Sam Foreman
Extending $μ$P: Spectral Conditions for Feature Learning Across Optimizers
Feb 24, 2026Abstract:Several variations of adaptive first-order and second-order optimization methods have been proposed to accelerate and scale the training of large language models. The performance of these optimization routines is highly sensitive to the choice of hyperparameters (HPs), which are computationally expensive to tune for large-scale models. Maximal update parameterization $(μ$P$)$ is a set of scaling rules which aims to make the optimal HPs independent of the model size, thereby allowing the HPs tuned on a smaller (computationally cheaper) model to be transferred to train a larger, target model. Despite promising results for SGD and Adam, deriving $μ$P for other optimizers is challenging because the underlying tensor programming approach is difficult to grasp. Building on recent work that introduced spectral conditions as an alternative to tensor programs, we propose a novel framework to derive $μ$P for a broader class of optimizers, including AdamW, ADOPT, LAMB, Sophia, Shampoo and Muon. We implement our $μ$P derivations on multiple benchmark models and demonstrate zero-shot learning rate transfer across increasing model width for the above optimizers. Further, we provide empirical insights into depth-scaling parameterization for these optimizers.
HiPerRAG: High-Performance Retrieval Augmented Generation for Scientific Insights
May 07, 2025Abstract:The volume of scientific literature is growing exponentially, leading to underutilized discoveries, duplicated efforts, and limited cross-disciplinary collaboration. Retrieval Augmented Generation (RAG) offers a way to assist scientists by improving the factuality of Large Language Models (LLMs) in processing this influx of information. However, scaling RAG to handle millions of articles introduces significant challenges, including the high computational costs associated with parsing documents and embedding scientific knowledge, as well as the algorithmic complexity of aligning these representations with the nuanced semantics of scientific content. To address these issues, we introduce HiPerRAG, a RAG workflow powered by high performance computing (HPC) to index and retrieve knowledge from more than 3.6 million scientific articles. At its core are Oreo, a high-throughput model for multimodal document parsing, and ColTrast, a query-aware encoder fine-tuning algorithm that enhances retrieval accuracy by using contrastive learning and late-interaction techniques. HiPerRAG delivers robust performance on existing scientific question answering benchmarks and two new benchmarks introduced in this work, achieving 90% accuracy on SciQ and 76% on PubMedQA-outperforming both domain-specific models like PubMedGPT and commercial LLMs such as GPT-4. Scaling to thousands of GPUs on the Polaris, Sunspot, and Frontier supercomputers, HiPerRAG delivers million document-scale RAG workflows for unifying scientific knowledge and fostering interdisciplinary innovation.
DeepSpeed4Science Initiative: Enabling Large-Scale Scientific Discovery through Sophisticated AI System Technologies
Oct 11, 2023



Abstract:In the upcoming decade, deep learning may revolutionize the natural sciences, enhancing our capacity to model and predict natural occurrences. This could herald a new era of scientific exploration, bringing significant advancements across sectors from drug development to renewable energy. To answer this call, we present DeepSpeed4Science initiative (deepspeed4science.ai) which aims to build unique capabilities through AI system technology innovations to help domain experts to unlock today's biggest science mysteries. By leveraging DeepSpeed's current technology pillars (training, inference and compression) as base technology enablers, DeepSpeed4Science will create a new set of AI system technologies tailored for accelerating scientific discoveries by addressing their unique complexity beyond the common technical approaches used for accelerating generic large language models (LLMs). In this paper, we showcase the early progress we made with DeepSpeed4Science in addressing two of the critical system challenges in structural biology research.
A Comprehensive Performance Study of Large Language Models on Novel AI Accelerators
Oct 06, 2023



Abstract:Artificial intelligence (AI) methods have become critical in scientific applications to help accelerate scientific discovery. Large language models (LLMs) are being considered as a promising approach to address some of the challenging problems because of their superior generalization capabilities across domains. The effectiveness of the models and the accuracy of the applications is contingent upon their efficient execution on the underlying hardware infrastructure. Specialized AI accelerator hardware systems have recently become available for accelerating AI applications. However, the comparative performance of these AI accelerators on large language models has not been previously studied. In this paper, we systematically study LLMs on multiple AI accelerators and GPUs and evaluate their performance characteristics for these models. We evaluate these systems with (i) a micro-benchmark using a core transformer block, (ii) a GPT- 2 model, and (iii) an LLM-driven science use case, GenSLM. We present our findings and analyses of the models' performance to better understand the intrinsic capabilities of AI accelerators. Furthermore, our analysis takes into account key factors such as sequence lengths, scaling behavior, sparsity, and sensitivity to gradient accumulation steps.
Applications of Machine Learning to Lattice Quantum Field Theory
Feb 10, 2022Abstract:There is great potential to apply machine learning in the area of numerical lattice quantum field theory, but full exploitation of that potential will require new strategies. In this white paper for the Snowmass community planning process, we discuss the unique requirements of machine learning for lattice quantum field theory research and outline what is needed to enable exploration and deployment of this approach in the future.
HMC with Normalizing Flows
Dec 02, 2021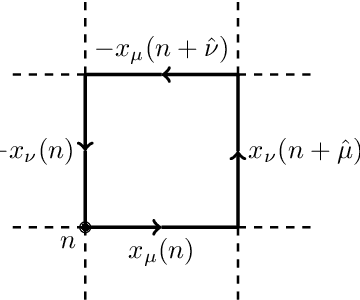
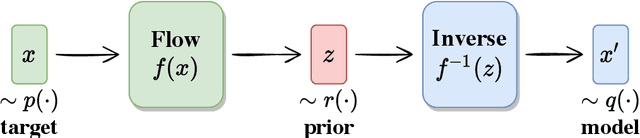
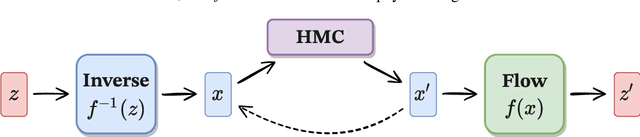
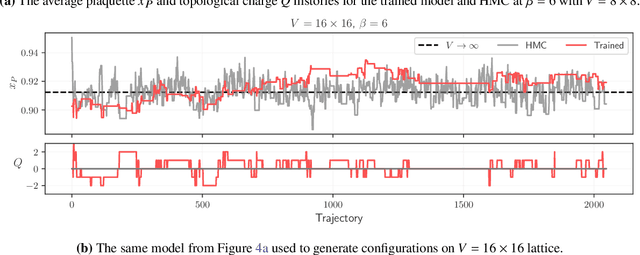
Abstract:We propose using Normalizing Flows as a trainable kernel within the molecular dynamics update of Hamiltonian Monte Carlo (HMC). By learning (invertible) transformations that simplify our dynamics, we can outperform traditional methods at generating independent configurations. We show that, using a carefully constructed network architecture, our approach can be easily scaled to large lattice volumes with minimal retraining effort. The source code for our implementation is publicly available online at https://github.com/nftqcd/fthmc.
LeapfrogLayers: A Trainable Framework for Effective Topological Sampling
Dec 02, 2021
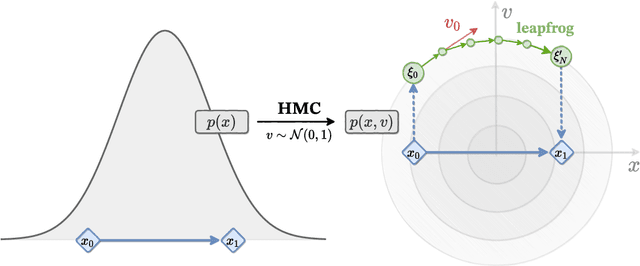
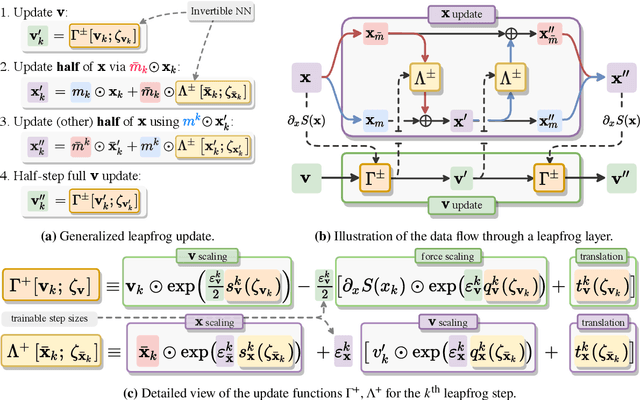
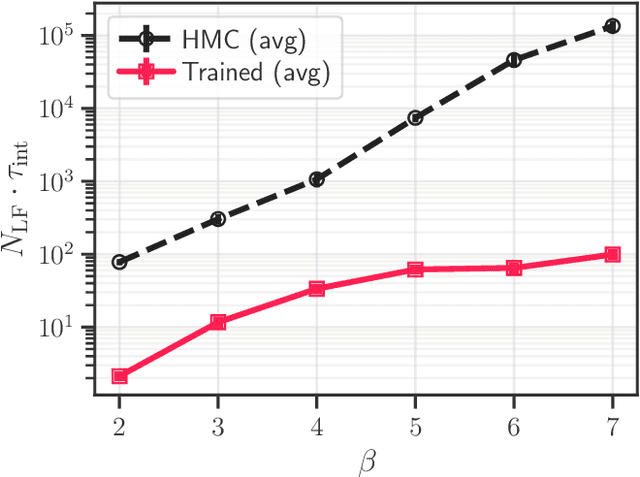
Abstract:We introduce LeapfrogLayers, an invertible neural network architecture that can be trained to efficiently sample the topology of a 2D $U(1)$ lattice gauge theory. We show an improvement in the integrated autocorrelation time of the topological charge when compared with traditional HMC, and propose methods for scaling our model to larger lattice volumes. Our implementation is open source, and is publicly available on github at https://github.com/saforem2/l2hmc-qcd
Deep Learning Hamiltonian Monte Carlo
May 07, 2021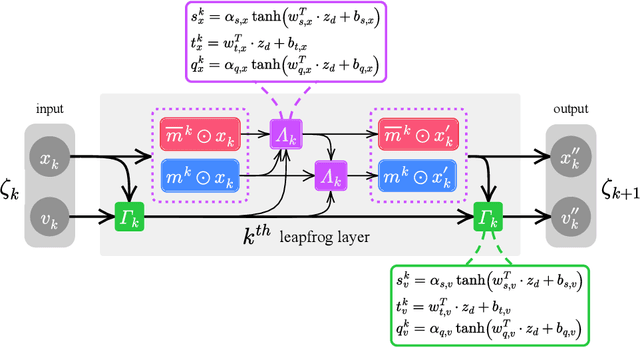
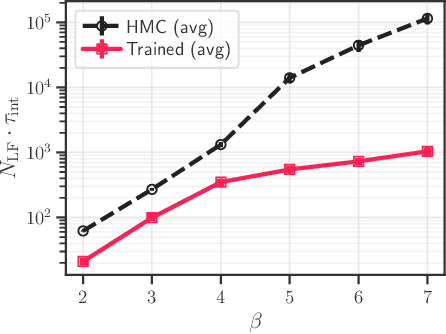

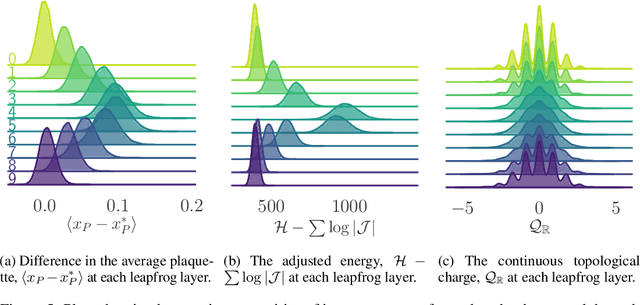
Abstract:We generalize the Hamiltonian Monte Carlo algorithm with a stack of neural network layers and evaluate its ability to sample from different topologies in a two dimensional lattice gauge theory. We demonstrate that our model is able to successfully mix between modes of different topologies, significantly reducing the computational cost required to generated independent gauge field configurations. Our implementation is available at https://github.com/saforem2/l2hmc-qcd .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge