refine nuclear structures and generate nuclear masks in conjunction with synthetic images. During inference, we combine the model with another parametric model for generating colon images and associated cellular counts as annotations given the grade of differentiation and cell densities of different cells. We assess the generated images quantitatively and report on feedback from trained pathologists who assigned realism scores to a set of images generated by the framework. The average realism score across all pathologists for synthetic images was as high as that for the real images. We also show that augmenting limited real data with the synthetic data generated by our framework can significantly boost prediction performance of the cellular composition prediction task.
Cellular Segmentation and Composition in Routine Histology Images using Deep Learning
Mar 04, 2022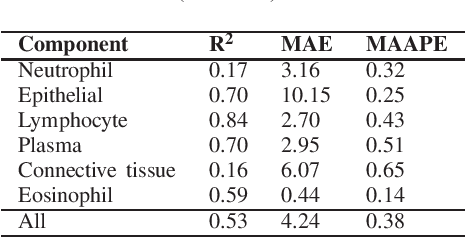
Abstract:Identification and quantification of nuclei in colorectal cancer haematoxylin \& eosin (H\&E) stained histology images is crucial to prognosis and patient management. In computational pathology these tasks are referred to as nuclear segmentation, classification and composition and are used to extract meaningful interpretable cytological and architectural features for downstream analysis. The CoNIC challenge poses the task of automated nuclei segmentation, classification and composition into six different types of nuclei from the largest publicly known nuclei dataset - Lizard. In this regard, we have developed pipelines for the prediction of nuclei segmentation using HoVer-Net and ALBRT for cellular composition. On testing on the preliminary test set, HoVer-Net achieved a PQ of 0.58, a PQ+ of 0.58 and finally a mPQ+ of 0.35. For the prediction of cellular composition with ALBRT on the preliminary test set, we achieved an overall $R^2$ score of 0.53, consisting of 0.84 for lymphocytes, 0.70 for epithelial cells, 0.70 for plasma and .060 for eosinophils.
SAFRON: Stitching Across the Frontier for Generating Colorectal Cancer Histology Images
Aug 11, 2020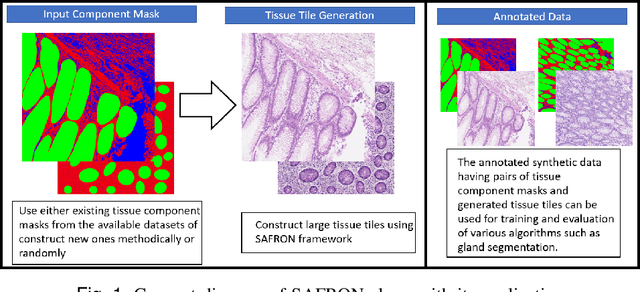
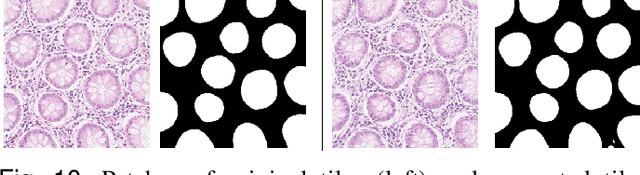
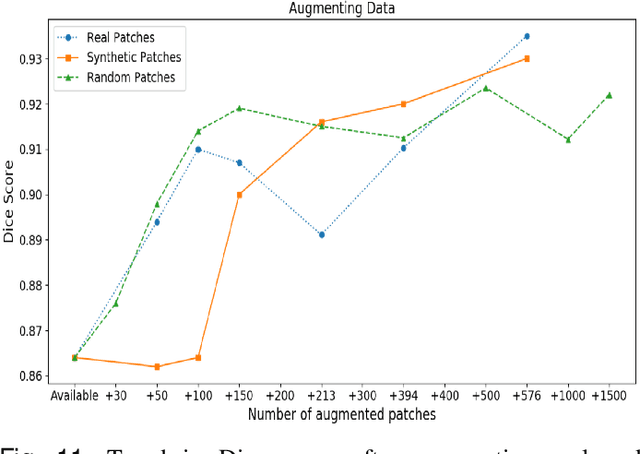

Abstract:Synthetic images can be used for the development and evaluation of deep learning algorithms in the context of limited availability of annotations. In the field of computational pathology where histology images are large and visual context is crucial, synthesis of large tissue images via generative modeling is a challenging task due to memory and computing constraints hindering the generation of large images. To address this challenge, we propose a novel framework named as SAFRON to construct realistic large tissue image tiles from ground truth annotations while preserving morphological features and with minimal boundary artifacts at the seams. To this end, we train the proposed SAFRON framework based on conditional generative adversarial networks on large tissue image tiles from the Colorectal Adenocarcinoma Gland (CRAG) and DigestPath datasets. We demonstrate that our model can generate high quality and realistic image tiles of arbitrary large size after training it on relatively small image patches. We also show that training on synthetic data generated by SAFRON can significantly boost the performance of a standard algorithm for gland segmentation of colorectal cancer tissue images. Sample high resolution images generated using SAFRON are available at the URL:https://warwick.ac.uk/TIALab/SAFRON
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge