Shadi Albarqouni
University Hospital Bonn, Venusberg-Campus 1, D-53127, Bonn, Germany, Helmholtz Munich, Ingolstädter Landstraße 1, D-85764, Neuherberg, Germany, Technical University of Munich, Boltzmannstr. 3, D-85748 Garching, Germany
Polyp-artifact relationship analysis using graph inductive learned representations
Sep 15, 2020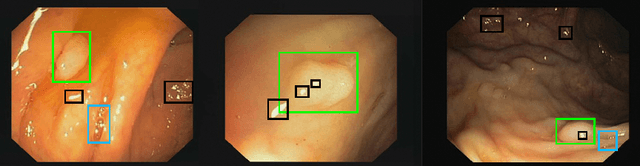
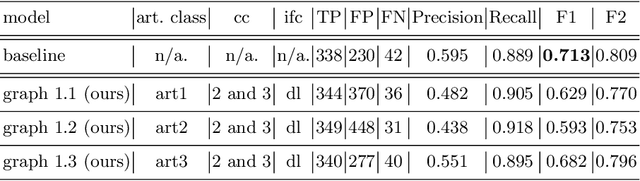

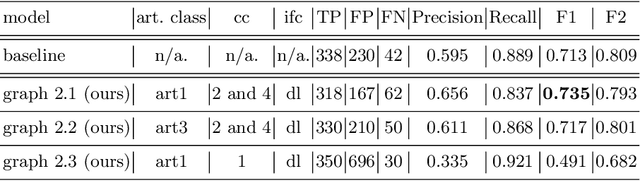
Abstract:The diagnosis process of colorectal cancer mainly focuses on the localization and characterization of abnormal growths in the colon tissue known as polyps. Despite recent advances in deep object localization, the localization of polyps remains challenging due to the similarities between tissues, and the high level of artifacts. Recent studies have shown the negative impact of the presence of artifacts in the polyp detection task, and have started to take them into account within the training process. However, the use of prior knowledge related to the spatial interaction of polyps and artifacts has not yet been considered. In this work, we incorporate artifact knowledge in a post-processing step. Our method models this task as an inductive graph representation learning problem, and is composed of training and inference steps. Detected bounding boxes around polyps and artifacts are considered as nodes connected by a defined criterion. The training step generates a node classifier with ground truth bounding boxes. In inference, we use this classifier to analyze a second graph, generated from artifact and polyp predictions given by region proposal networks. We evaluate how the choices in the connectivity and artifacts affect the performance of our method and show that it has the potential to reduce the false positives in the results of a region proposal network.
Inverse Distance Aggregation for Federated Learning with Non-IID Data
Aug 17, 2020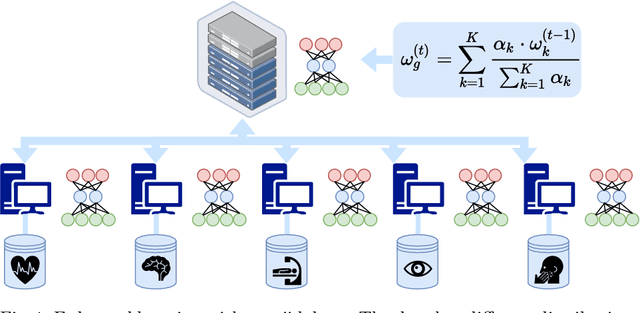
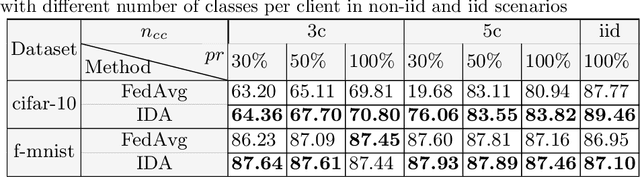
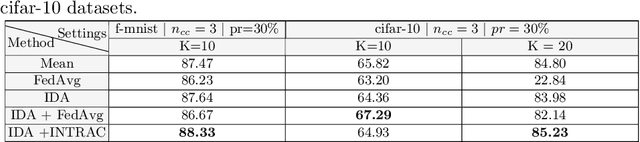
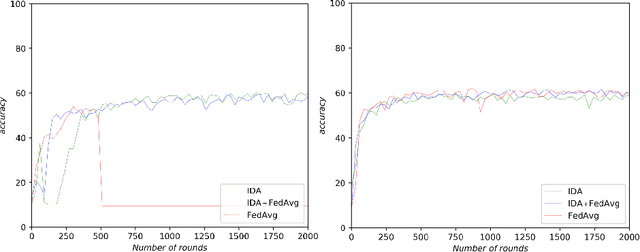
Abstract:Federated learning (FL) has been a promising approach in the field of medical imaging in recent years. A critical problem in FL, specifically in medical scenarios is to have a more accurate shared model which is robust to noisy and out-of distribution clients. In this work, we tackle the problem of statistical heterogeneity in data for FL which is highly plausible in medical data where for example the data comes from different sites with different scanner settings. We propose IDA (Inverse Distance Aggregation), a novel adaptive weighting approach for clients based on meta-information which handles unbalanced and non-iid data. We extensively analyze and evaluate our method against the well-known FL approach, Federated Averaging as a baseline.
Attention based Multiple Instance Learning for Classification of Blood Cell Disorders
Jul 22, 2020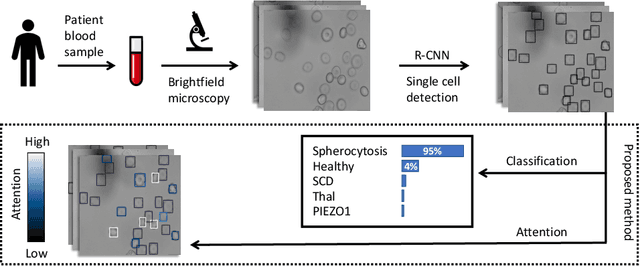
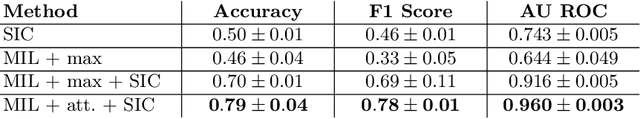
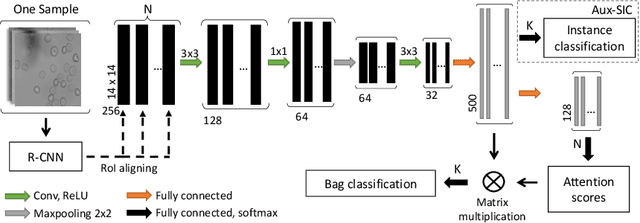
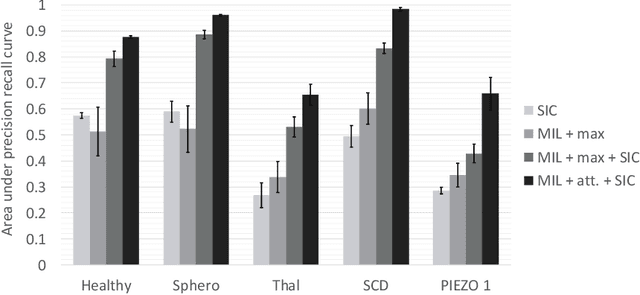
Abstract:Red blood cells are highly deformable and present in various shapes. In blood cell disorders, only a subset of all cells is morphologically altered and relevant for the diagnosis. However, manually labeling of all cells is laborious, complicated and introduces inter-expert variability. We propose an attention based multiple instance learning method to classify blood samples of patients suffering from blood cell disorders. Cells are detected using an R-CNN architecture. With the features extracted for each cell, a multiple instance learning method classifies patient samples into one out of four blood cell disorders. The attention mechanism provides a measure of the contribution of each cell to the overall classification and significantly improves the network's classification accuracy as well as its interpretability for the medical expert.
Scale-Space Autoencoders for Unsupervised Anomaly Segmentation in Brain MRI
Jun 23, 2020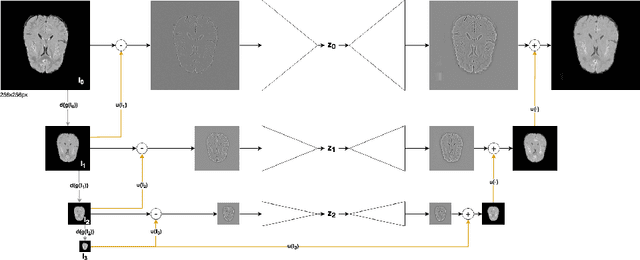
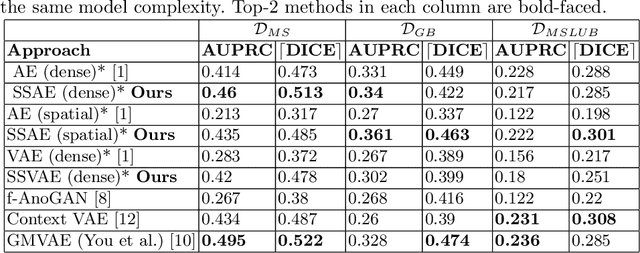
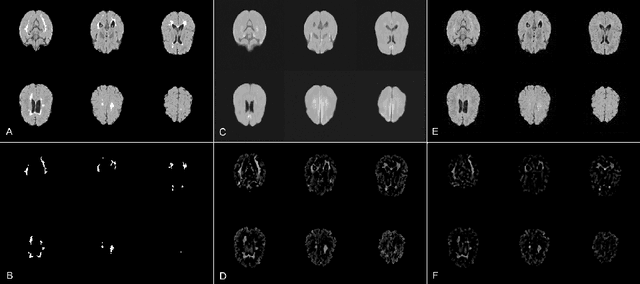
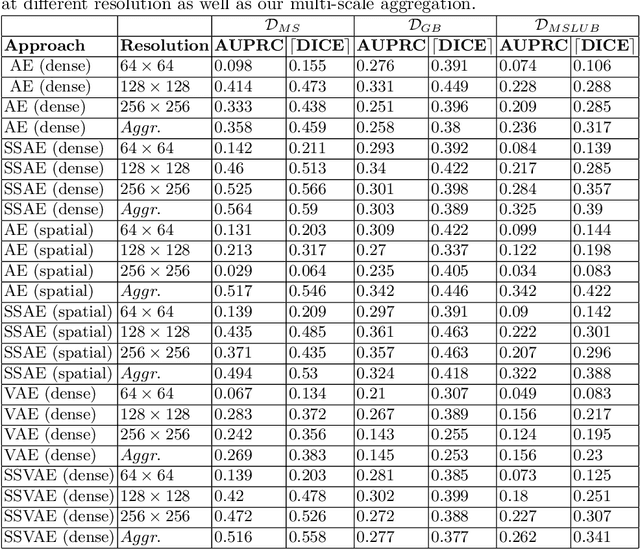
Abstract:Brain pathologies can vary greatly in size and shape, ranging from few pixels (i.e. MS lesions) to large, space-occupying tumors. Recently proposed Autoencoder-based methods for unsupervised anomaly segmentation in brain MRI have shown promising performance, but face difficulties in modeling distributions with high fidelity, which is crucial for accurate delineation of particularly small lesions. Here, similar to these previous works, we model the distribution of healthy brain MRI to localize pathologies from erroneous reconstructions. However, to achieve improved reconstruction fidelity at higher resolutions, we learn to compress and reconstruct different frequency bands of healthy brain MRI using the laplacian pyramid. In a range of experiments comparing our method to different State-of-the-Art approaches on three different brain MR datasets with MS lesions and tumors, we show improved anomaly segmentation performance and the general capability to obtain much more crisp reconstructions of input data at native resolution. The modeling of the laplacian pyramid further enables the delineation and aggregation of lesions at multiple scales, which allows to effectively cope with different pathologies and lesion sizes using a single model.
6D Camera Relocalization in Ambiguous Scenes via Continuous Multimodal Inference
Apr 09, 2020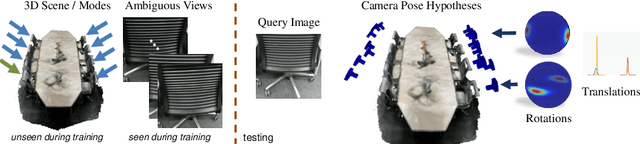
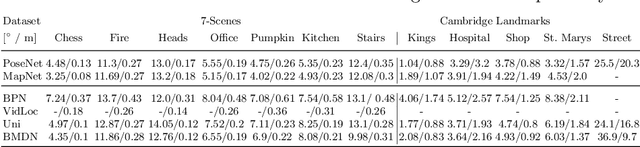
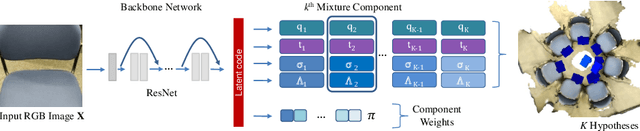
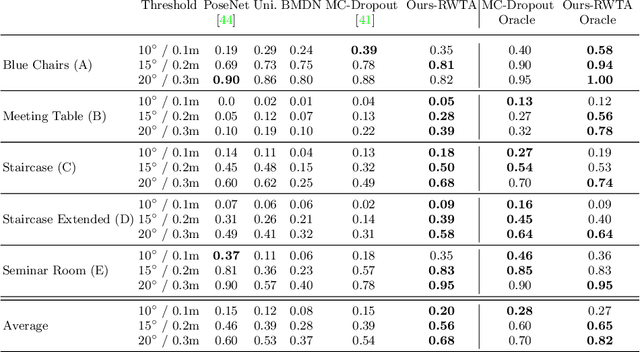
Abstract:We present a multimodal camera relocalization framework that captures ambiguities and uncertainties with continuous mixture models defined on the manifold of camera poses. In highly ambiguous environments, which can easily arise due to symmetries and repetitive structures in the scene, computing one plausible solution (what most state-of-the-art methods currently regress) may not be sufficient. Instead we predict multiple camera pose hypotheses as well as the respective uncertainty for each prediction. Towards this aim, we use Bingham distributions, to model the orientation of the camera pose, and a multivariate Gaussian to model the position, with an end-to-end deep neural network. By incorporating a Winner-Takes-All training scheme, we finally obtain a mixture model that is well suited for explaining ambiguities in the scene, yet does not suffer from mode collapse, a common problem with mixture density networks. We introduce a new dataset specifically designed to foster camera localization research in ambiguous environments and exhaustively evaluate our method on synthetic as well as real data on both ambiguous scenes and on non-ambiguous benchmark datasets. We plan to release our code and dataset under $\href{https://multimodal3dvision.github.io}{multimodal3dvision.github.io}$.
Autoencoders for Unsupervised Anomaly Segmentation in Brain MR Images: A Comparative Study
Apr 08, 2020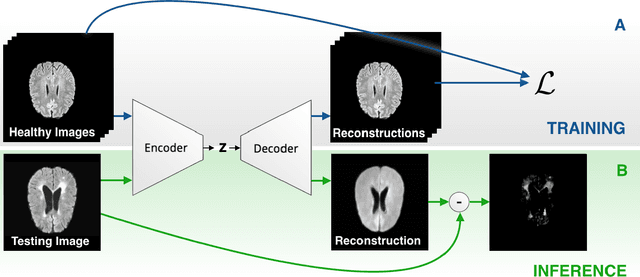
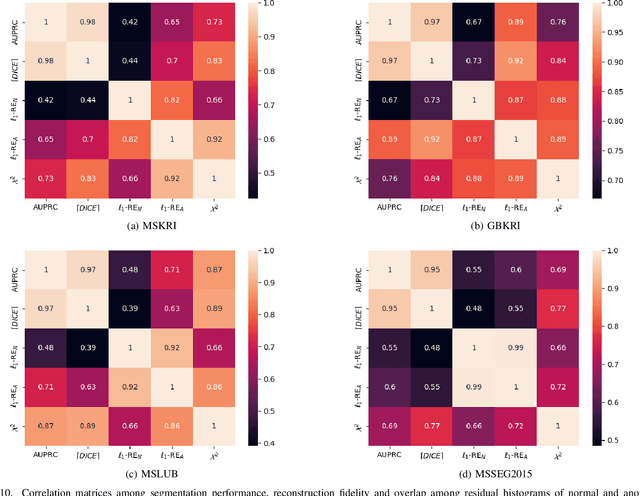
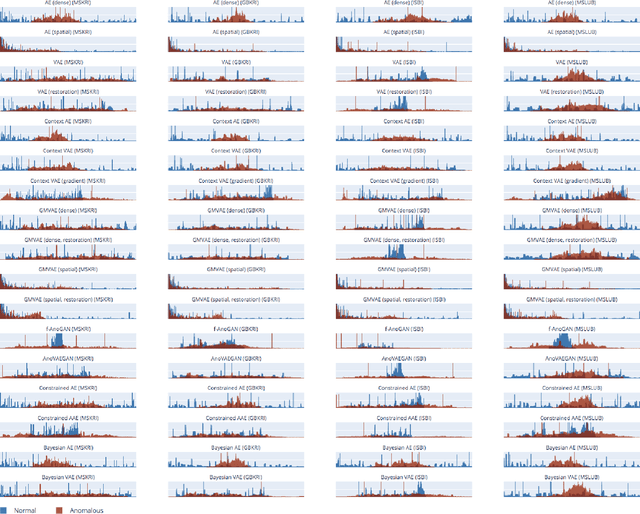
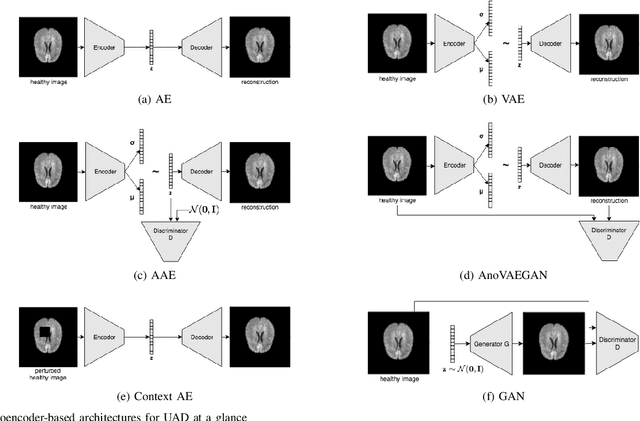
Abstract:Deep unsupervised representation learning has recently led to new approaches in the field of Unsupervised Anomaly Detection (UAD) in brain MRI. The main principle behind these works is to learn a model of normal anatomy by learning to compress and recover healthy data. This allows to spot abnormal structures from erroneous recoveries of compressed, potentially anomalous samples. The concept is of great interest to the medical image analysis community as it i) relieves from the need of vast amounts of manually segmented training data---a necessity for and pitfall of current supervised Deep Learning---and ii) theoretically allows to detect arbitrary, even rare pathologies which supervised approaches might fail to find. To date, the experimental design of most works hinders a valid comparison, because i) they are evaluated against different datasets and different pathologies, ii) use different image resolutions and iii) different model architectures with varying complexity. The intent of this work is to establish comparability among recent methods by utilizing a single architecture, a single resolution and the same dataset(s). Besides providing a ranking of the methods, we also try to answer questions like i) how many healthy training subjects are needed to model normality and ii) if the reviewed approaches are also sensitive to domain shift. Further, we identify open challenges and provide suggestions for future community efforts and research directions.
Fairness by Learning Orthogonal Disentangled Representations
Mar 22, 2020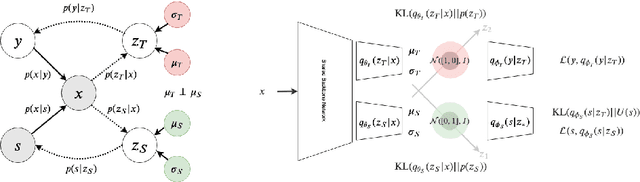
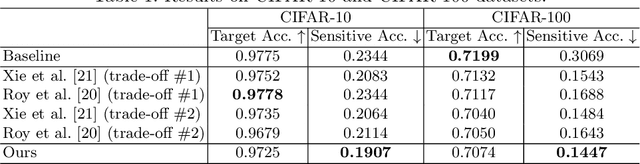
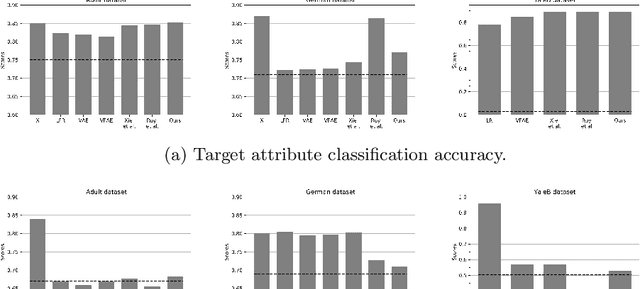
Abstract:Learning discriminative powerful representations is a crucial step for machine learning systems. Introducing invariance against arbitrary nuisance or sensitive attributes while performing well on specific tasks is an important problem in representation learning. This is mostly approached by purging the sensitive information from learned representations. In this paper, we propose a novel disentanglement approach to invariant representation problem. We disentangle the meaningful and sensitive representations by enforcing orthogonality constraints as a proxy for independence. We explicitly enforce the meaningful representation to be agnostic to sensitive information by entropy maximization. The proposed approach is evaluated on five publicly available datasets and compared with state of the art methods for learning fairness and invariance achieving the state of the art performance on three datasets and comparable performance on the rest. Further, we perform an ablative study to evaluate the effect of each component.
ROAM: Random Layer Mixup for Semi-Supervised Learning in Medical Imaging
Mar 20, 2020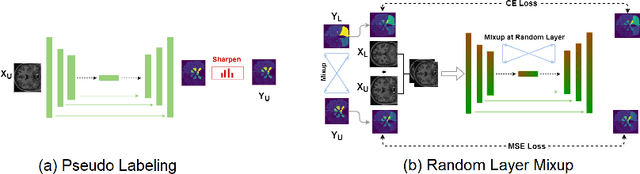
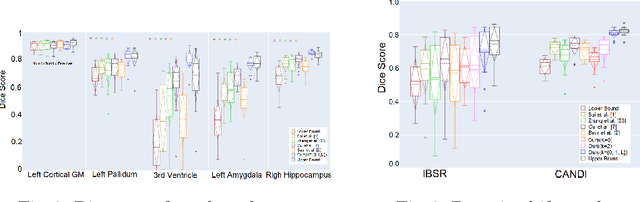
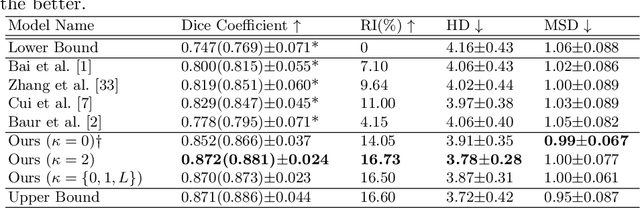

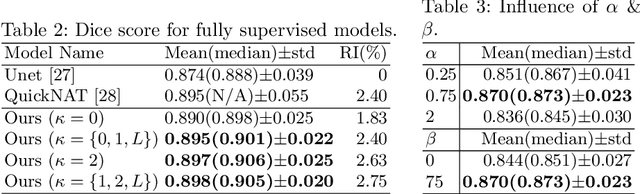
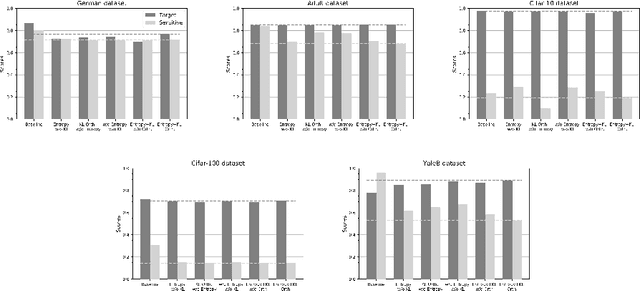
Abstract:Medical image segmentation is one of the major challenges addressed by machine learning methods. Yet, deep learning methods profoundly depend on a huge amount of annotated data which is time-consuming and costly. Though semi-supervised learning methods approach this problem by leveraging an abundant amount of unlabeled data along with a small amount of labeled data in the training process. Recently, MixUp regularizer [32] has been successfully introduced to semi-supervised learning methods showing superior performance [3]. MixUp augments the model with new data points through linear interpolation of the data at the input space. In this paper, we argue that this option is limited, instead, we propose ROAM, a random layer mixup, which encourages the network to be less confident for interpolated data points at randomly selected space. Hence, avoids over-fitting and enhances the generalization ability. We validate our method on publicly available datasets on whole-brain image segmentation (MALC) achieving state-of-the-art results in fully supervised (89.8%) and semi-supervised (87.2%) settings with relative improvement up to 2.75% and 16.73%, respectively.
The Future of Digital Health with Federated Learning
Mar 18, 2020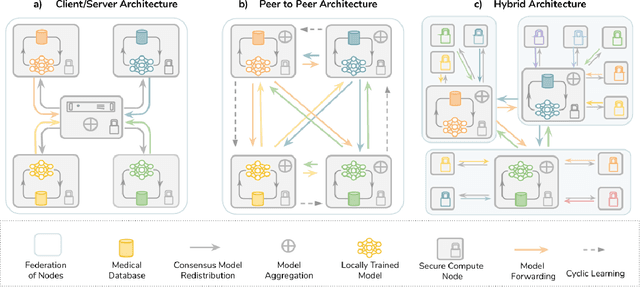
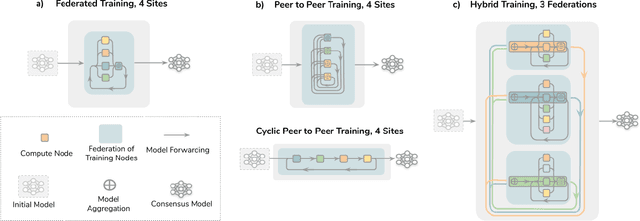
Abstract:Data-driven Machine Learning has emerged as a promising approach for building accurate and robust statistical models from medical data, which is collected in huge volumes by modern healthcare systems. Existing medical data is not fully exploited by ML primarily because it sits in data silos and privacy concerns restrict access to this data. However, without access to sufficient data, ML will be prevented from reaching its full potential and, ultimately, from making the transition from research to clinical practice. This paper considers key factors contributing to this issue, explores how Federated Learning (FL) may provide a solution for the future of digital health and highlights the challenges and considerations that need to be addressed.
A learning without forgetting approach to incorporate artifact knowledge in polyp localization tasks
Feb 11, 2020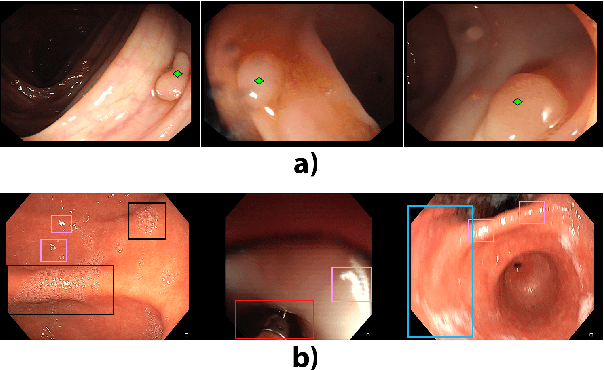
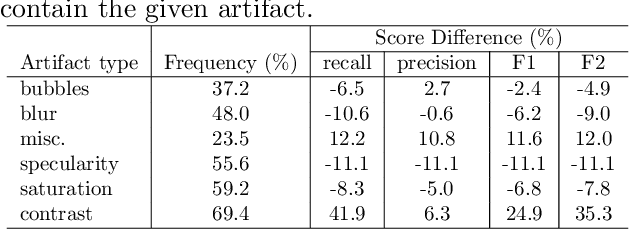
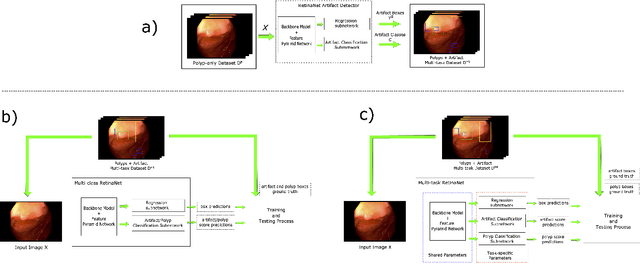
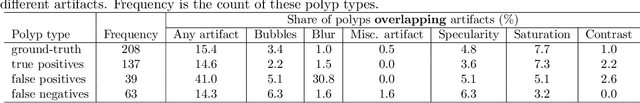
Abstract:Colorectal polyps are abnormalities in the colon tissue that can develop into colorectal cancer. The survival rate for patients is higher when the disease is detected at an early stage and polyps can be removed before they develop into malignant tumors. Deep learning methods have become the state of art in automatic polyp detection. However, the performance of current models heavily relies on the size and quality of the training datasets. Endoscopic video sequences tend to be corrupted by different artifacts affecting visibility and hence, the detection rates. In this work, we analyze the effects that artifacts have in the polyp localization problem. For this, we evaluate the RetinaNet architecture, originally defined for object localization. We also define a model inspired by the learning without forgetting framework, which allows us to employ artifact detection knowledge in the polyp localization problem. Finally, we perform several experiments to analyze the influence of the artifacts in the performance of these models. To our best knowledge, this is the first extensive analysis of the influence of artifact in polyp localization and the first work incorporating learning without forgetting ideas for simultaneous artifact and polyp localization tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge